N-glycosylation site occupancy is a physiological feature of glycoproteins and mainly contributes N-glycan. The degree of N-glycosylation site occupancy by itself correlates with the severity of the disease. Mass spectrometry (MS) has been widely used to analyze site occupancy in complex or clinical samples. Based on MS, label-free and labeling methods are commonly applied in this field. Labeling methodologies include SILAC (stable isotope labeling by amino acids in cell culture), TMT (tandem mass tags), dimethylation, and iTRAQ (isobaric tags for absolute and relative quantification). Label-free methods have the advantage of experimental simplicity in sample preparation.
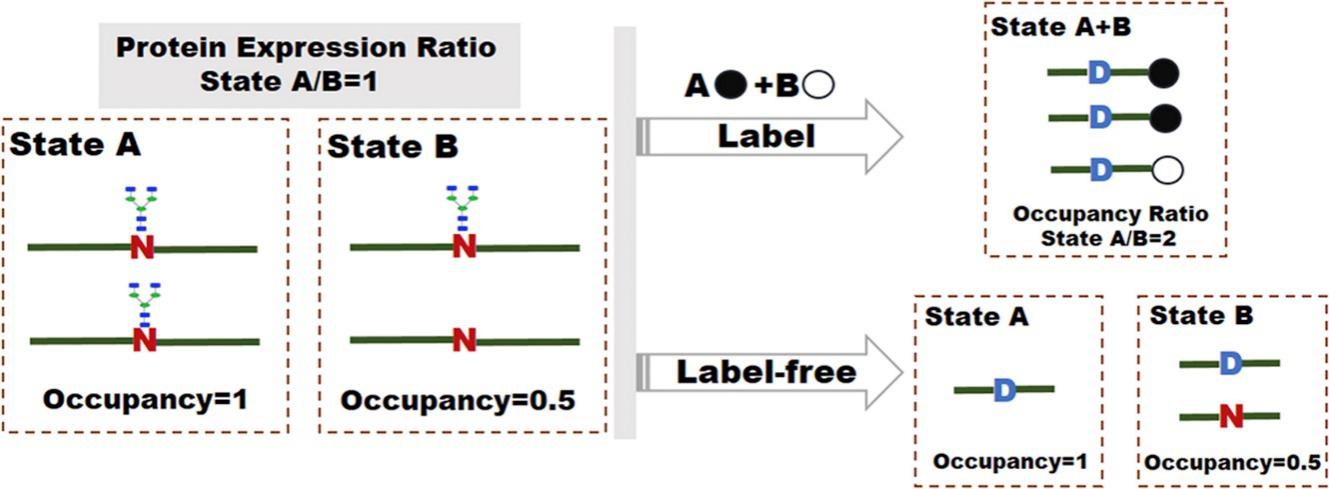 Figure 1. MS-based labeling and label-free technologies for quantification of N-glycosylation site occupancy (Zhang et al., 2017).
Figure 1. MS-based labeling and label-free technologies for quantification of N-glycosylation site occupancy (Zhang et al., 2017).
Overview of N-Glycosylation Site Occupation Analysis Service
Characterizing the site-specific N-glycosylation including N-glycosylation site occupancy and site-specific glycan structure is important for understanding glycoprotein biosynthesis and function. Quantification of N-glycosylation site occupancy is also important for disease research and drug development as N-glycosylation site occupancy could be physiological, affected by human disease or modified by heterologous protein overexpression. At Creative Proteomics, we provide both labeling and labeling-free technologies for the quantification of N-glycosylation site occupancy.
- Labeling quantification of N-glycosylation site occupancy
The significant advantage of labeling methods is the ability to simultaneously and efficiently analyze differentially labeled samples in a single run. We can provide trypsin catalyzed 18O, SILAC, and iTRAQ labeling approaches for N-glycosylation site occupancy analysis. Isotope-coded glycosylation site-specific tagging (IGOT) can be used for large-scale N-glycosylation site occupancy analysis.
- Label-free quantification of N-glycosylation site occupancy
Label-free quantification of N-glycosylation site occupancy is generally based on peptide-ion intensity or peak areas. And its significant advantage is the simple workflow. The general strategy of variable N-glycosylation site occupancy analysis in glycoproteins comprises the following procedures.
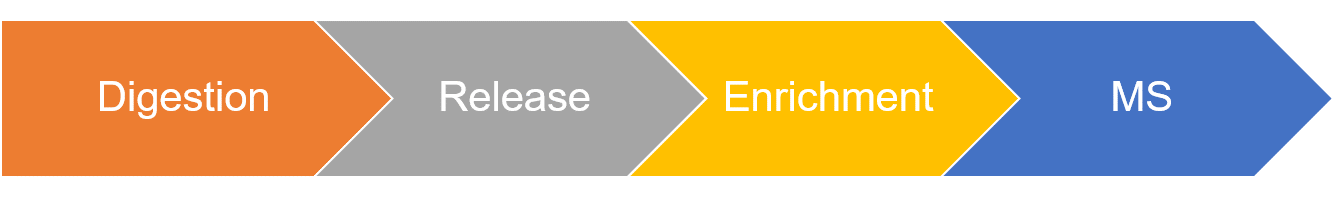 Figure 2. The workflow for label-free quantification of N-glycan site occupancy rates.
Figure 2. The workflow for label-free quantification of N-glycan site occupancy rates.
- Protein digestion of proteins to generate peptides.
- N-glycan release using PNGase F. PNGase F specifically release N-glycans linked to the nitrogen of Asn, and converts Asn residue to Asp with mass increasing of 0.984 Da.
- Enrichment of N-glycopeptide via HILIC (Hydrophilic interaction liquid chromatography).
- Determination of N-linked glycosylation site occupancy using LC-ESI-MS/MS.
Sample Requirement
We work with a range of sample sources as follows.
- Protein: 100 ug
- Cells: 1×107
- Animal tissues: 1 g
- Plant tissues: 200 mg
- Anticoagulated blood (EDTA): 1 mL
- Serum: 0.2-0.5 mL
- Urine: 2 mL
- Microbial sample: 200 mg (dry weight)
Advantages of N-Glycosylation Site Occupancy Analysis
- High coverage of N-glycans
- State-of-the-art platforms and strategies
- Customized N-glycosylation site occupancy analysis services based on your specific project
- Delivers Comprehensive report containing raw data, parameters, data interpretation, and bioinformatics analysis
Creative Proteomics provides the N-linked glycans site occupancy analysis to obtain truly glycosylated peptides in the whole mixture of a number of potentially N-glycosylated sites that are may or may not be glycosylated. Please feel free to contact us to discuss your project.
Reference
- Su Y L, Wang B, Hu M D, et al. Site-Specific N-Glycan Characterization of Grass Carp Serum IgM. Frontiers in Immunology, 2018, 9.
- Thannhauser T W, Shen M, Sherwood R, et al. A workflow for large‐scale empirical identification of cell wall N‐linked glycoproteins of tomato (Solanum lycopersicum) fruit by tandem mass spectrometry. Electrophoresis, 2013, 34(16): 2417-2431.
- Zhang S, Li W, Lu H, et al. Quantification of N-glycosylation site occupancy status based on labeling/label-free strategies with LC-MS/MS. Talanta, 2017, 170: 509-513.


















