El-Nashar HAS, Eldahshan OA, El Hassab MA, Zengin G, Elhawary EA. "UPLC/MSn analysis of Bougainvillea glabra leaves and investigation of antioxidant activities and enzyme inhibitory properties." Scientific Reports 15:28272 (2025). https://doi.org/10.1038/s41598-025-11851-9
Background
Bougainvillea glabra is an ornamental plant with documented traditional uses, but its detailed phytochemical profile remained incompletely characterized. The authors aimed to apply UPLC/MSn for comprehensive structural elucidation of its secondary metabolites, including flavonoids, phenolic acids, and other compound classes.
Methods
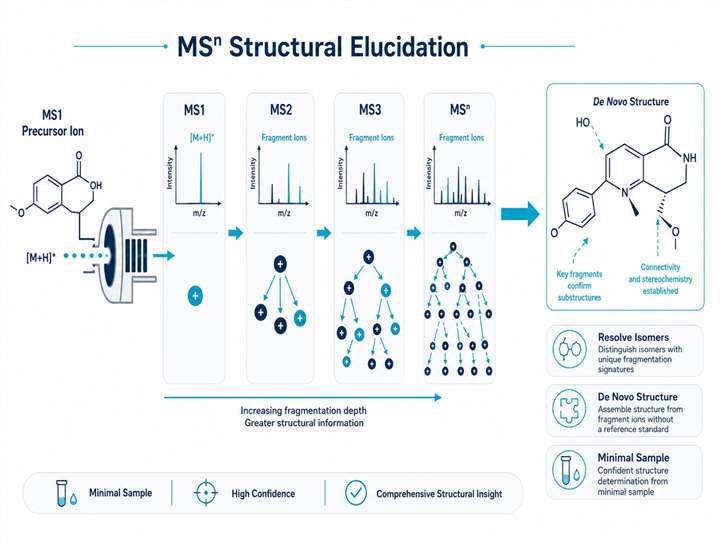
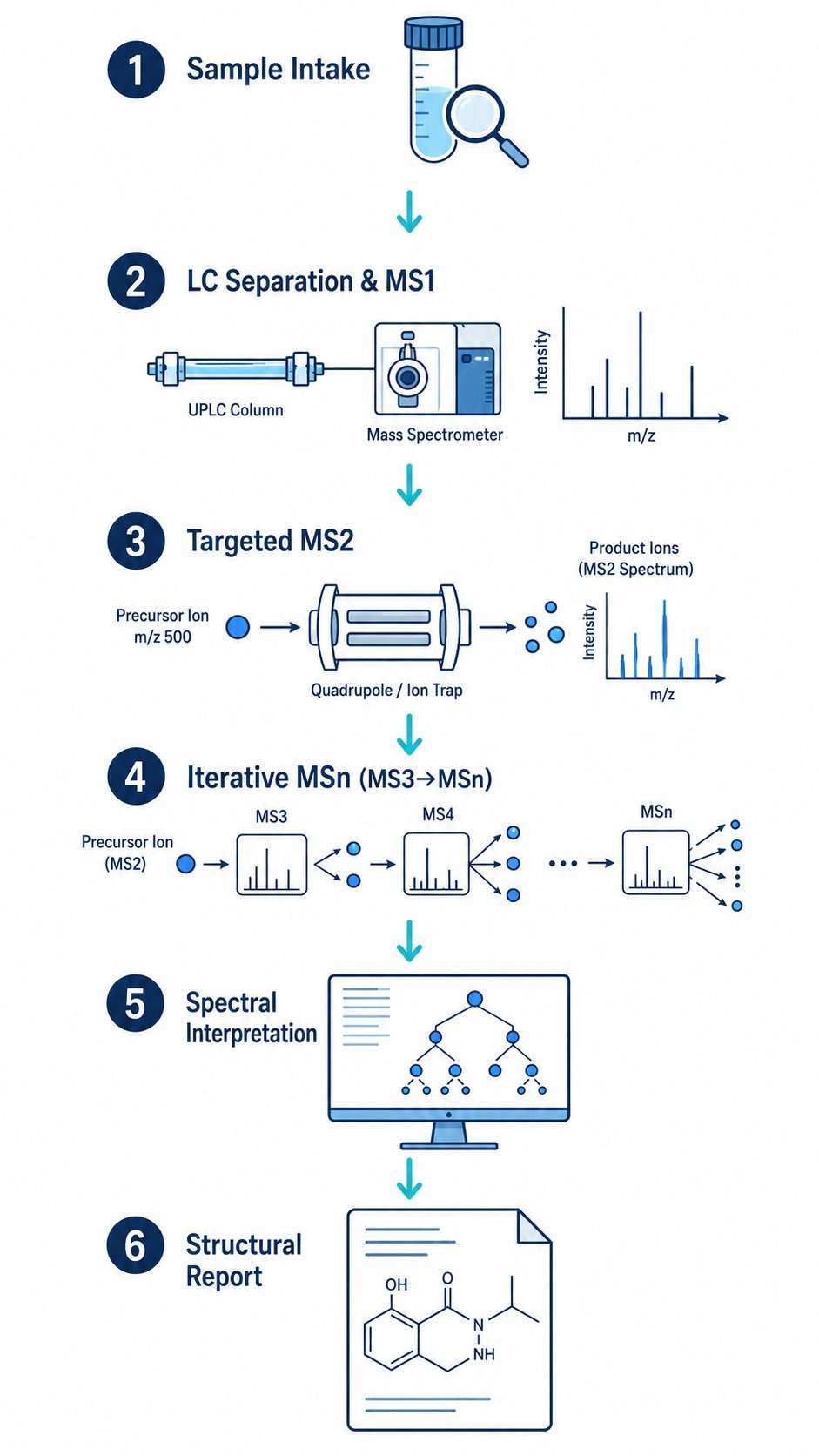
Methanol leaf extracts were analyzed using UPLC-ESI-MS/MS in both positive and negative ionization modes. The MSn workflow involved:
- Full-scan MS1 acquisition for precursor ion detection and molecular formula assignment
- Data-dependent MS2 fragmentation of detected precursor ions
- Manual interpretation of MSn fragmentation patterns for structural assignment
- Comparison of retention times, molecular ions, and fragment ion series with published literature data
Results
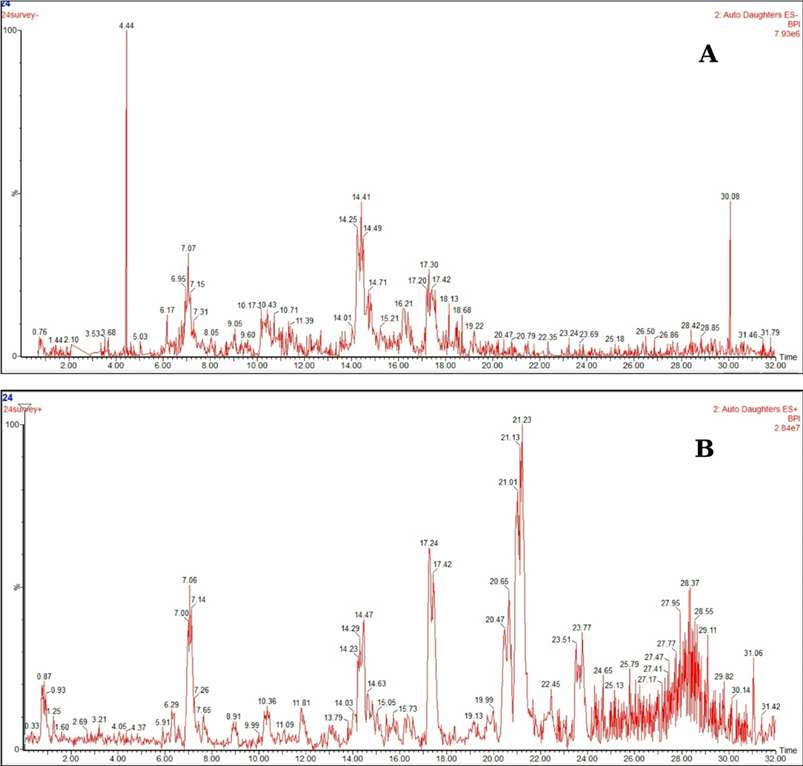
The UPLC/MSn analysis detected 23 chromatographic peaks, of which 21 compounds were tentatively identified (Fig. 1). Key structural assignments included:
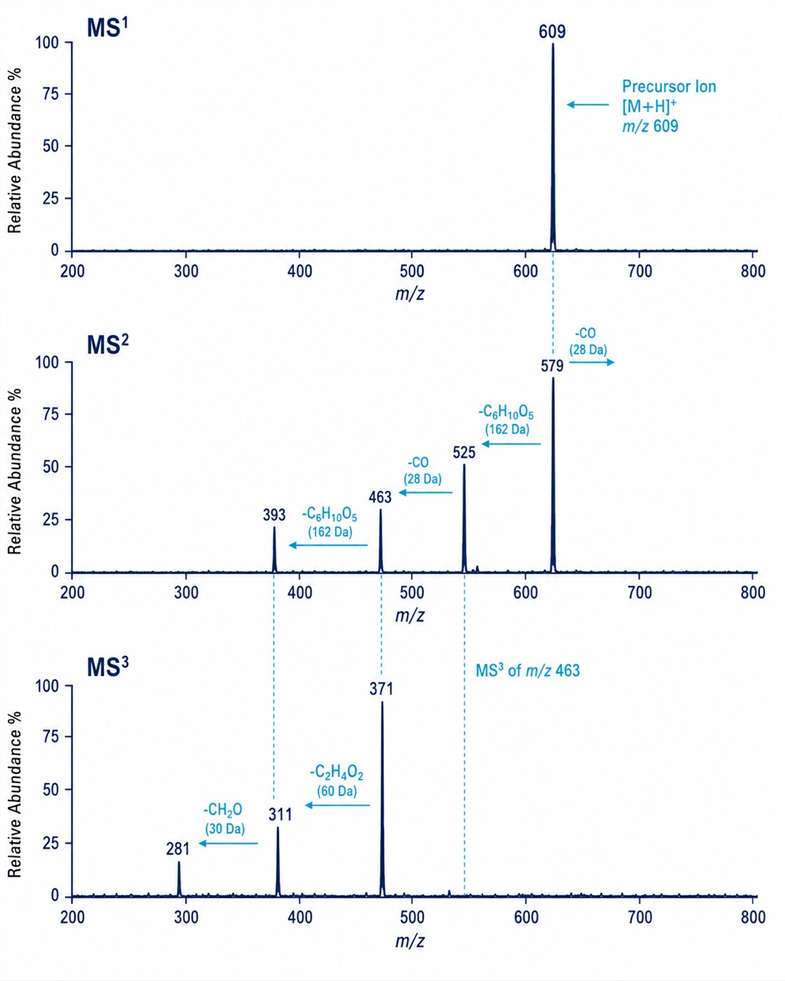
- Flavonoids: luteolin robinobioside (m/z 593 → fragment at m/z 285 confirming luteolin aglycone), rhamnocitrin-O-rutinoside (m/z 609 → MSn fragments at m/z 579, 525, 463, 393, 371, 311, 281), apigenin-C-hexoside-C-pentoside (m/z 563), and biflavonoids including di-hydro-bi-apigenin methyl ether (m/z 553) and manniflavanone (m/z 589)
- Phenolic acids: caffeic acid hexoside (m/z 341), tri-O-caffeoyl shikimic acid (m/z 659), chlorogenic acid (m/z 353), and rosmarinic acid (m/z 359)
- Other classes: (+)-syringaresinol hexoside (lignan, m/z 579), delphinidin hexosyl pentosyl malonate (anthocyanin, m/z 683), oleanolic acid (triterpene, m/z 455), and speedoside (iridoid, m/z 685)
The fragmentation patterns at each MSn stage provided diagnostic ions that distinguished between closely related structural isomers — for example, the specific fragment series of tri-O-caffeoyl shikimic acid (m/z 571, 475, 397, 329, 299, 285) confirmed the sequential loss of caffeoyl moieties and established the substitution pattern.
Conclusions
The study demonstrated that UPLC/MSn is a powerful and efficient approach for the structural elucidation of diverse natural product classes from a single plant extract. The MSn fragmentation data enabled confident identification of 21 compounds across multiple chemical classes — flavonoids, phenolic acids, lignans, anthocyanins, triterpenes, and iridoids — without the need for compound isolation or NMR analysis. The authors noted that MSn provided the structural depth necessary to distinguish between isomeric flavonoid glycosides and to assign modification patterns that would have been ambiguous from MS2 data alone.