miniTurbo Protein Interaction Mapping Service for Drug Discovery
Some interactions don't wait around. Capture them in minutes, not hours.
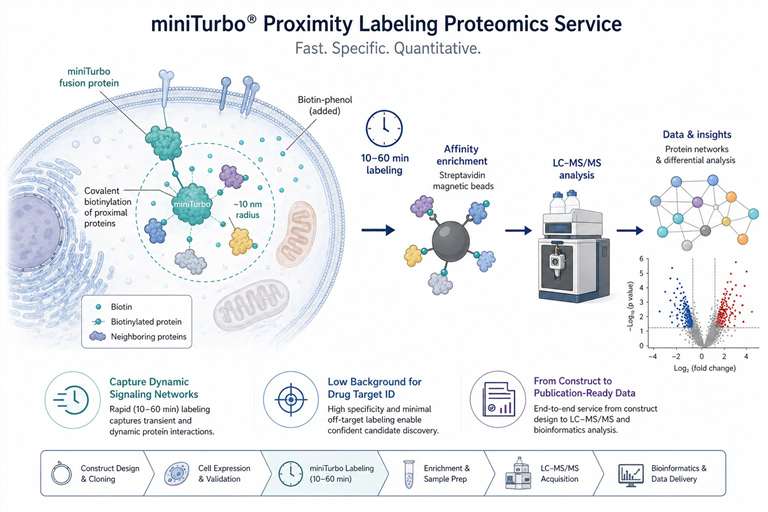
Our miniTurbo proximity labeling proteomics service delivers fast, low-background interactome maps for dynamic signaling networks and PROTAC targets.
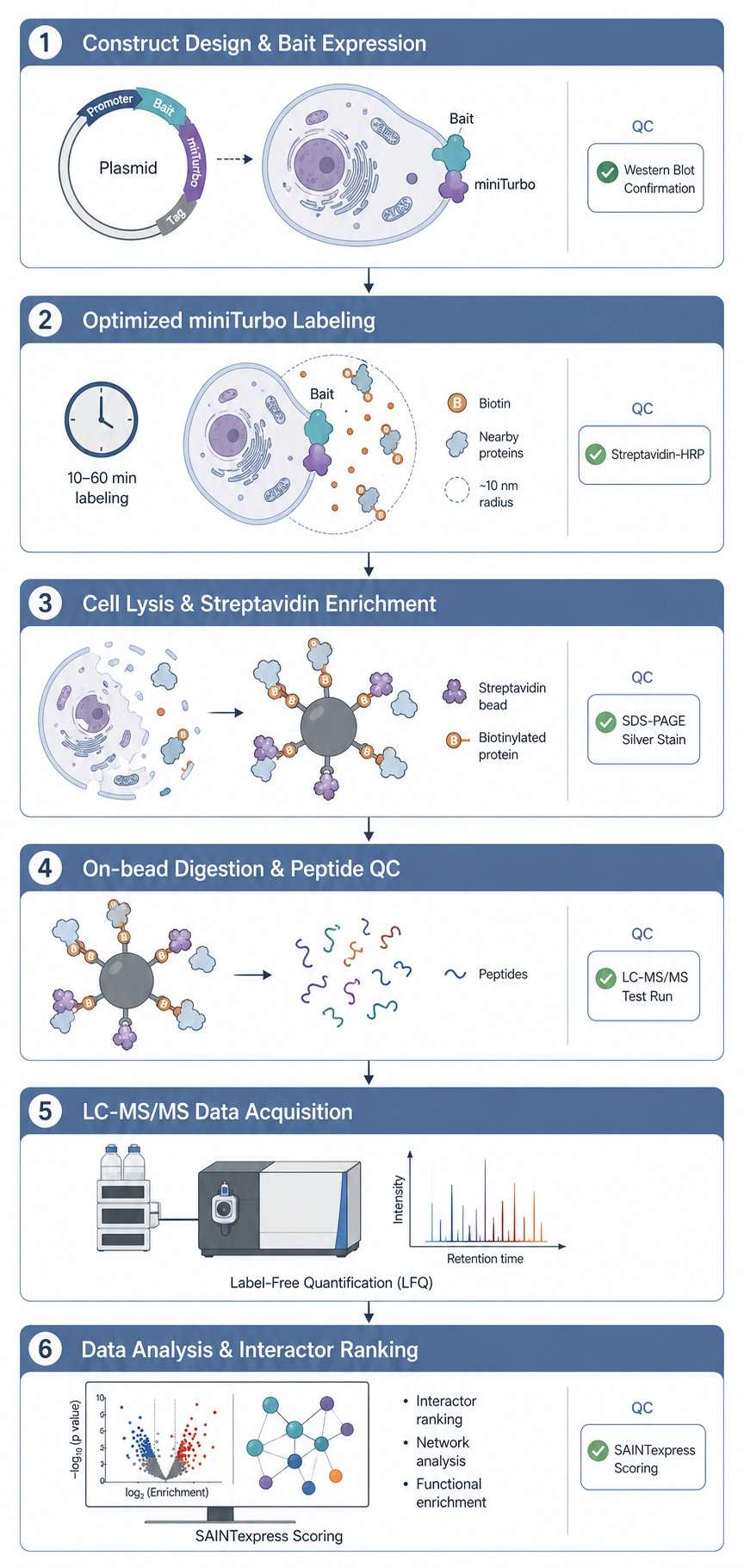
From construct design to publication-ready data, we help you map the interactions that matter—with clear QC at every step.
Some interactions don't wait around. A kinase touches its substrate for seconds. A PROTAC brings two proteins together just long enough to tag one for degradation. If your labeling window is 18 hours, you miss it.
That's where miniTurbo changes the equation. It's the fastest, lowest-background proximity labeling enzyme we work with, and we've built a full service around it. From your construct to a publication-ready interaction network, the result isn't just a list of proteins. It's a snapshot of what was actually happening in your cells during that narrow, biologically relevant window—the one your drug was designed to perturb.
Key Advantages:
- 10–60 min labeling captures dynamic interactors
- Low background, high-confidence hits
- Optimized for low-input, complex samples
- End-to-end service, raw data delivered