Ligandability Mapping for Difficult Targets and Binding-Site Discovery
Many discovery programs begin with a biologically important target, but the next question is often harder: can the target be engaged by a useful small molecule, fragment, probe, or covalent ligand? Ligandability mapping is designed to answer this early question with experimental evidence.
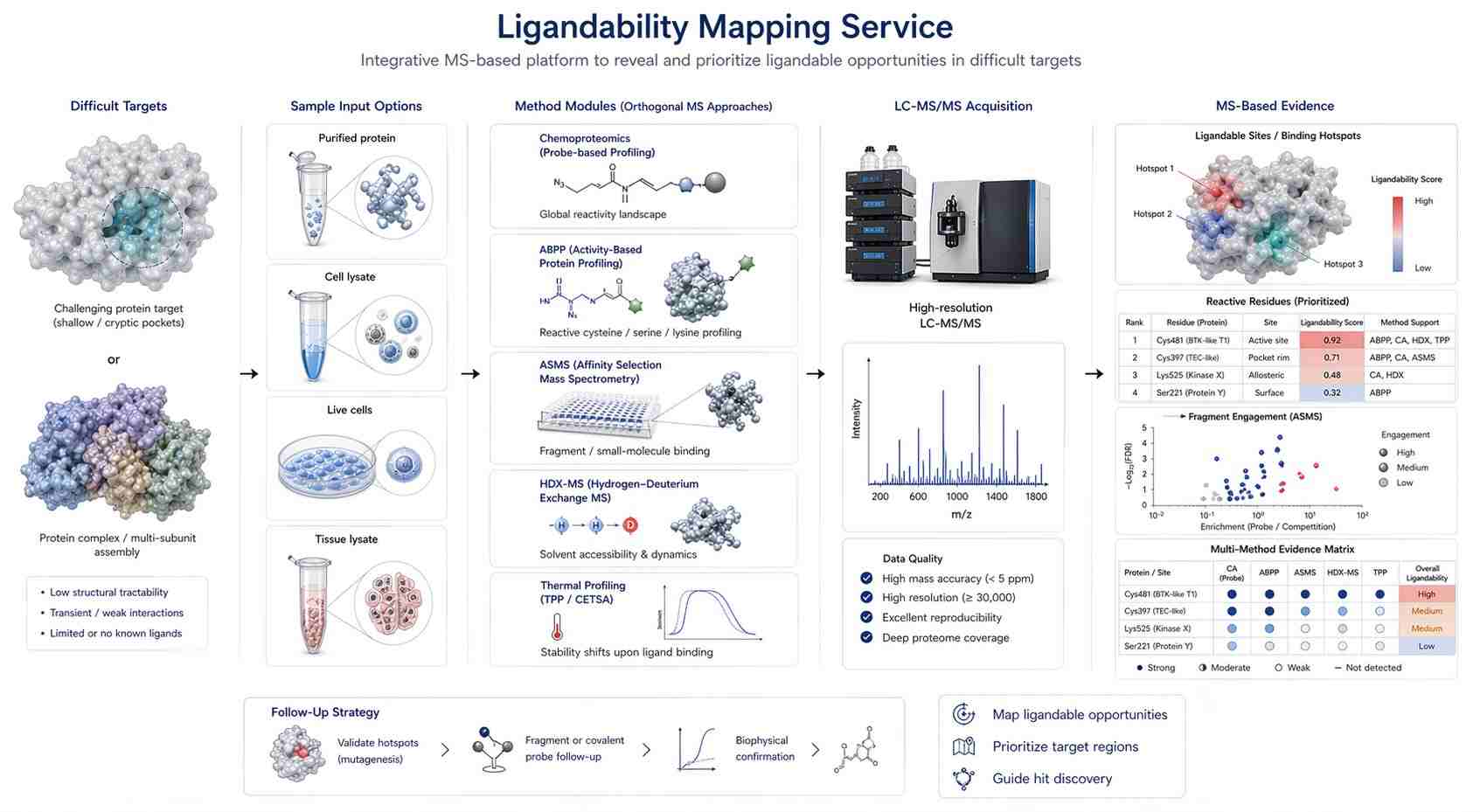
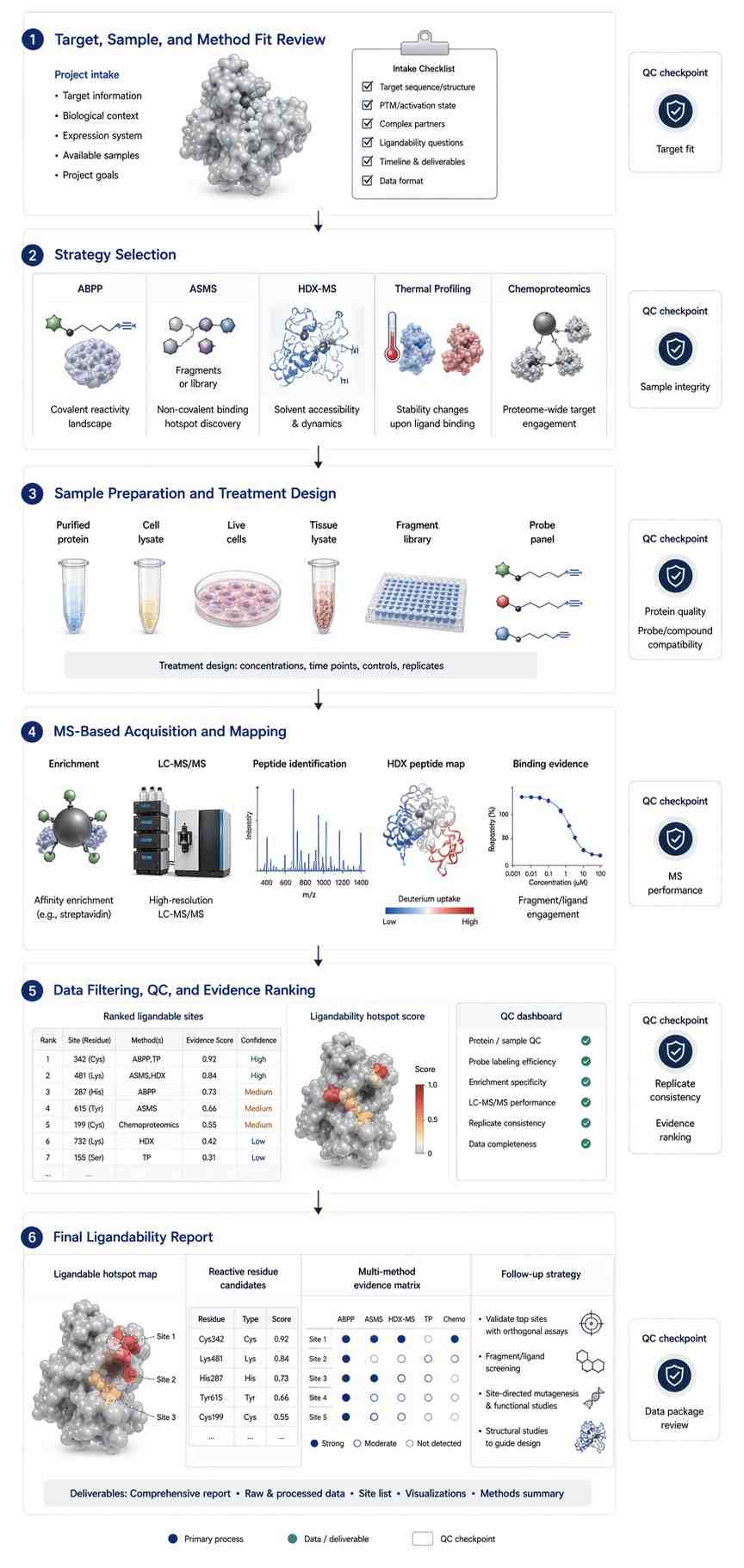
Our ligandability mapping service helps you evaluate whether a target, protein region, reactive residue, or binding hotspot shows evidence of ligand engagement. Instead of relying only on a predicted pocket or a single assay readout, we use MS-based approaches to build a practical evidence map around the target.
This service is especially useful for difficult targets, protein complexes, targets without mature functional assays, reactive residue discovery, fragment-based programs, and covalent ligand discovery.
What Ligandability Mapping Can Reveal
A well-designed ligandability mapping study can help answer several practical questions:
- Does the target contain a site that can be engaged by a ligand?
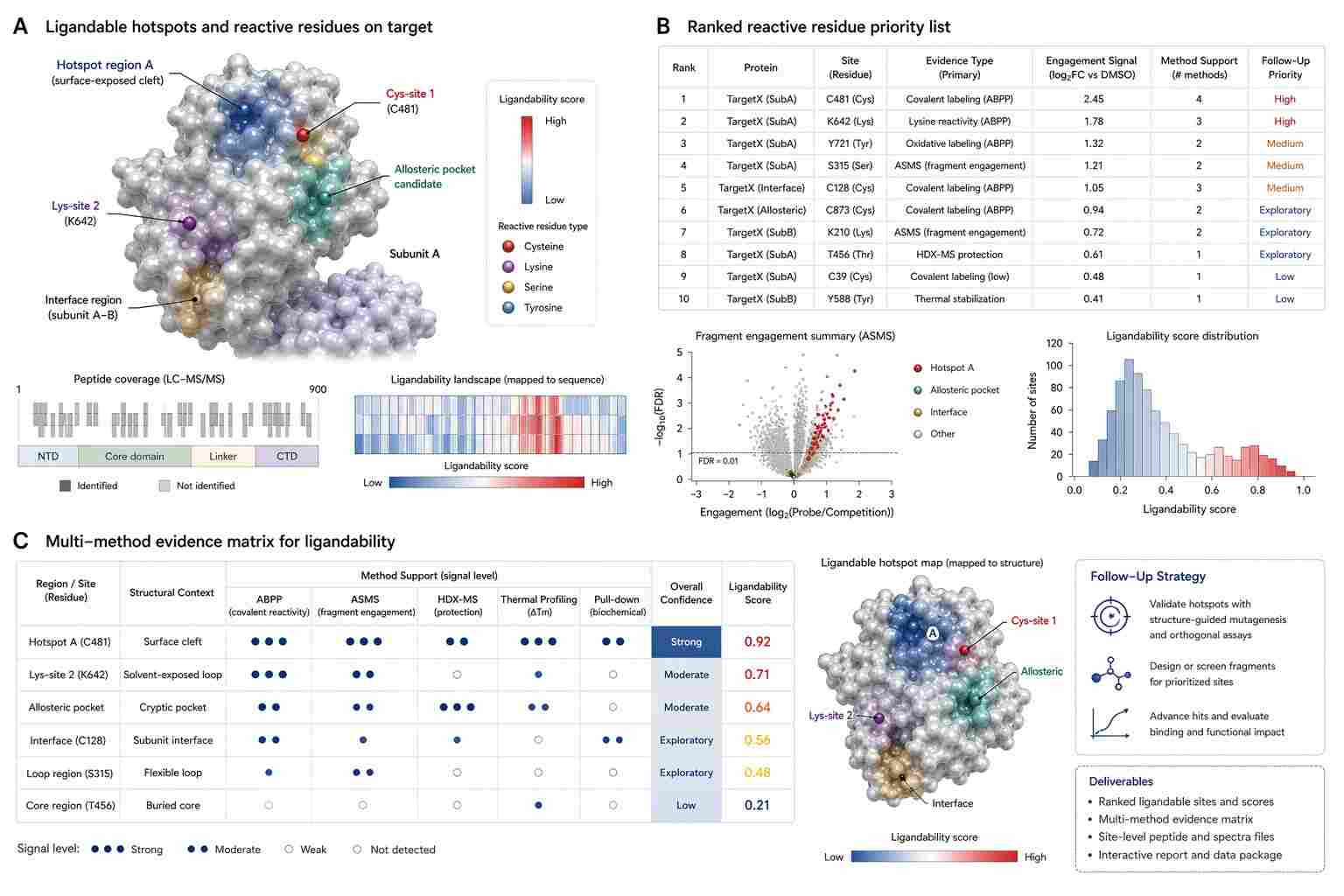
- Are there reactive residues that may support covalent ligand discovery?
- Can weak fragments or early hits be connected to a binding region?
- Is the evidence protein-level only, or does it support a specific peptide, residue, or hotspot?
- Which method is most appropriate: ABPP, reactive residue profiling, ASMS, HDX-MS, thermal profiling, or another MS-based strategy?
- Which sites or target regions should be prioritized for follow-up screening or validation?
For many teams, the most useful output is not simply a list of proteins. It is a ranked view of ligandable opportunities, evidence strength, sample context, and next-step recommendations.
Ligandability Is Not the Same as Druggability
Ligandability and druggability are related, but they are not the same. Ligandability asks whether a site, protein region, or target can be engaged by a ligand. Druggability requires additional downstream evidence, such as potency, selectivity, cellular activity, functional relevance, and developability.
We treat ligandability mapping as an early evidence layer. It can help your team decide whether a target is worth deeper screening, which site or region deserves attention, and which follow-up method should come next. It does not replace later optimization or functional validation, but it can reduce uncertainty before larger discovery investments.