Background
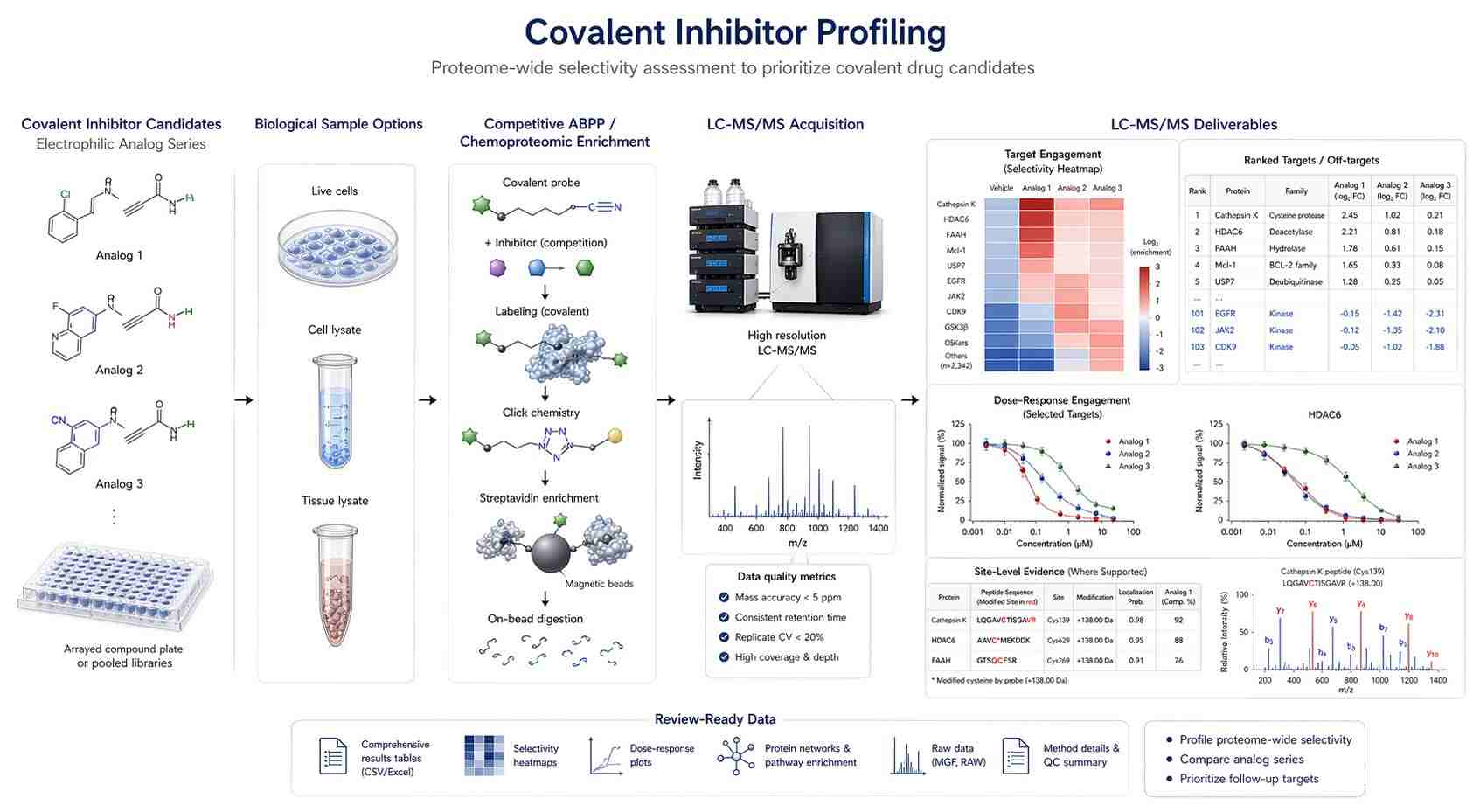
Covalent inhibitors are increasingly important in drug discovery, but their development requires careful balancing of potency and selectivity. A reactive warhead can support durable target engagement, but excessive or poorly controlled reactivity can increase off-target labeling across the proteome.
Lin and colleagues addressed this challenge in COOKIE-Pro: covalent inhibitor binding kinetics profiling on the proteome scale. The study introduced COOKIE-Pro, a proteomics workflow designed to quantify irreversible covalent inhibitor binding kinetics and selectivity across the proteome.
Methods
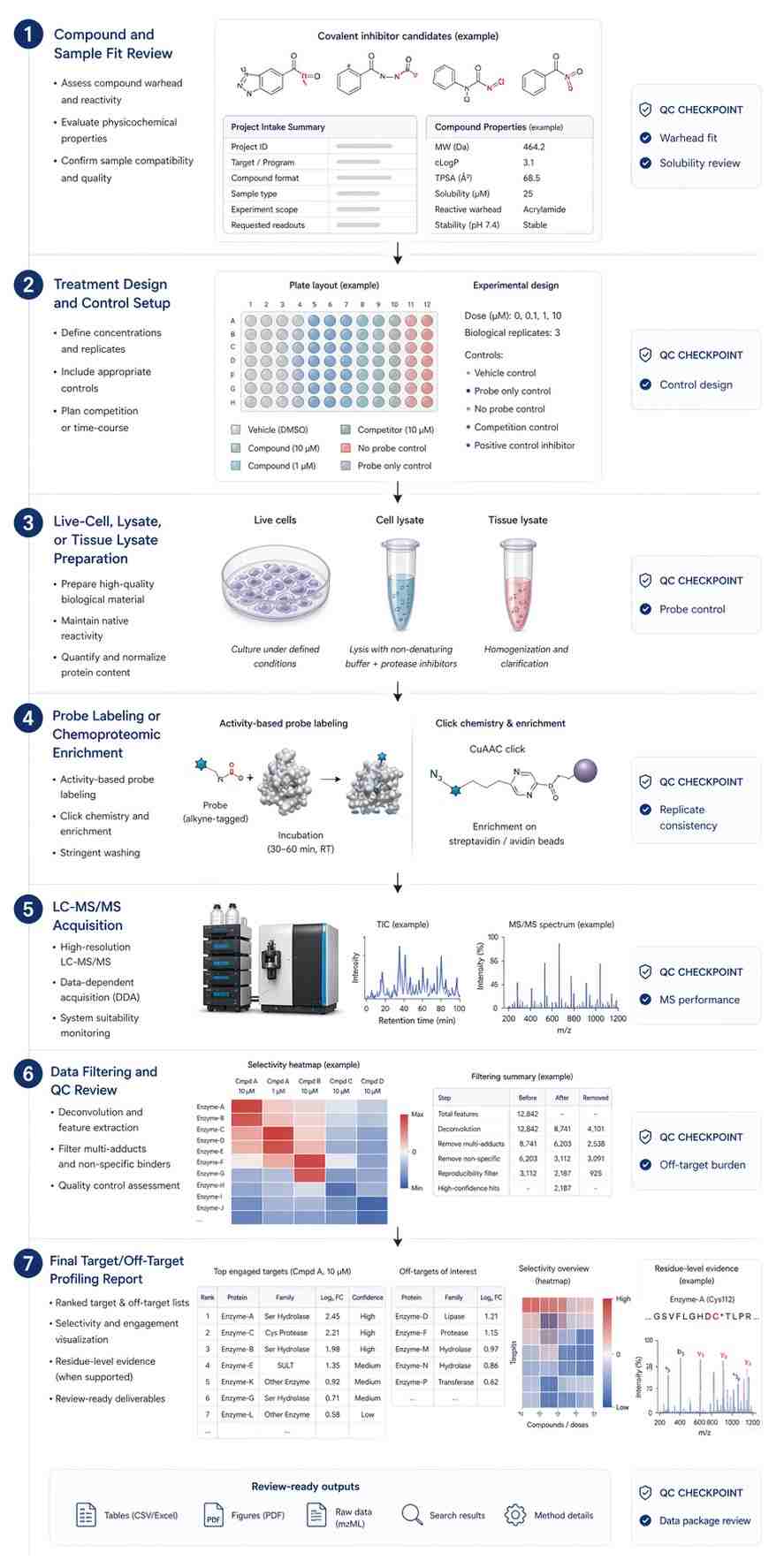
The study used a two-step covalent occupancy kinetics enrichment strategy. In the first step, permeabilized cells were preincubated with a covalent inhibitor at defined concentrations and time points. In the second step, a desthiobiotin probe was used to enrich unoccupied target and off-target proteins. The enriched proteins were then analyzed by mass spectrometry-based proteomics.
The authors used BTK inhibitors, including spebrutinib and ibrutinib, to validate the approach. They also used controls such as DMSO and saturating inhibitor conditions to support occupancy interpretation. TMT-based quantification helped compare treatment groups and reduce injection-to-injection variation.
Results
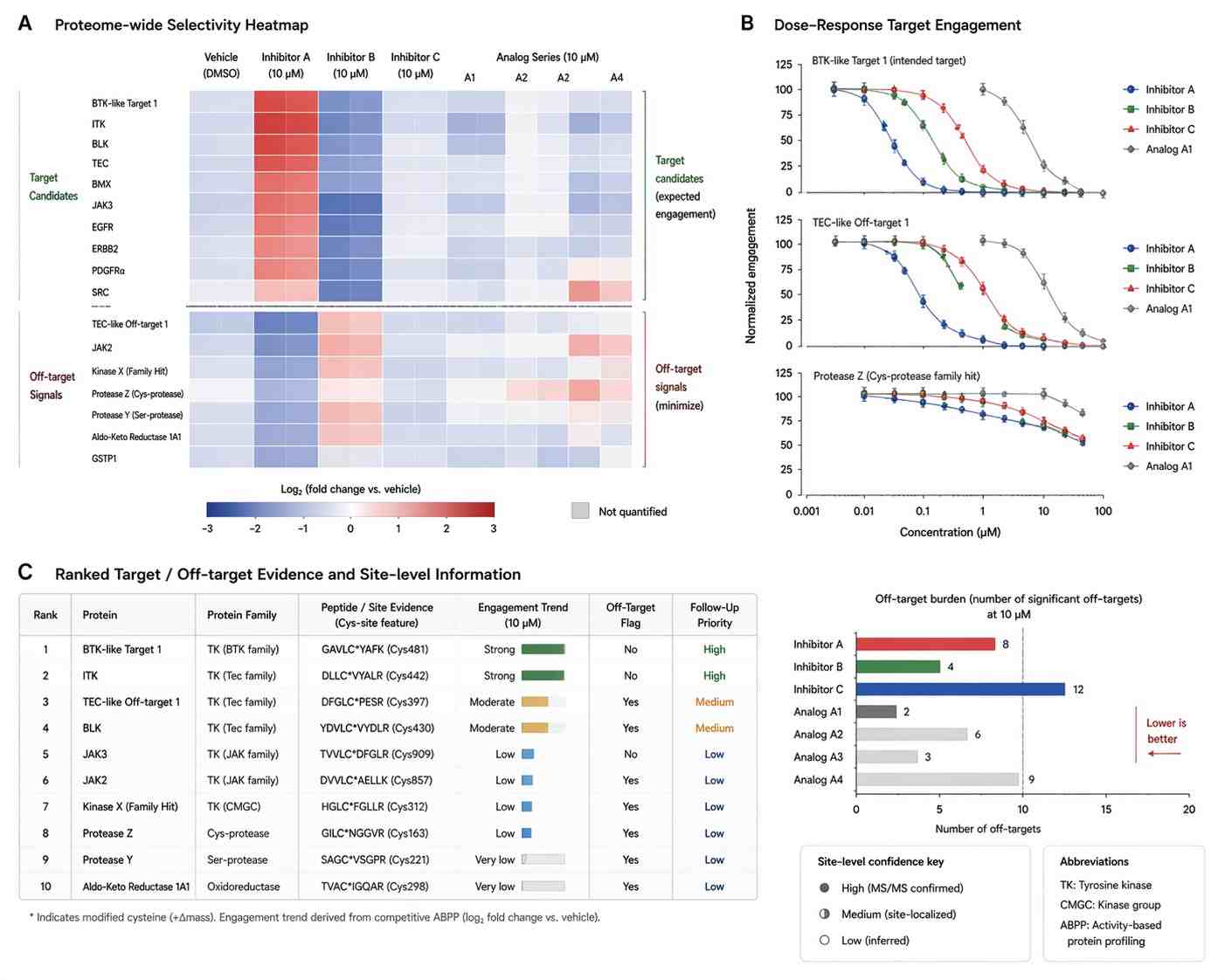
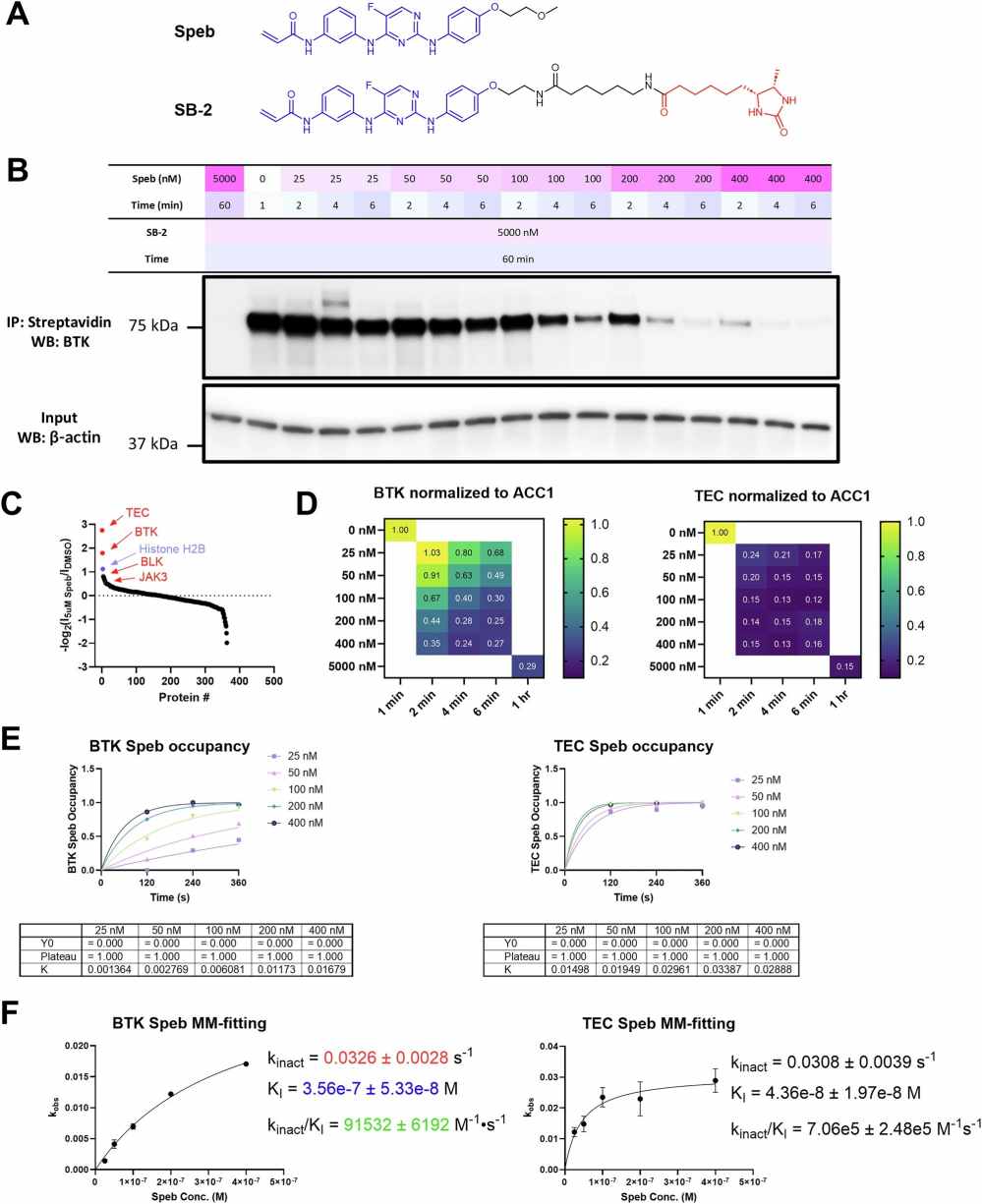
The paper reported that COOKIE-Pro could profile covalent inhibitor binding kinetics and selectivity on the proteome scale. In the abstract, the authors state that the method reproduced known kinetic parameters for BTK inhibitors and identified both expected and unreported off-targets. They also reported that spebrutinib showed more than 10-fold higher potency for TEC kinase compared with its intended target BTK.
Figure 3 focuses on COOKIE-Pro profiling for spebrutinib in Ramos cells using the SB-2 desthiobiotin probe. It includes the spebrutinib and SB-2 probe structures, the experiment layout, a ranked proteomics off-target list, kinase off-target heatmaps, and kinetic fitting for BTK and TEC. In this case, the figure helps show how a covalent inhibitor profiling workflow can move from compound treatment to off-target ranking and kinetic interpretation.
The same study also applied a streamlined two-point strategy to a library of 16 covalent fragments. This generated thousands of kinetic profiles and supported quantitative separation of intrinsic chemical reactivity from binding affinity at scale.
Conclusion
This case shows why covalent inhibitor profiling should not rely only on single-target potency. A useful profiling workflow should connect target engagement, off-target ranking, concentration or time behavior, and data interpretation. For discovery teams, the value is not only identifying whether a compound works, but understanding how selectively it works across the proteome and which findings should guide the next round of chemistry or validation.