Automated Compound–Target Binding HT-MS Screening
Label-free binding detection at scale — automated affinity selection mass spectrometry for hit identification across diverse target classes.
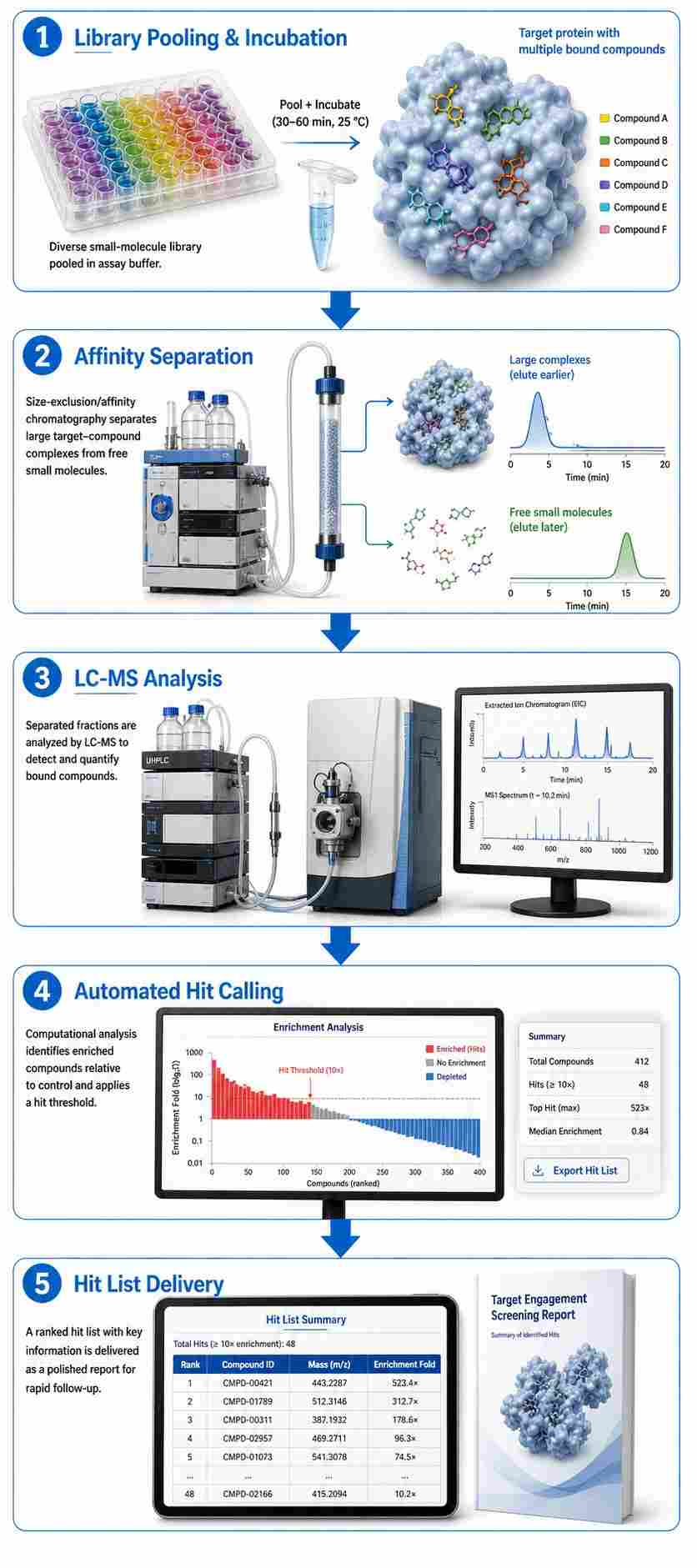
Automated compound-target binding HT-MS combines affinity selection chemistry with high-resolution mass spectrometry to directly detect ligand-target binding events from pooled compound libraries — without fluorescent tags, immobilized targets, or DNA barcodes. Our platform integrates automated liquid handling, affinity separation, and multi-plexed MS readout into a single, standardized workflow capable of screening thousands of compounds per day against soluble proteins, membrane proteins, and protein complexes.
At Creative Proteomics, our MassTarget™ HT-MS binding platform deploys multiple affinity selection formats — SEC-based, ultrafiltration-based, and bead-based — selected to match the biophysical properties of each target. The result: direct, label-free hit identification with rich mass spectrometric data for each binding event.
Core Capabilities:
- Label-free binding detection — no fluorescent tags, no immobilisation, no DNA barcodes
- Pooled library screening — 100–1,000 compounds per run with deconvolution by accurate mass
- Multiple affinity formats — SEC-based, ultrafiltration-based, and bead-based ASMS
- Broad target compatibility — soluble proteins, membrane proteins, protein complexes, RNA
- Automated workflow — from plate loading to ranked hit list with minimal hands-on time