This literature-supported case study is based on van Rooden et al., Mapping in vivo target interaction profiles of covalent inhibitors using chemical proteomics with label-free quantification. It is not a Creative Proteomics customer case.
Background
The paper addresses a central problem in covalent inhibitor development: researchers need to understand both intended target engagement and off-target activity in biological systems. The authors describe ABPP as a chemical proteomics method that can guide covalent drug development by assessing on-target engagement and off-target activity.
The article also discusses why this matters for safety and interpretation. The authors previously used ABPP to determine the serine hydrolase interaction landscape of BIA 10-2474, an experimental drug, and reported that this provided a potential explanation for the adverse side effects observed with that compound. The protocol then focuses on a label-free quantitative proteomics workflow that can compare biological samples and map inhibitor interaction profiles in native proteomes.
Methods
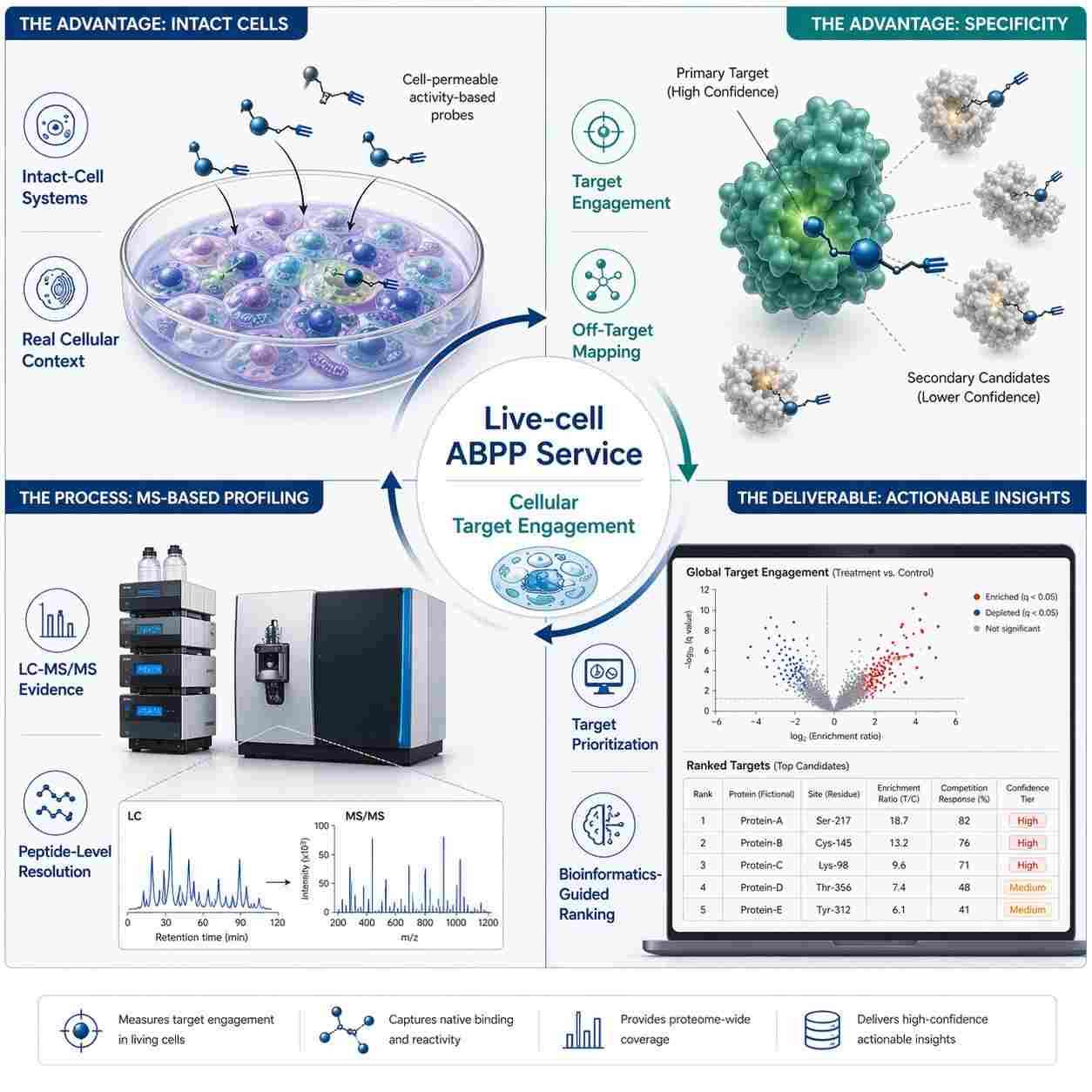
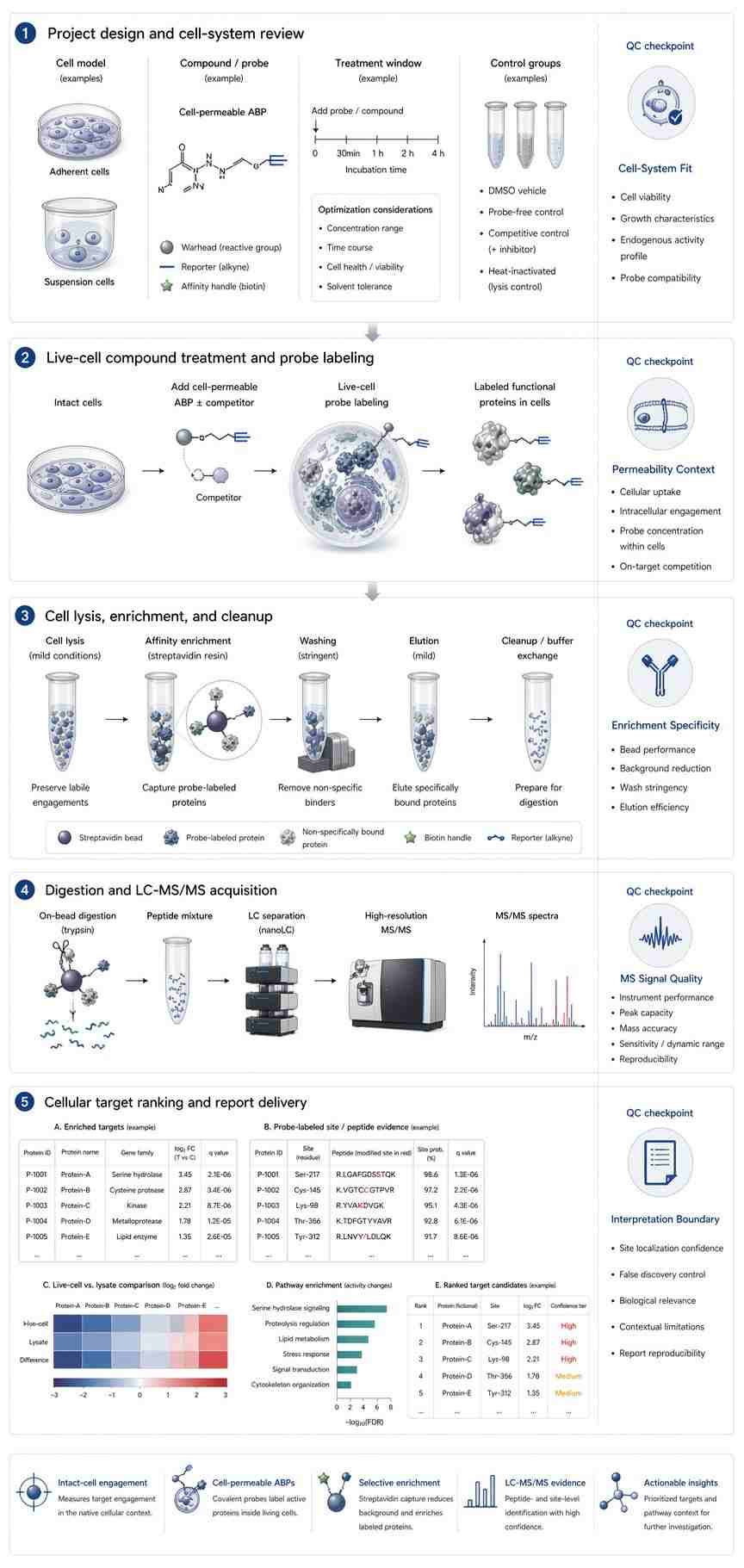
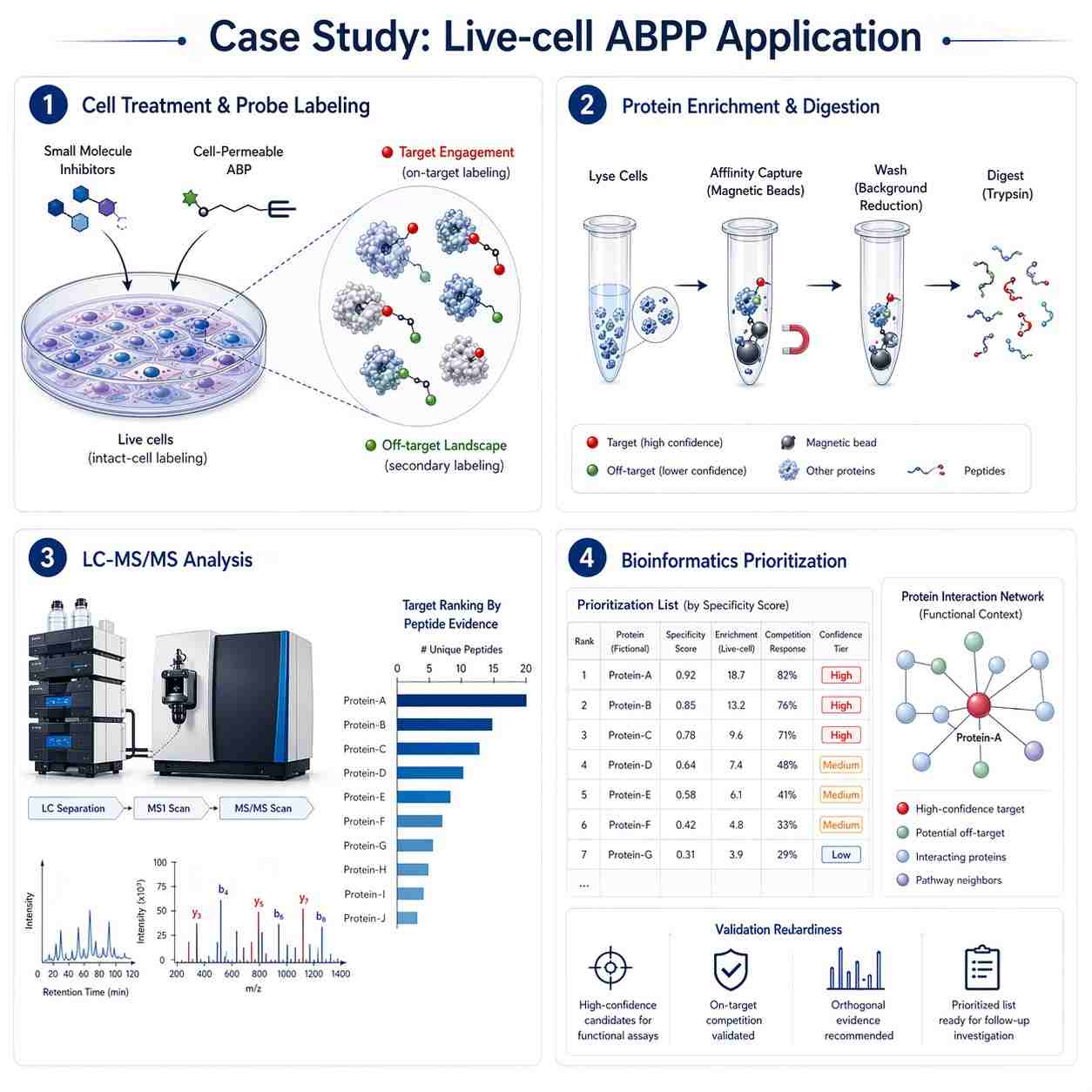
The protocol uses chemical proteomics with label-free quantification to identify the in vivo selectivity profile of covalent inhibitors. The workflow includes tissue lysis, probe incubation, target enrichment, sample preparation, LC-MS measurement, data processing, and analysis. The authors describe this approach as suitable for studying target engagement in a native proteome and identifying potential off-targets.
For the demonstrated experiment, the authors assessed the protein interaction landscape of the diacylglycerol lipase inhibitor DH376 in mouse brain, liver, kidney, and testes. Figure 1 presents the chemical proteomics workflow, including inhibitor and probe structures used in the study. Figure 2 presents results from the competitive ABPP experiment in mice treated with DH376 and a probe cocktail, including hierarchical clustering of probe targets. Figure 3 shows protein and peptide abundance data for selected probe targets.
The paper also provides public proteomics data access through the Proteomics Identifications Database accession PXD007965, which strengthens transparency and reuse.
Results
The article reports that ABPP can map protein interaction landscapes of inhibitors in cells, tissues, and animal models. In this protocol, the authors demonstrate the method by profiling DH376 across four mouse tissues: brain, liver, kidney, and testes.
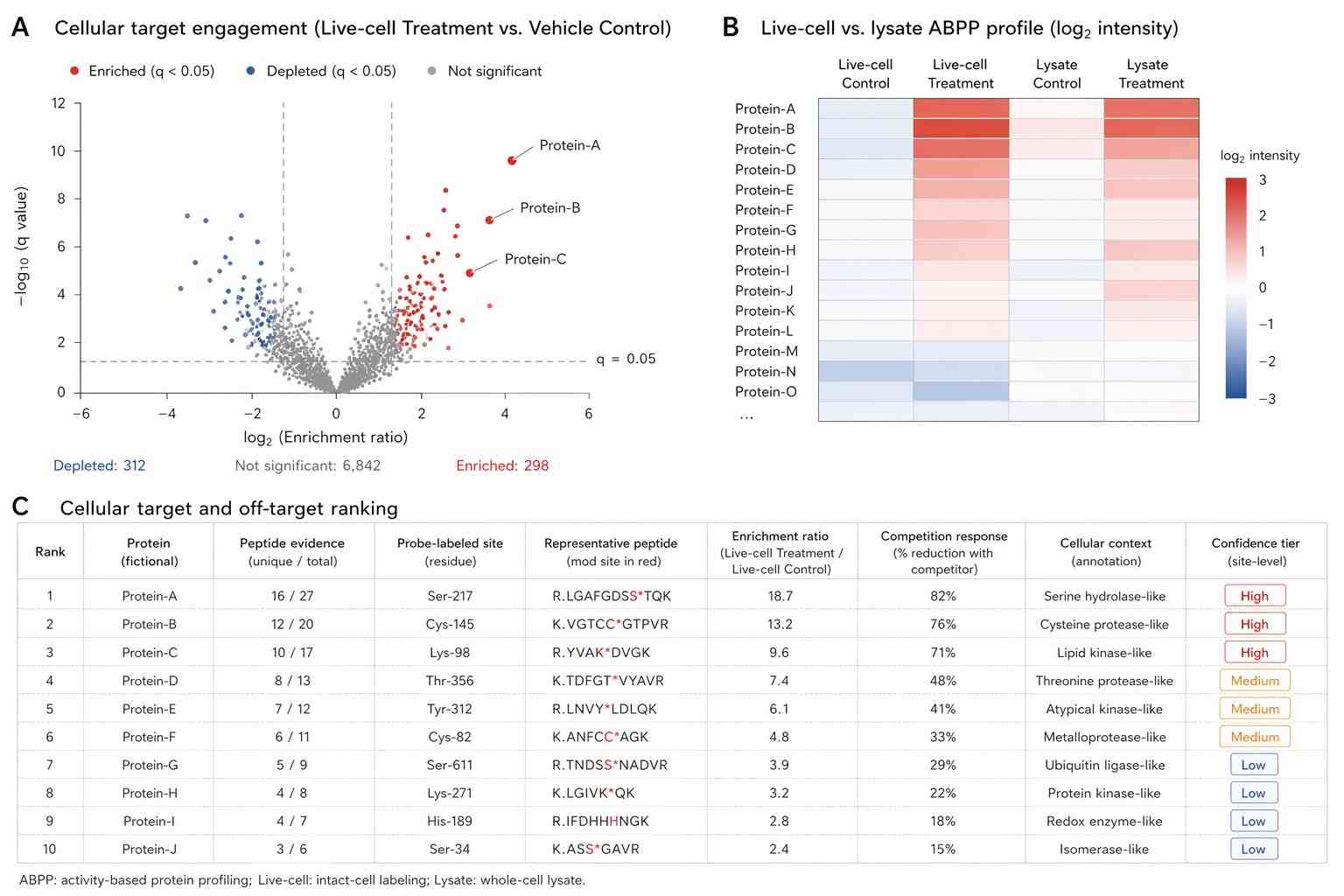
Figure 2 is the key result figure for the demonstrated competitive ABPP experiment. It shows how DH376 treatment affects probe-target activity patterns and uses hierarchical clustering to organize probe targets across tissues. This figure supports the central idea that ABPP can move beyond a single target and reveal a broader target interaction landscape.
The paper also links this workflow to a prior BIA 10-2474 study. The authors state that ABPP was used to determine the serine hydrolase interaction landscape of BIA 10-2474, which provided a potential explanation for adverse side effects. This is important because it shows how ABPP can help identify not only expected target engagement but also possible off-target activity.
The protocol emphasizes that optimization for label-free quantification results in high proteome coverage and allows comparison across multiple biological samples. For service-page interpretation, the important scientific point is that the workflow includes controlled stages from sample processing to LC-MS and data analysis.
Conclusion
The authors conclude that ABPP with label-free quantitative proteomics can be used to study target engagement in native proteomes and identify potential off-targets for covalent inhibitors. Their example with DH376 shows how inhibitor interaction landscapes can be assessed across multiple tissues, while the BIA 10-2474 context shows how ABPP can help explain off-target liability.
For live-cell ABPP service planning, this case supports three practical lessons. First, target engagement should be evaluated in biologically relevant systems instead of relying only on simplified biochemical assays. Second, LC-MS/MS chemoproteomics can reveal a broader interaction landscape than single-target testing. Third, controls, tissue or cell context, probe behavior, and quantitative analysis are essential for separating likely engagement from background or indirect effects.