Global Leader in Multi-Omics CRO Services: Empowering Research Discovery
Creative Proteomics delivers high-impact analytical solutions for global research teams. We combine state-of-the-art instrumentation with PhD-led expertise to provide end-to-end support across proteomics, metabolomics, lipidomics, glycomics, and spatial omics, with robust multi-omics integration and reporting. Beyond omics, we also support mass-spectrometry-enabled drug screening, drug metabolism and metabolite profiling (DMPK), drug characterization, cytokine profiling (Luminex, MSD, Simoa), and molecular interaction/biophysical binding studies (SPR, BLI, MST, ITC). As a multi-division group, we bring specialized teams for omics, bioinformatics, and biopharma analytics under one umbrella—enabling you to mix and match workflows from discovery to verification with consistent QC, clear deliverables, and scalable throughput.



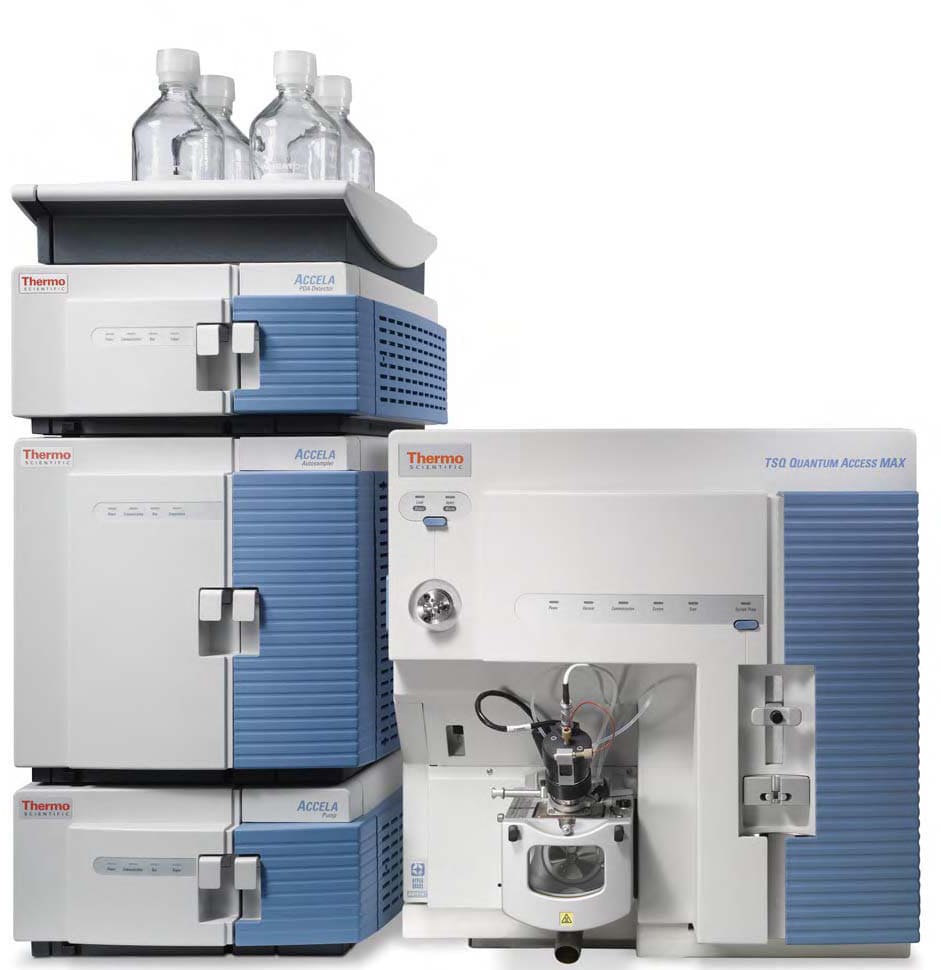
Advanced Mass Spectrometry Platform
- Orbitrap Fusion Lumos: Premier sensitivity for PTM mapping and intact protein analysis.
- Orbitrap Exploris 480: Optimized for discovery proteomics and large-scale research cohort studies.
- Bruker timsTOF Pro 2: 4D-Proteomics leveraging PASEF for superior identification depth.
- Sciex QTRAP 6500+: Industry gold standard for absolute targeted quantification.
- MS-Enabled Drug Screening: Label-free profiling, target engagement support, and structure/impurity-aware characterization (RUO).
- Drug Metabolism (DMPK): In vitro metabolism assessment, metabolite identification, and bioanalytical quantification workflows.


Biophysical & Structural Analytics
- Circular Dichroism (CD): Protein secondary structure and stability analysis.
- NMR Spectroscopy: High-resolution insight into molecular structure and purity.
- FT-IR Spectroscopy: Comprehensive functional group molecular fingerprinting.
- Nano-LC & UPLC: Precision separation via EASY-nLC 1200 and Waters ACQUITY.
- Cytokine & Biomarker Panels: Multiplex immunoassays on Luminex, MSD, and Simoa platforms (RUO).
- Molecular Interaction: Binding kinetics and affinity using SPR/BLI, plus orthogonal biophysics (MST, ITC).
Bioinformatics & Multi-Omics Data Hub
- Standardized Pipelines: Deep data mining via MaxQuant, Skyline, and Mascot clusters.
- Functional Enrichment: Pathway mapping (KEGG/Reactome) and PPI network analysis.
- Multi-omics Integration: Systematic correlation analysis of proteome-metabolome datasets.
- HPC Infrastructure: Dedicated server clusters ensuring rapid data turnaround times.
- Multi-omics Joint Analysis: Cross-omics correlation, causal pathway inference, and multi-layer reporting.
- Spatial Omics Analytics: Spatial feature extraction, region-specific differential analysis, and cell neighborhood mapping.
- Targeted & Panel Data: Olink/immunoassay-style panel processing, QC, normalization, and downstream interpretation.
- Sequencing-Compatible Workflows: Illumina-ready library/data handoff and integrated analysis with omics outputs.
- Interaction & Network Biology: Protein–protein interaction analysis and system-level network modeling.

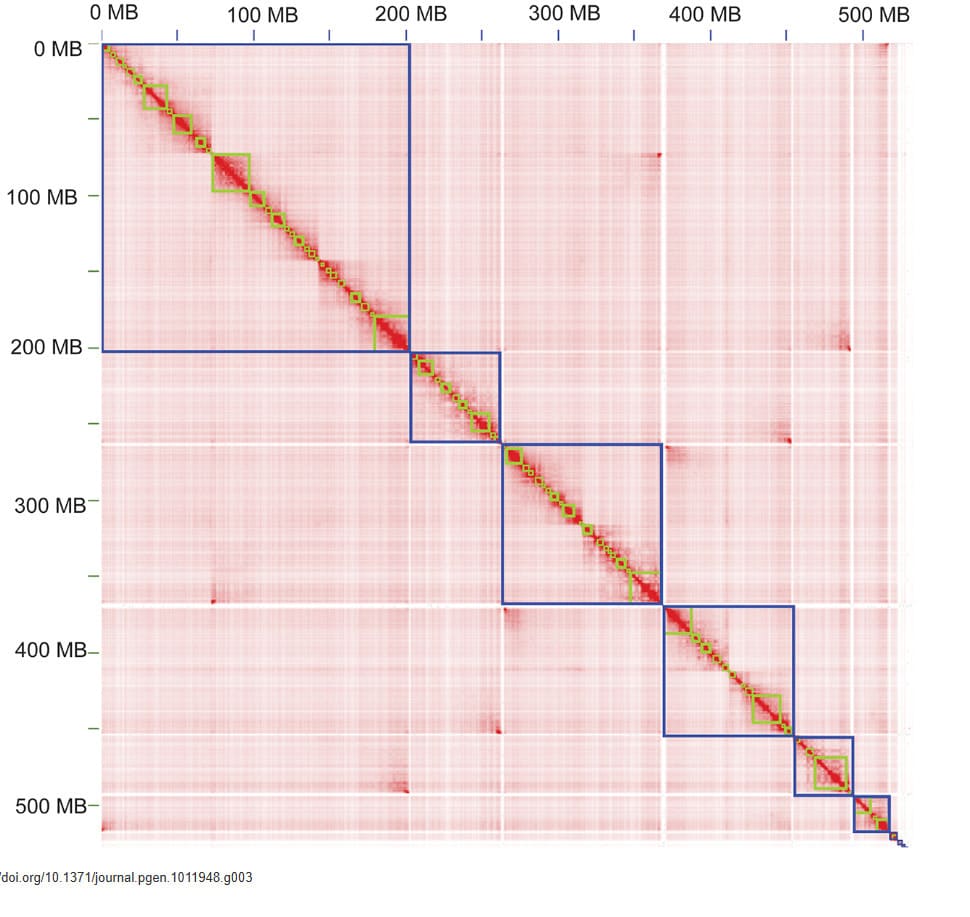
Transcriptomic profiling identifies molecular markers for insect developmental timing; Hi-C–enabled assembly/QC visuals support robust genomics analysis in research workflows.
Transcriptomics | Hi-C / 3D Genomics | Genome Assembly QC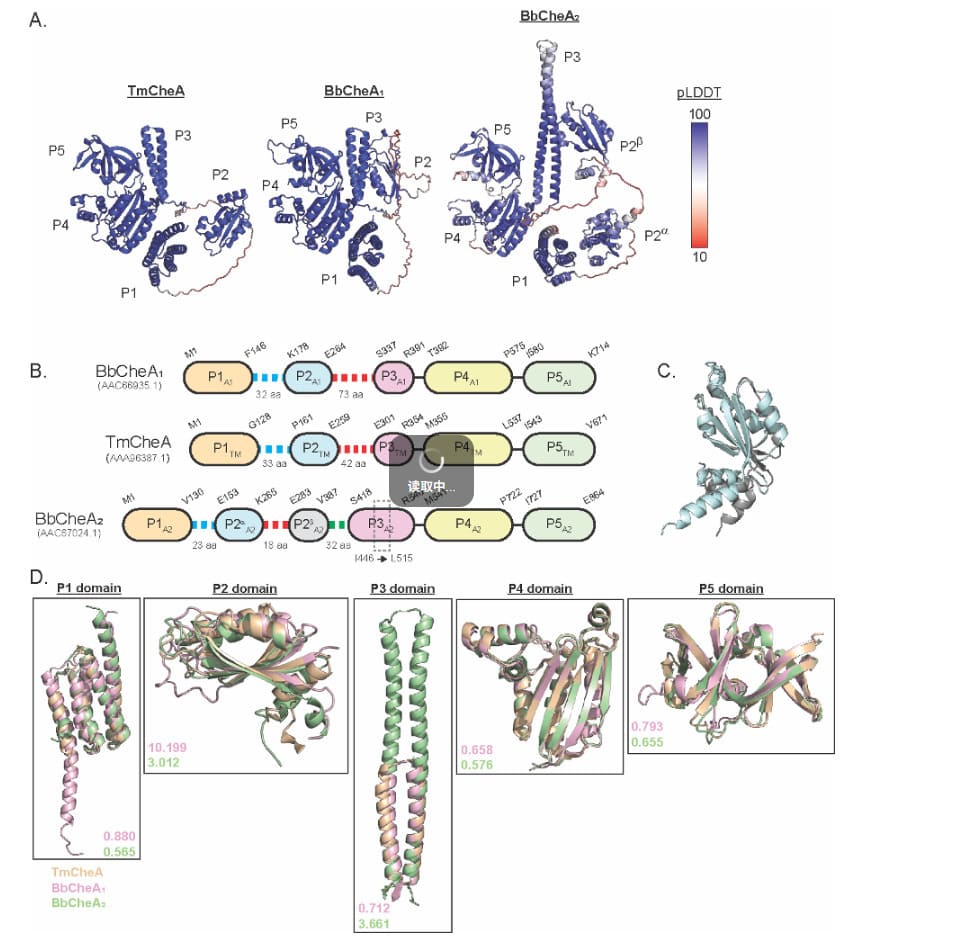
Mechanism-oriented analysis links signaling components to phenotype; structure/domain comparisons provide clear, publication-grade visuals for research interpretation.
Protein Modeling | Mechanism Mapping | Functional Genomics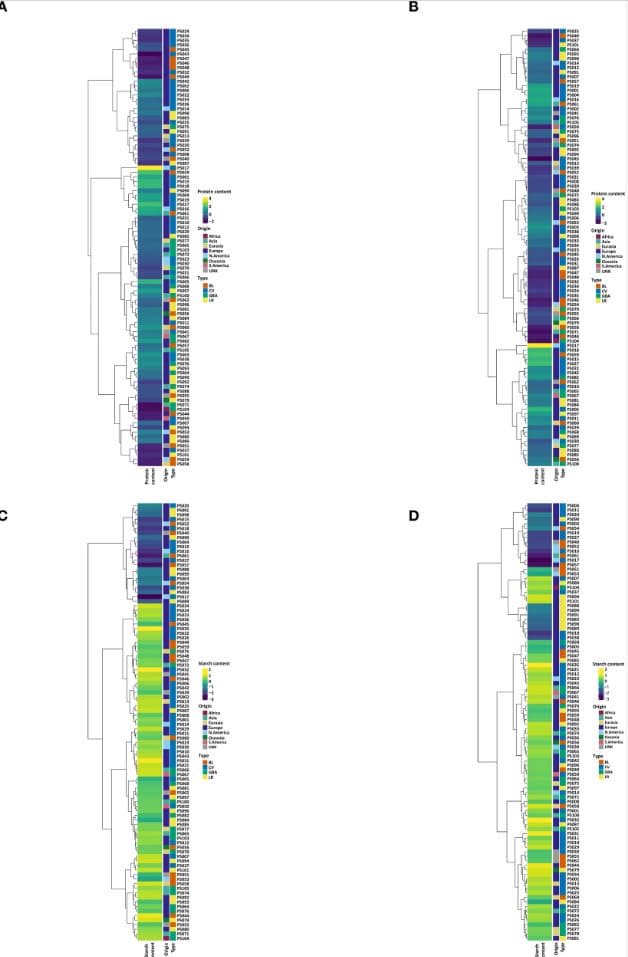
SNP-driven population structure and clustering align genetic variants with measured seed composition traits for research and breeding workflows.
Genotyping / SNP Analysis | PCA | Trait Association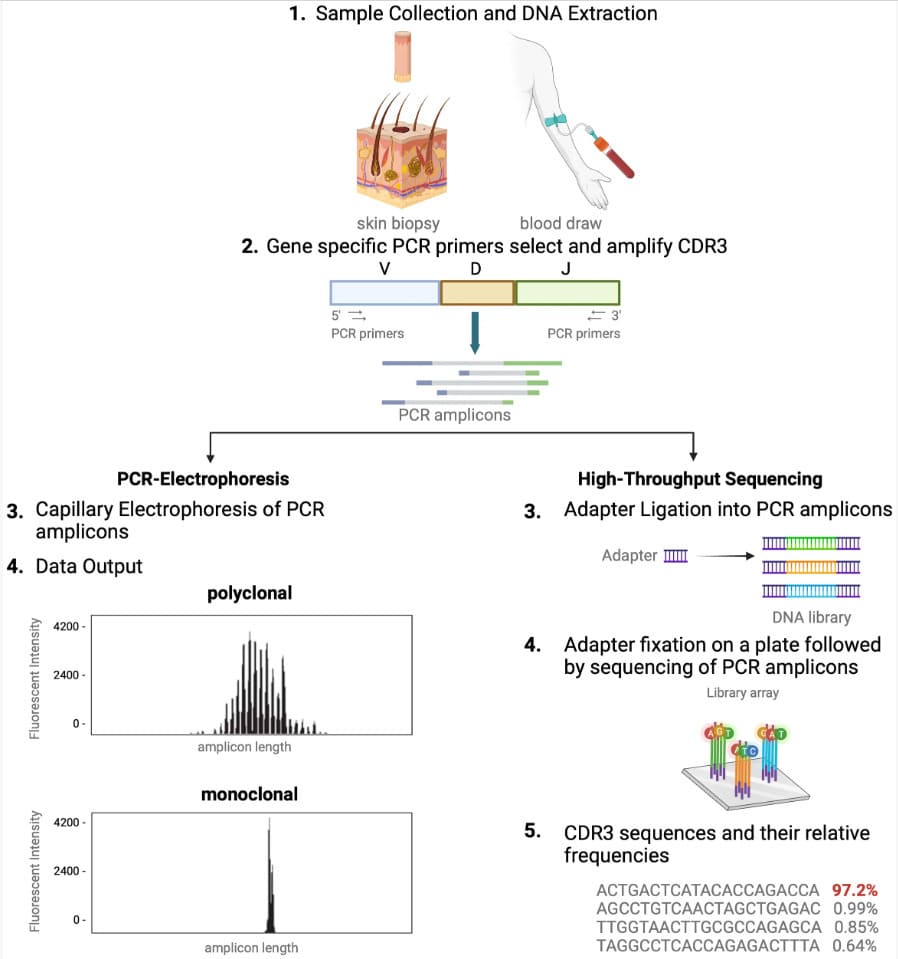
A side-by-side workflow schematic clarifies sample-to-readout steps and highlights why sequencing-based clonotype profiling improves resolution for immune repertoire research.
Immune Repertoire Sequencing | NGS Workflow | Clonotype Profiling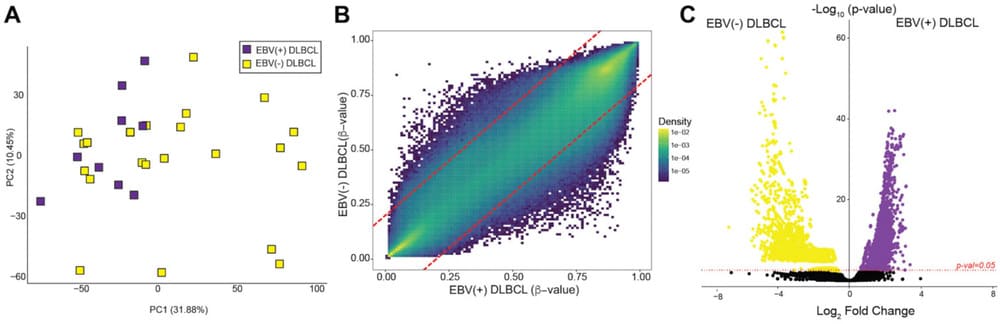
Methylome-level ordination (PCA) separates groups and illustrates how epigenomic profiles stratify research cohorts for downstream differential analysis.
DNA Methylation | Epigenomics | PCA Stratification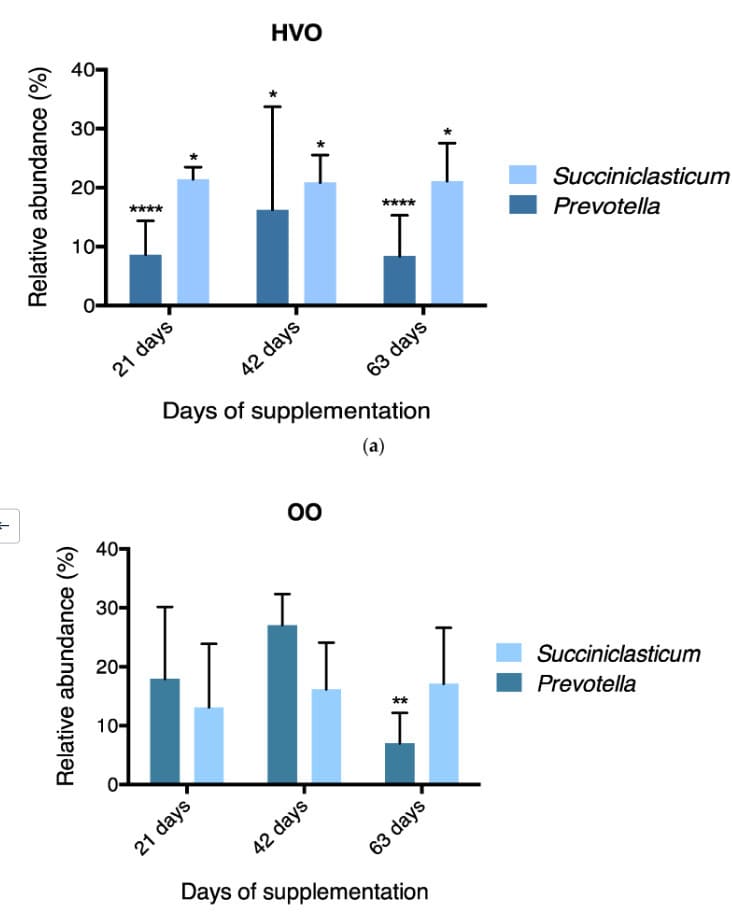
Longitudinal community profiling shows how dietary lipid sources shift dominant taxa over time, providing a clear summary for microbiome research designs.
16S Microbiome | Longitudinal Design | Differential Abundance"We combined proteomics + metabolomics with their joint analysis workflow and got a coherent mechanism story in one report."
"Their Olink panel planning, sample QC, and downstream statistics were tightly executed. Professional B2B service."
"We needed spatial omics plus follow-up protein validation. The team handled the data integration end-to-end."
"For drug characterization, they supported binding/interaction readouts which helped us prioritize candidates quickly."
"The cytokine profiling service was rock-solid—clean QC, low batch effects, and clear interpretation across conditions."
"Their lipidomics coverage and annotation depth exceeded what we could do in-house, and the turnaround stayed predictable."
"We ran Illumina-based assays alongside proteomics and they standardized the reporting so everything lined up."
"Protein–protein interaction experiments were publication-ready, with transparent controls and raw data packages."