Bioinformatics plays an important role in all aspects of protein analysis, including sequence analysis, structure analysis, and evolution analysis. In sequence analysis, several bioinformatics techniques can be used to provide the sequence comparisons, in which new sequences can be compared to those with known functions to study the biology of an organism. In structure analysis, many available databases and bioinformatics techniques can help to find the factors governing the folding and stability of proteins and to predict the secondary structures from amino acid sequence. In evolution analysis, we use the technique like phylogenetic trees to find homologous proteins and conservative functional regions. With bioinformatics techniques and databases, function, structure and evolutionary history of proteins can be easily identified.
Bioinformatics for Protein at Creative Proteomics
Bioinformatics has been applied to protein research for many years and endeavored great contributions in sequence, structure and evolution analysis of proteins. At Creative Proteomics, we can provide various professional bioinformatics techniques for protein analysis.
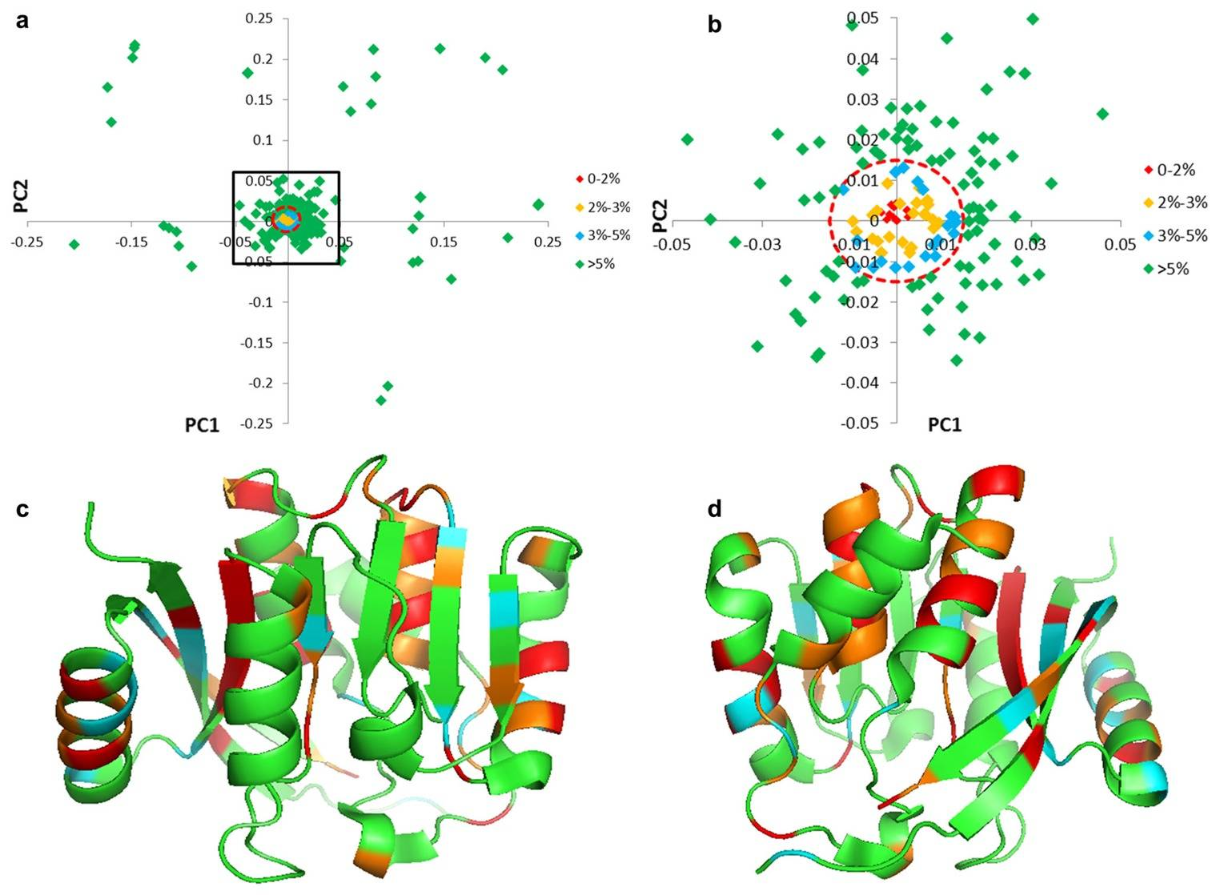 Clustering of amino acids
Clustering of amino acids
Protein Sequencing Analysis Services
The analysis of protein sequences provides the information about the preference of amino acid residues and their distribution along the sequences for understanding the secondary and tertiary structures of proteins and their functions. Besides, the identification of conserved regions in protein sequences support to predict the structurally or functionally important regions. We provide several tools for protein sequence analysis, such as PSI-BLAST (Position-Specific Iterative BLAST), a more suitable method to search a database of sequences similar to a query sequence, and ClustalW, which is a multiple sequence alignment program for biomolecules.
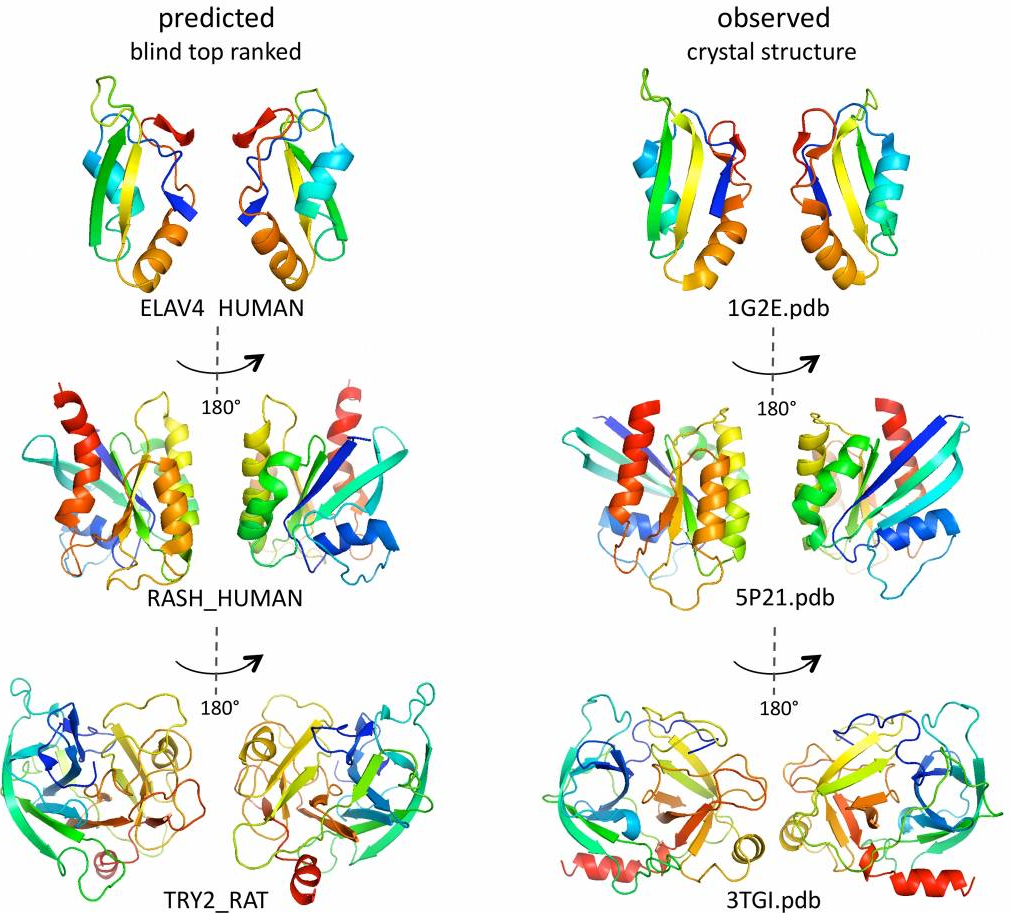 Predicted 3D structures for three representative proteins
Predicted 3D structures for three representative proteins
Protein Structure Analysis Services
The analysis on protein structures provides data about the factors associated with the folding and stability of proteins, the interactions between amino acid residues and with the surrounding medium, the preferred amino acid residues in protein environment, and the location of residues in the interior/surface of a protein and amino acid clusters. The information obtained from the protein structure analysis will enhance the prediction the secondary and tertiary structures of proteins. In addition, secondary structure information enables to predict three-dimensional structures of proteins and protein stability upon mutations. We use ACCESS to compute ASA (The solvent accessible surface area) and Naccess to calculate the atomic accessible area.
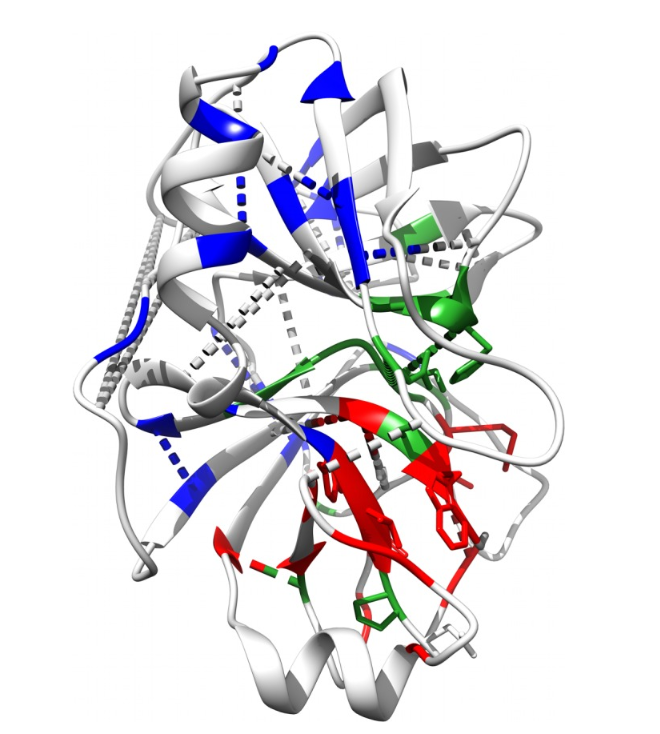 Co-evolving residues in rat trypsin
Co-evolving residues in rat trypsin
Protein Evolution Analysis Services
The analysis of protein evolution engaged the study on homologous protein, functionally conserved regions, and the evolutionary history of protein. We use the phylogenetic tree for the protein evolution analysis. Phylogenetic tree, which consists of a root (the last common ancestor of the protein family), nodes, branches, and tips, can use the ancestral protein sequence reconstruction to infer the history of a protein family and detect the coevolution between protein families. The phylogenetic tree can be constructed and analyzed with robust scripting tools, which include R package “APE”(Analyses of Phylogenetics and Evolution), ETE (a Python Environment for phylogenetic Tree Exploration) and Bio.Phylo for Biopython. We will choose the best tool for specific requirements.
Our Advantages
- A wide range of bioinformatics techniques for all aspects of protein analysis, including sequence, structure and evolution analysis of protein.
- Various techniques for each aspect of protein analysis, and we will choose the suitable method to meet your needs.
- Analysis visualization to help you get visible results suitable for research and display.
- Fast and cheap bioinformatics protein analysis services in a time-saving and cost-efficient manner.
Creative Proteomics is a leading biotech company, specialized in offering a set of services for protein analysis. We provide a wide range of bioinformatics techniques for all aspects of proteins. Our experienced scientists and advanced techniques enable us to deliver and high-quality data with cost-efficiency. If you have any questions or specific requirements, please do not hesitate to contact us.
References
- Bo Wang.; et al. Principal components analysis of protein sequence clusters. Journal of structural and functional genomics.2014; 15(1):1-11.
- Hensen U.; et al. Exploring protein dynamics space: the dynasome as the missing link between protein structure and function. PLoS One. 2012; 7(5): e33931.
- Sikosek T.; et al. Biophysics of protein evolution and evolutionary protein biophysics. Journal of the Royal Society Interface. 2014; 11(100): 20140419.
- Marks DS.; et al. Protein 3D structure computed from evolutionary sequence variation. PLoS One. 2011; 6(12): e28766.




