What is Label Transfer?
It is one of the techniques used to study protein-protein interactions (PPIs) in biological systems, involving the transfer of a label from one protein to another upon interaction. The technique is frequently employed to investigate weak or transient interactions that are often imperceptible using other methods, such as Co-IP or pull down. In a typical assay, the bait protein (interested protein) is first reacted with a label transfer reagent (LTR), such as fluorogenic, radioisotope, biotin, or chromogenic label. The labeled bait protein then incubates with prey protein (target protein) in vitro, and then they are exposed to ultraviolet light to activate the photosensitive group part on the cross-linker. Secondly, another terminal of the LTR covalently binds to the prey protein. Finally, label transfer is accomplished by cleaving the cross-linker spacer arm to release the attached bait protein. Overall, the technology plays a crucial role in the study of PPIs, holding immense potential in providing valuable insights into the underlying molecular mechanisms. This, in turn, facilitates a deeper comprehension of cellular processes, propels biomedical research, and potentially contributes to advancements in therapeutic interventions.
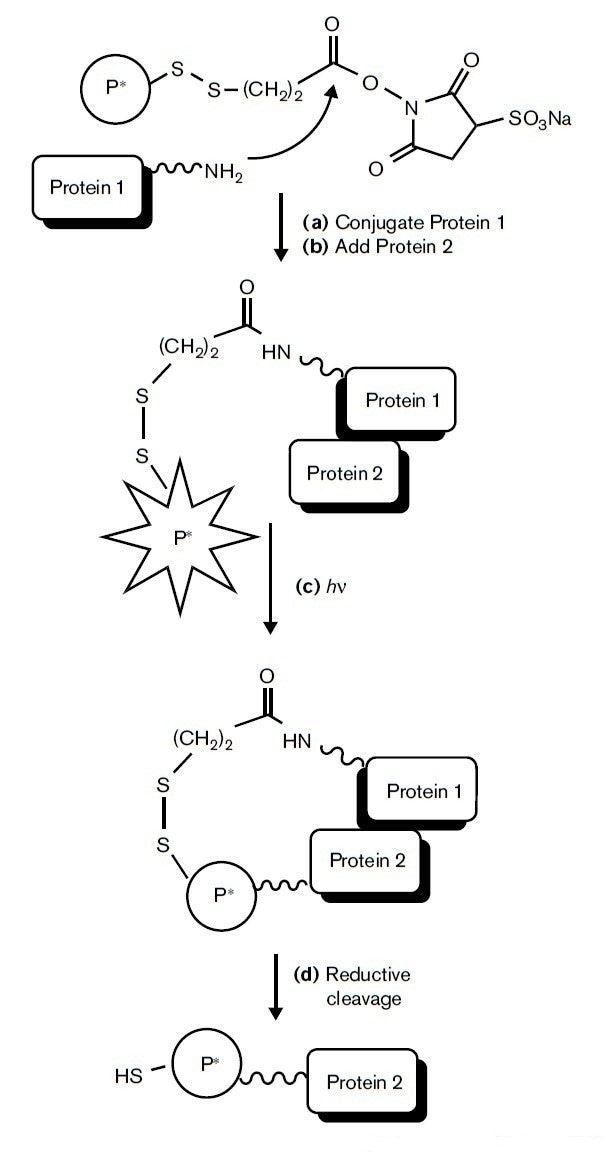 Figure 1. The principle of identification of target proteins by label transfer [1].
Figure 1. The principle of identification of target proteins by label transfer [1].
The analysis of complete PPIs analysis follows a specific process (figure 1): (i) a target protein (Protein 1) is first derivatized on one end of a hetero-bifunctional cross-linking reagent via a nucleophilic attack, for example, on a succinimide. (ii) The protein derivative is then added back to its original multiprotein complex and (iii) photolyzed (hν) causing a radio-labeled photo-probe (P*) to cross-link with a nearest neighbor protein (Protein 2). (iv) Specifically purify and isolate the target PPIs complexes. (v) The disulfide bond between the interacting proteins can be reduced by a reducing agent, transferring the radio-label to Protein 2 and completing the transfer process. Therefore, the label transfer strategy is highly sensitive, making it an excellent approach for studying weak or transient PPIs.
Label Transfer Protein Interaction Analysis in Creative Proteomics
With over a decade of experience in the field of proteomics and PPIs research, Creative Proteomics possesses exceptional expertise and have a well-established repertoire of technologies for Label Transfer Protein Interaction Analysis. We are proficient in various LTRs techniques, including Sulfo-SBED. The combination of label transfer with western blot and/or LC-MS/MS techniques facilitates the accurate mapping of interacting sites and PPI networks. Moreover, the incorporation of labels enables efficient purification of both decoy and target proteins. Importantly, we possess the capability to handle various sample types and provide accurate and comprehensive identification as well as quantification information pertaining to PPIs and their respective binding sites.
 Figure 2. Identification of PPIs through combination of label transfer and LC-MS/MS [2].
Figure 2. Identification of PPIs through combination of label transfer and LC-MS/MS [2].
Advantages
- Reliable in study weak or transient PPIs;
- Suitable for both in vivo and in vitro applications;
- The strategy can significantly enhance confidence in identifying PPIs and enable the simultaneous mapping of the binding interface under physiological conditions.
Creative Proteomics also provides the following PPIs analysis services
- Co-immunoprecipitation/mass spectrometry (Co-IP/MS)
- Crosslinking Protein Interaction Analysis
- Far-Western Blot Analysis
- Pull-Down Assay
- BioID-MS Service
- Chemical Cross-linking Mass Spectrometry (CX-MS) Service
- Tandem Affinity Purification (TAP)-MS Service
- TurboID Service
Technologies platform
1. Skilled methodologies in studying PPIs: The incidence of false-positive identifications is minimal.
2. Professional detection and analysis capability: The research team is equipped with specialized expertise in proteomics, quantitative proteomics, and bioinformatics data preprocessing and analysis.
3. High specificity: Optimization of experimental design and methods with strict quality control.
4. High stability and reproducible: Achieve consistent and reproducible results in both inter- and intra-assay analyses for data interpretation.
5. High resolution and sensitivity: Triple TOF 5600, Q-Exactive, Q-Exactive HF, Orbitrap Fusion™ Tribrid™, etc.
Results Delivery
1. Detailed report, including experiment procedures, parameters, etc.
2. MS raw data and and data analysis results (e.g. identified proteins and peptides, site mapping, and optional bioinformatics analysis results, such as volcano plot, PCA, GO, KEGG, Protein Interaction Networks, etc.).
How to place an order
The provision of comprehensive support tailored to your specific requirements for Label Transfer Protein Interaction Analysis is our area of expertise. Please do not hesitate to contact us via email for any discussions regarding your specific requirements. The project we have will be efficiently managed to ensure cost-effectiveness and timely completion. Our customer service representatives are available 24 hours a day, from Monday to Sunday.

References
- Fancy DA. Elucidation of protein-protein interactions using chemical cross-linking or label transfer techniques. Current Opinion in Chemical Biology. 2000 Feb;4(1):28-33.
- Yang Y, Song H, He D, et al. Genetically encoded protein photocrosslinker with a transferable mass spectrometry-identifiable label. Nature Communications. 2016 Jul 27;7:12299.














