Limited Proteolysis–Mass Spectrometry (LiP-MS) Service
See protein conformational and accessibility changes that conventional abundance-focused proteomics can miss.
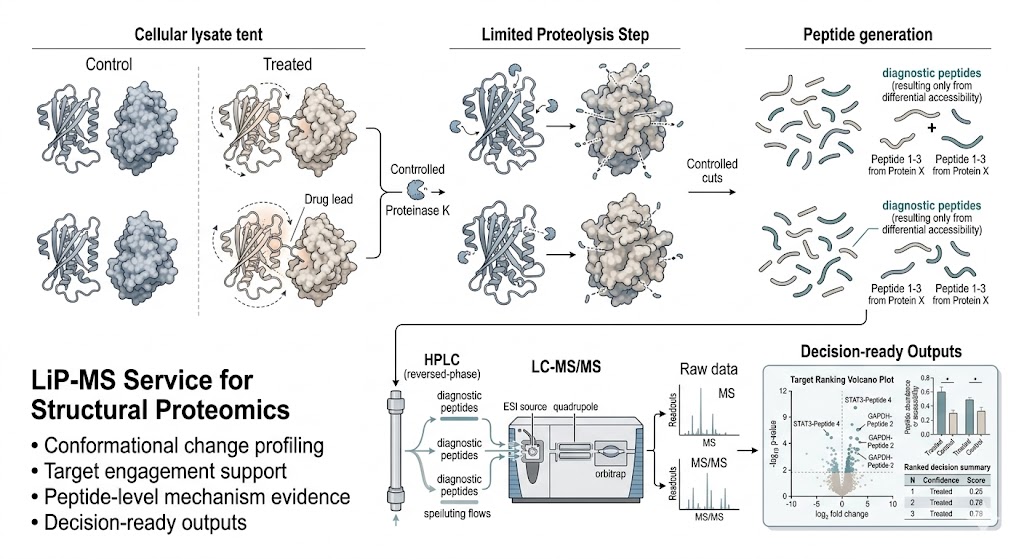
Limited Proteolysis–Mass Spectrometry (LiP-MS) Service helps you see protein structural responses that conventional abundance-focused proteomics can miss. When your project question is about conformational change, accessibility shift, target engagement support, or mechanism-related perturbation effects, LiP-MS can add peptide-level evidence that helps move from observation to interpretation.
At MassTarget™, we use LiP-MS to support discovery-stage studies where treated and control samples need to be compared in a way that connects protein structural behavior to mechanism-focused decision-making. Our goal is not just to generate differential peptide outputs, but to organize LiP-MS results into a format your team can review, challenge, and use for the next experimental decision.
Key advantages:
- Proteome-scale structural response profiling under treated-versus-control designs
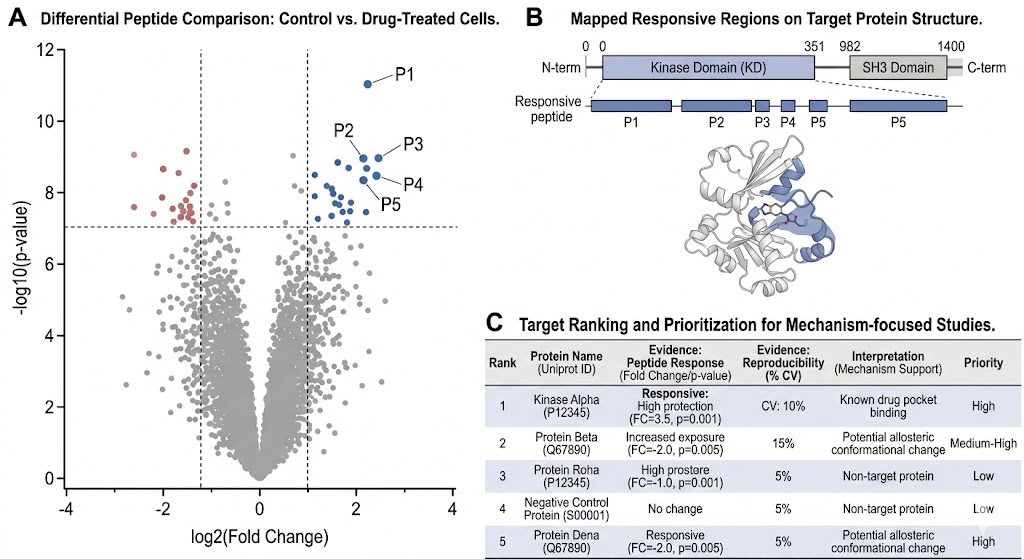
- Peptide-level evidence for target engagement and mechanism-focused interpretation
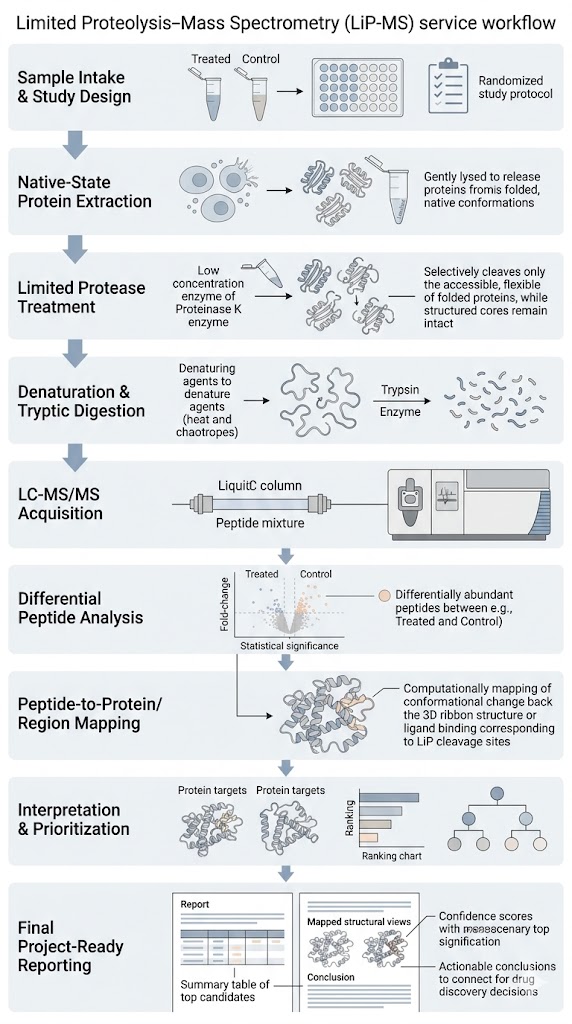
- Study-fit workflow from native-state extraction to interpretable peptide outputs
- Decision-ready deliverables beyond differential peptide tables
- Integration paths to thermal-shift and structural follow-up methods