- Service Details
- Case Study
What is Glycome and Glycomics
The glycome is defined as all polysaccharides produced by cells, whether they are bound to proteins/lipids or exist in a free form. Vertebrates synthesize glycoproteins, glycolipids, proteoglycans, glycosaminoglycans (GAGs), and glycosylphosphatidylinositol (GPI) anchors, which are covalently attached to proteins and free oligosaccharides. Similar to the proteome, each cell type possesses its unique glycome, determined by the local environment and the metabolic state of the cell. While there is no specific size established for a cell's glycome, the myriad possibilities of combinations of numerous glycan structures on various glycoconjugates imply that defining a "complete" glycome is not straightforward. Glycans can be studied as a whole (glycomics) or individually, one glycan or glycoconjugate at a time when changes occur during development, cancer progression, infections, and many other disease conditions. Many glycan-binding proteins, such as lectins, oligomerize on the cell surface and interact with arrays of glycans on the same or neighboring cells. At times, multiple glycans and their corresponding glycan-binding proteins work together to mediate cell adhesion or transmit signals between cells. Therefore, the term "glycomics" was coined to describe the many facets of glycobiology that can only be understood through systematic analysis of the glycome.
Glycomics represents a methodical exploration aimed at unraveling the intricacies of glycans and glycoconjugates' structure and function, along with the identification of the genetic codes responsible for glycoprotein synthesis within cells, tissues, organisms, or living entities. A plethora of methods and analytical tools have been innovated and put into practice to conduct comprehensive examinations of the glycome on a grand scale. These methodologies hold the promise of yielding profound revelations regarding the multifaceted roles played by glycans in the natural world and their intricate involvement in every facet of biology, well-being, and pathology.
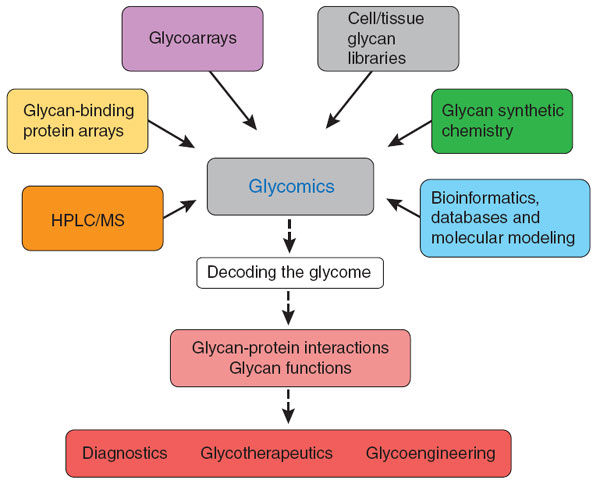
Glycomics Analysis
Glycomics analysis encompasses glycan profiling, glycan class characterization, and comprehensive structural analysis of glycans released from proteins.
Glycan Profiling involves the separation of complex mixtures of glycans to provide a simplified overview or snapshot of the glycans present in a sample. Techniques employed include High-Performance Liquid Chromatography (HPLC) for separation based on physicochemical parameters such as hydrophilicity, size, or charge, capillary electrophoresis for the separation of mass/charge-labeled glycans, and Matrix-Assisted Laser Desorption/Ionization (MALDI) and/or Electrospray Ionization (ESI) Mass Spectrometry (MS) for the separation of unlabeled or labeled glycans based on mass or charge.
Glycan Class Characterization techniques separate glycan mixtures into different glycan types based on structural features. Examples include MS separation of galactosylated and agalactosylated IgG glycans or weak anion-exchange (WAX) Liquid Chromatography (LC) separation of neutral, mono-, di-, tri-, and tetra-sialylated structures based on charge. These methods facilitate the highlighting of specific key features and provide relative quantitation of different glycan classes. LC-MS analysis can also be used to separate released N-glycan structures into mannose oligosaccharides, fucosylated oligosaccharides, sialylated and non-sialylated oligosaccharides, among others.
Comprehensive Structural Analysis involves the determination of the sequence and any modifications of monosaccharides, branch points, anomeric isomers, and glycosidic linkages within glycans present in the glycome. This detailed analysis typically requires multiple orthogonal techniques to establish preliminary structures and subsequently confirm them. If sample preparation for MS may disrupt glycans, it may be necessary to release and analyze glycans containing unstable modifications such as O-acetylation or sialylation separately. Comprehensive structural analysis may also include the absolute or relative quantitation of specific glycan structures, such as the levels of core fucose on antibodies designed for Antibody-Dependent Cellular Cytotoxicity (ADCC), levels of α-galactose residues on antigens, sialyl Lewis x epitopes as potential useful markers for inflammation and metastasis, or specific structures binding to bacterial proteins.
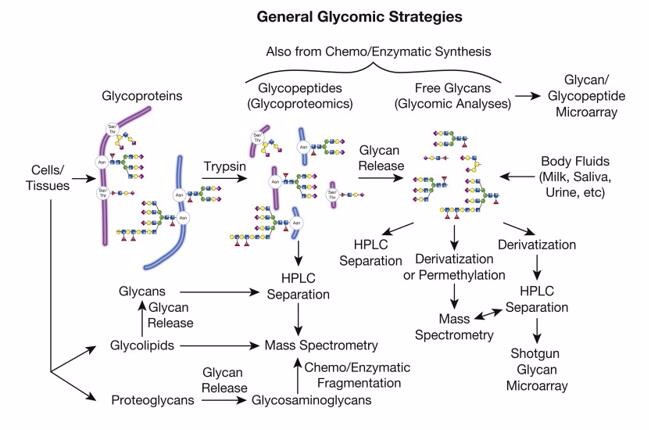
Our Glycomics Analysis Services
Creative Proteomics use advanced analytical techniques such as mass spectrometry, liquid chromatography, microarray, fluorescence and NMR spectroscopy to identify putative targets. We offer a wild range of glycomics services, including:
Mass Spectrometry-Based Glycomics Analysis Workflow
Release of Glycans from Glycoproteins:
The starting material for glycomics analysis can encompass glycoproteins embedded in gels, protein extracts from whole cell lysates, homogenized tissues, enriched membrane fractions, or serum and other bodily fluids. For high-throughput glycome analysis, complete N-glycans are typically released from glycoproteins using enzymes, such as Peptide-N-Glycosidase F (PNGase F).
Derivatization of Glycans:
Glycan derivatives refer to modifications made to polysaccharide (polymeric sugar) molecules, either chemically or biologically, to alter their chemical properties, functions, or utility. These modifications can involve different aspects of polysaccharides, including sugar units, polymer length, branching, glycan chain structures, among others. Glycan derivatives are compounds created by changing the structure or properties of polysaccharides to suit various research or application requirements.
Mass Spectrometry Analysis of Glycans:
One advantage of mass spectrometry-based glycan profiling is that different glycans can be simultaneously identified by their mass and diagnostic fragment ions, increasing the throughput of glycomics analysis. Determining the molecular weight of glycans using MALDI or ESI-MS provides a molecular distribution of glycans and allows for quantitative comparisons of glycosylation between samples. MS/MS fragmentation has become the gold standard for glycan structural characterization. The goal is to generate informative fragmentation mass spectra, which enable unambiguous glycan structure confirmation.
Post-Glycosylation Modifications:
Another challenge is that many critical glycan modifications, such as O-acetylation, acetone sialylation, and others, are unstable and/or absent in current analytical methods, potentially leading to database entries filled with misleading or biased information. For example, although many databases assume that sialic acid at the termini of vertebrate glycan chains is N-acetylneuraminic acid (Neu5Ac), in reality, there are dozens of naturally occurring sialic acid modifications, which can have profound effects on biological function. This includes variations in sialic acids due to modifications such as sulfation on N- and O-glycans on HexNAc and hexose residues. These analytical challenges continue, and, in many cases, precise localization of modifications on glycan chains remains highly challenging.
Glycoproteomics:
Most glycoproteomics methods rely on accurate mass analysis of the monoisotopic glycopeptide precursor ions (typically low ppm) to precisely determine the molecular mass of intact glycopeptides. Fragment ions also benefit from high-resolution detection for aiding in spectrum interpretation. Innovative LC-MS/MS acquisition strategies are increasingly being employed to enhance the performance of glycoproteomics experiments. Similar to peptide quantitation in proteomics, the quantitation of intact glycopeptides can be achieved using label-free or label-assisted (e.g., TMT) strategies. Quantitation may involve relative quantitation of all glycans at each glycosylation site (glycoproteome), comparison of glycan spectra at individual sites between conditions (comparative glycoproteomics), or simply determining the relative abundance of a single glycan form across two or more conditions (biomarker discovery). Extracting biologically relevant information from glycoproteomics data often requires multiple quantitation methods, along with information about protein levels and site occupancy.
Data Analysis

Creative Proteomics, staffed by highly experienced biological scientists in omics studies, can provide our customers a series of glycomics and bioinformatics analysis services. Please feel free to contact us and see how we can help you address your problems.
How to place an order

*If your organization requires signing of a confidentiality agreement, please contact us by email.
Customer Case: Identification of the O-Glycan Epitope Targeted by the Anti-Human Carcinoma Monoclonal Antibody (mAb) NEO-201
Journal: Cancers
Published: 2022
Method: Glycomics, O-glycan profile
Abstract
Glycosylation, an important post-translational modification found in mammalian proteins and lipids, can be disrupted in cancer cells. Specifically, the O-glycosylation pattern is often disrupted in both solid and liquid tumors, leading to poor prognosis and tumor progression. To counteract tumor growth, monoclonal antibodies targeting truncated O-glycans in cancer cells, such as the IgG1-humanized mAb NEO-201, have been employed as an effective strategy. Previous studies have shown that NEO-201 binds specifically to tumor-associated variants of CEACAM5 and CEACAM6 expressed in various types of cancer but not in normal tissues. In this article, we evaluated whether the epitope recognized by NEO-201 is an O-glycan, given that CEACAMs are highly glycosylated proteins. Our study demonstrated that NEO-201 binds to core 1 O-glycans and has the ability to target and eliminate cancer cells expressing both core 1 and extended core 1 O-glycans. This suggests that NEO-201 binds to O-glycans attached to any protein containing amino acid regions with serine and threonine, where a GalNAc residue can be added.
Results
1. NEO-201 Binds to Mammalian-Expressed rhCEACAM6 but Not to Bacterial-Expressed rhCEACAM6 by ELISA

2. NEO-201 Binds to O-Glycans

3. Human Cancer Cell Lines and Human Neutrophils Recognized by NEO-201 Express Core 1 and Extended Core 1 O-Glycans

4. NEO-201 Mediates ADCC to Kill Target Cells Expressing Core 1 and Extended Core 1 O-Glycans

Conclusions
This study revealed that NEO-201 has the ability to recognize various types of O-glycans, with core 1 and extended core 1 O-glycans showing the highest binding affinity. Cells that are reactive to NEO-201, such as CFPAC-1 (a pancreatic cell line), human neutrophils, HL60, and U937 (human hematological neoplastic cells), predominantly express core 1 and/or extended core 1 O-glycan profiles. NK-mediated ADCC enables NEO-201 to kill these cells. Abnormal expression of O-glycans is linked to the development, progression, invasion, and metastasis of cancer, including both epithelial-derived solid tumors and hematological malignancies like AML and multiple myeloma. By specifically binding to tumor-associated proteins carrying core 1 and/or extended core 1 O-glycans, NEO-201 could serve as a promising and innovative immunotherapy for the treatment of both solid and liquid tumors.
References
- Pauline M. Rudd, Niclas G. Karlsson, Kay-Hooi Khoo, Morten Thaysen-Andersen, Lance Wells, and Nicolle H. Packer. Chapter 51Glycomics and Glycoproteomics. In Essentials of Glycobiology [Internet]. 4th edition. Varki A, Cummings RD, Esko JD, et al., editors. Cold Spring Harbor (NY): Cold Spring Harbor Laboratory Press; 2022.
- Tsang, K.Y.; Fantini, M.; Zaki, A.; Mavroukakis, S.A.; Morelli, M.P.; Annunziata, C.M.; Arlen, P.M. Identification of the O-Glycan Epitope Targeted by the Anti-Human Carcinoma Monoclonal Antibody (mAb) NEO-201.


















