Zheng, X., et al. "Collision Cross Section (CCS) Measurement and Prediction Methods in Omics." (2023) https://pmc.ncbi.nlm.nih.gov/articles/PMC10530098/
Background
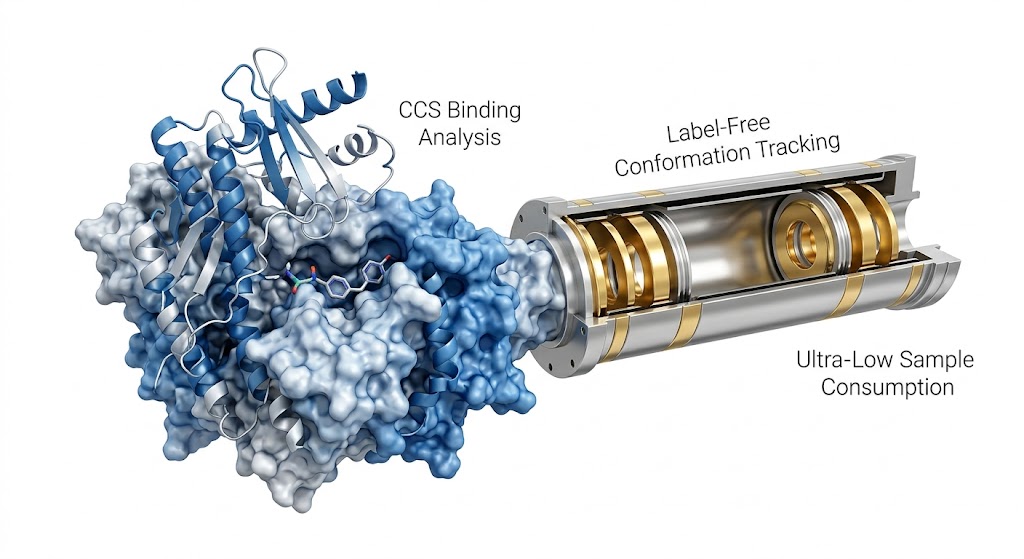
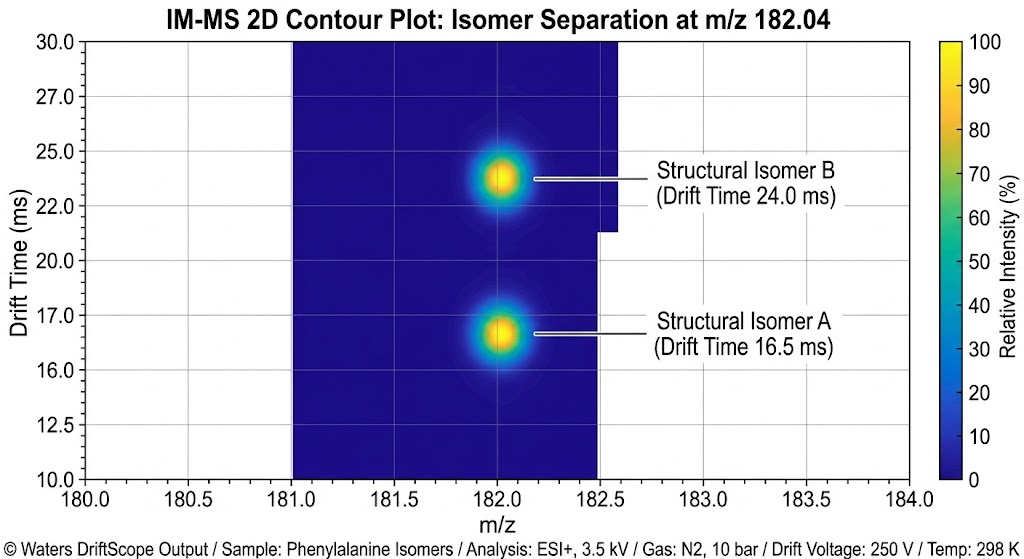
Understanding the extreme structural diversity of biomolecules is often limited by the resolving power of traditional analytical techniques. In modern omics and drug discovery, researchers frequently encounter highly complex isomeric mixtures or large, non-covalent multi-protein assemblies. Because these different structural states often share identical mass-to-charge (m/z) ratios, standard mass spectrometry cannot distinguish between them, leaving critical structural dynamics hidden.
Methods
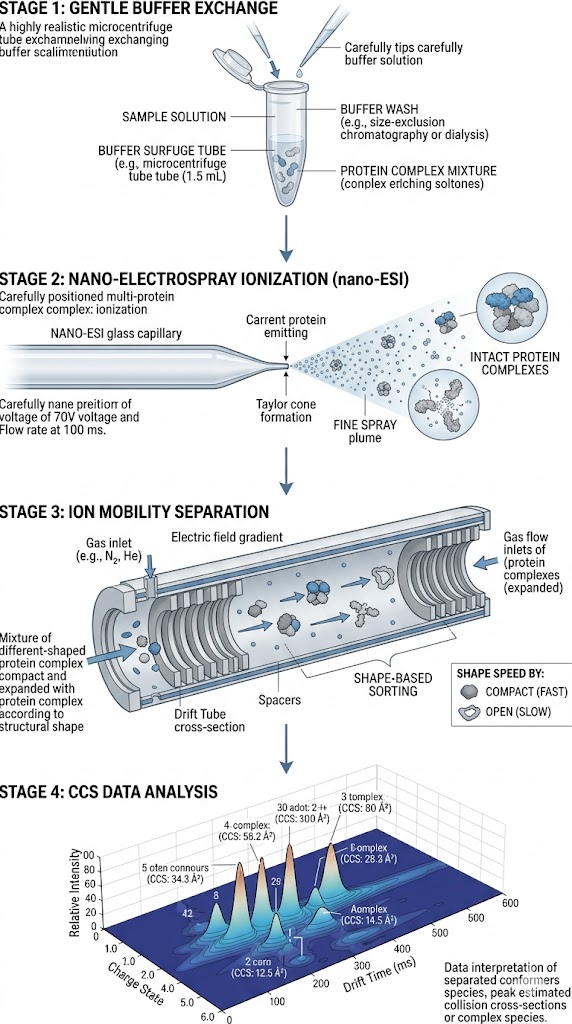
To overcome this fundamental limitation, researchers utilized Ion Mobility-Mass Spectrometry (IM-MS) combined with predictive modeling. This multidimensional approach separates ions not just by their mass, but by their physical size, shape, and charge distribution (their collision cross-section) as they travel through a gas-filled drift region. By doing so, the team could simultaneously acquire mass spectra and ion mobility drift times, allowing for a highly detailed mapping of the conformational landscape.
Results
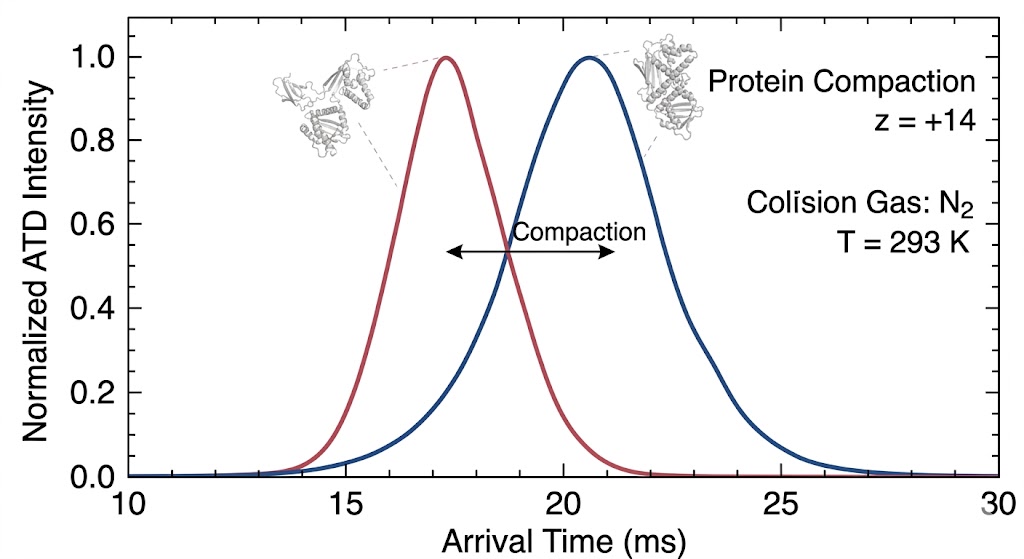
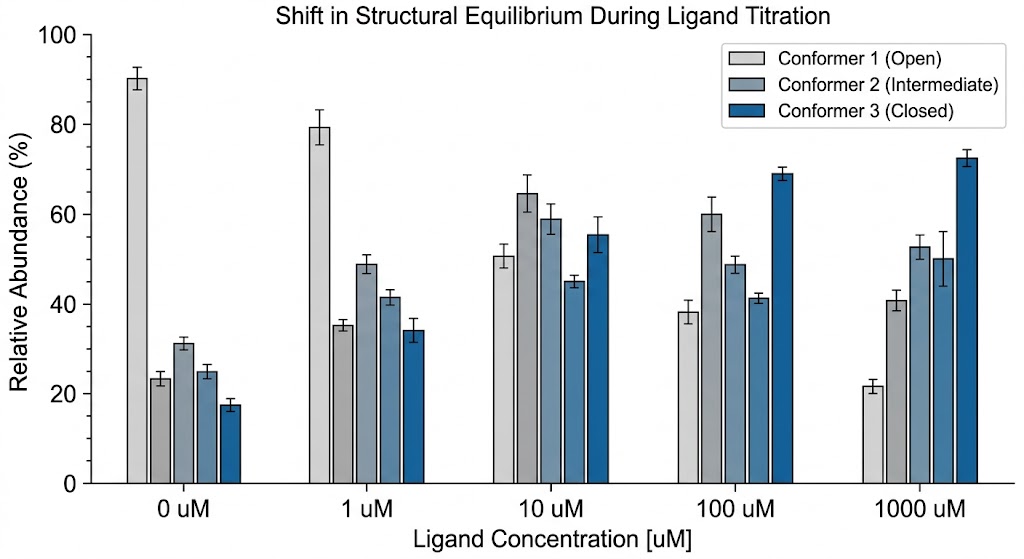
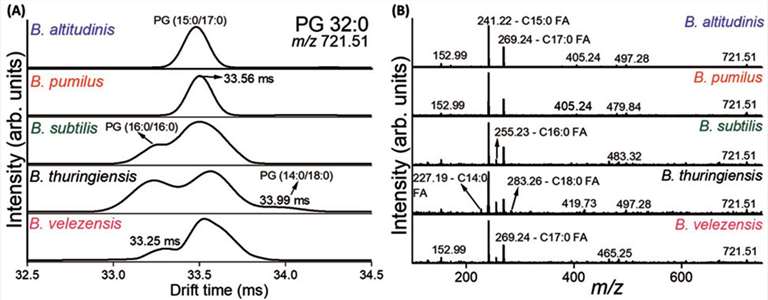
As detailed in Figure 2 of the published study, the IM-MS data clearly illustrates the successful separation of structural conformers that could not be resolved by standard MS alone. The resulting Arrival Time Distribution (ATD) profiles generated distinct, measurable peaks for different folding states. This provided unique, quantifiable CCS signatures for the distinct binding states and molecular architectures present in the complex mixture.
Conclusions
Accurate CCS measurement serves as a highly robust orthogonal descriptor in structural biology. It validates dynamic conformational changes, resolves heterogeneous mixtures, and significantly enhances confidence in molecular target interactions during early-stage discovery, proving invaluable for characterizing targets that evade traditional structural biology tools.