Lipidomics Service: Cutting-Edge Solutions for Exploring Lipid Pathways & Networks
Lipidomics services allow researchers to analyze the full lipid profile of biological samples. These services are critical for understanding lipid metabolism, uncovering disease mechanisms, and identifying novel biomarkers.
At Creative Proteomics, we offer comprehensive lipidomics services, including Targeted Lipidomics, Untargeted Lipidomics, and specialized services for plants, yeast, mammals, and even exosomes.
Submit Your Request Now
×- Intruduction
- Types of Services
- Workflow
- Platform
- Feature
- Application
- Sample Requirement
- Case Study
- FAQ
- Publication
Understanding Lipids: More Than Just Cell Membranes
In recent years, scientists have discovered that lipids do far more than build cell membranes or store energy. These complex molecules also play key roles in essential cellular functions—from regulating inflammation to transmitting signals between cells.
Disruptions in lipid metabolism have now been linked to several major health issues, including:
Alzheimer's disease and other neurodegenerative conditions
- Type 2 diabetes and obesity
- Cancer
- Certain infectious diseases
What's driving this research forward is lipidomics—the science of identifying and studying lipids at scale.
What Are Lipids?
Lipids are a diverse group of fat-like molecules found in every living organism. They're usually grouped into eight major categories, each with specific biological roles:
Fatty acids (e.g. linoleic acid, arachidonic acid) – Key components of metabolic energy and signaling.
Glycerolipids (e.g. triglycerides, diglycerides) – Energy storage molecules.
Glycerophospholipids (e.g. PC, PE, PG, PA) – Major building blocks of cell membranes.
Sphingolipids (e.g. ceramides, sphingomyelins) – Important in signal transmission and cell recognition.
Sterolipids (e.g. cholesterol) – Regulate membrane fluidity and serve as hormone precursors.
Glycolipids (e.g. MGDG, SQDG) – Involved in cell signaling and immune responses.
Prenol lipids (e.g. Coenzyme A derivatives) – Crucial for metabolic pathways.
Polyethylene-linked lipids – A newer class with emerging biological significance.
Each lipid class contributes to a different aspect of health—and when these molecules fall out of balance, disease often follows.
What Is Lipidomics?
Lipidomics is the large-scale study of pathways and networks of cellular lipids in biological systems. It involves identifying and quantifying thousands of lipid species, helping researchers understand their roles in health and disease. Compared to metabolomics and proteomics, lipidomics focuses solely on lipids—key molecules in cell structure, energy storage, and signaling.
What Is Lipidomic Analysis?
Lipidomic analysis is the high-throughput study of lipids in biological samples—like tissues, cells, or blood plasma. It goes beyond simple identification to explore how lipids behave, how they're structured, and what roles they play in health and disease.
Here's what lipidomics can reveal:
Lipid structures: Detailed information on chain length, saturation, and functional groups
Lipid interactions: How lipids interact with proteins, metabolites, and other lipids
Metabolic networks: How lipids are synthesized, broken down, or altered in disease
Functional insights: The biological roles of specific lipids under different physiological conditions
Rather than isolating a few key molecules, lipidomics looks at the big picture—offering a panoramic view of lipid metabolism and its influence on human health.
Importance of Lipidomics in Modern Biomedical Research
Lipids play central roles in many diseases, including cancer, diabetes, and neurodegenerative disorders. Lipidomics helps:
- Detect early disease biomarkers
- Monitor metabolic disorders
- Understand lipid-related drug responses
- Support precision medicine initiatives
By profiling lipid changes, scientists can gain deeper insights into disease mechanisms and develop more effective treatments.
Types of Our Lipidomics Analysis
Targeted Lipidomics
- Targeted Lipidomics
- Untargeted lipidomics
- MALDI-Imaging Lipidomics
- Exosome Lipidomics
Targeted Lipidomics Service
Key Lipid Classes Analyzed
Lipidomics services typically analyze a wide array of biologically significant lipid classes. At Creative Proteomics, our targeted lipidomics platform covers:
Fatty Acids & Their Derivatives
From essential omega-3s to inflammatory mediators, fatty acids play vital roles in cardiovascular health, metabolism, and immune function.
Services include:
- Short Chain Fatty Acids, Medium- and Long-chain Fatty Acids, and Very Long Chain Fatty Acids
- Saturated Fatty Acids vs. Unsaturated Fatty Acids
- Free Fatty Acids (FFAs)
- Branched & Straight Chain Fatty Acids
- Total TRANS-fatty Acid quantification
- Linoleic Acid & α-Linolenic Acid analysis
- Omega-3 and Omega-6 Fatty Acids profiling
Metabolic derivatives:
Eicosanoids (e.g., Prostaglandins, Leukotrienes, HETEs)
CYP450 Metabolites
For pathway-level insights, we also offer:
Sphingolipid Profiling
Sphingolipids are crucial for neurobiology, immune signaling, and apoptosis regulation, with emerging roles in neurodegenerative and autoimmune diseases.
Our Sphingolipid Analysis Services cover:
- Ceramides
- Sphingomyelin
- Sphingosine Bases
- Sphingosine-1-phosphate (S1P)
- Gangliosides
- Hexosylceramide
Sterol Lipids
Sterol lipids regulate membrane integrity and serve as precursors to steroid hormones and vitamin D.
Explore our Sterol Lipids Analysis Service, including:
- Total Cholesterol
- Free Cholesterol
- Cholesterol Esters
- Oxysterols
- Steroids
Glycerolipids & Glycerophospholipids
These lipid classes are foundational to membrane dynamics, energy storage, and signaling cascades.
- Glycerolipids Analysis Services include:
- Triglycerides (TGs)
- Diacylglycerols (DAGs)
- Monoacylglycerols (MAGs)
- Glycerophospholipids Analysis Services include:
Major headgroups: PC, PE, PG, PA, and more
- Cardiolipins—key to mitochondrial function
Other Specialized Lipid Classes
In addition to major lipid categories, we also analyze rare and bioactive lipid species via our Other Analysis Services:
- Lipid CLASS Profiles – Overview of lipid diversity
- Phospholipids & wax esters
- Hydroxycholesterols – Oxidized cholesterol forms
- Polyprenols – Important for protein glycosylation
- Olive Oil Phenols – Nutraceutical lipid profiling
Untargeted Lipidomics
Untargeted Lipidomics detects and quantifies a broad range of lipids without prior knowledge. It supports:
- Biomarker discovery
- Hypothesis generation
- Exploratory research
Check out our Untargeted Lipidomics, including options for plants, yeast, and mammals.
Advanced Techniques: MALDI-Imaging Lipidomics
MALDI-Imaging Lipidomics combines spatial imaging with lipid detection. It allows researchers to map lipid distributions across tissue sections—providing valuable context for disease progression, tumour microenvironments, or organ-specific metabolism.
For example, in liver disease research, MALDI imaging has revealed distinct zonal lipid distributions linked to fibrosis development.
Emerging Frontier: Exosome Lipidomics
Exosomes are nano-sized vesicles that carry lipids, proteins, and RNA between cells. Analysing their lipid content can uncover:
- Mechanisms of intercellular communication
- Biomarkers for cancer and neurodegenerative diseases
- Clues to drug resistance pathways
At Creative Proteomics, our Exosome Lipidomics Service isolates exosomal lipids and characterizes them using high-resolution mass spectrometry—offering a new layer of insight in biomarker discovery and therapeutic monitoring.
Workflow of a Lipidomics Service
Sample Preparation and Submission
- Accepted samples:
- Blood/plasma/serum
- Tissue homogenates
- Cell cultures
- Exosomes
Lipid Extraction and Quantification
- Solvent-based extraction (Bligh-Dyer, Folch methods)
- Analysis via LC-MS, UPLC-MS, or MALDI-Imaging Lipidomics
Data Analysis and Reporting
- Quantification of lipid species and classes
- Multivariate statistics (PCA, PLS-DA)
- Volcano plots, heatmaps
- Interactive visualizations
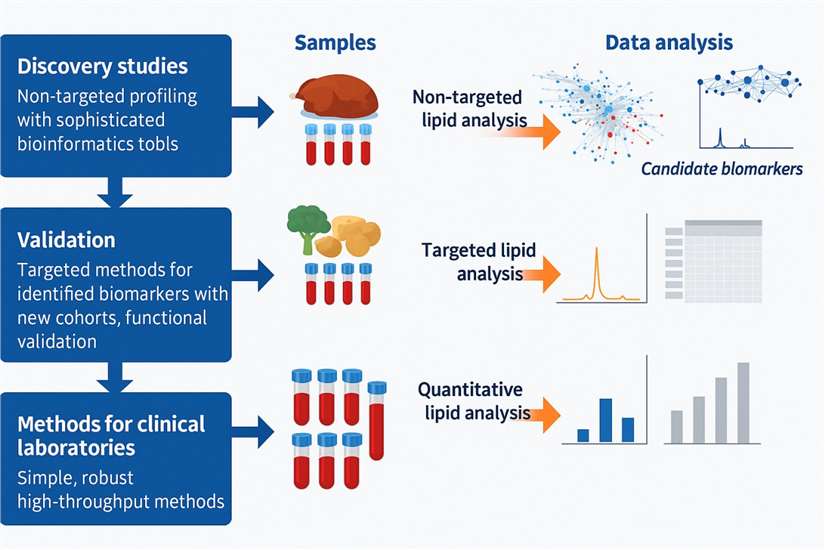 Figure 1. Analytical Workflows in Lipidomics
Figure 1. Analytical Workflows in Lipidomics
Lipidomics Data Analysis Workflow
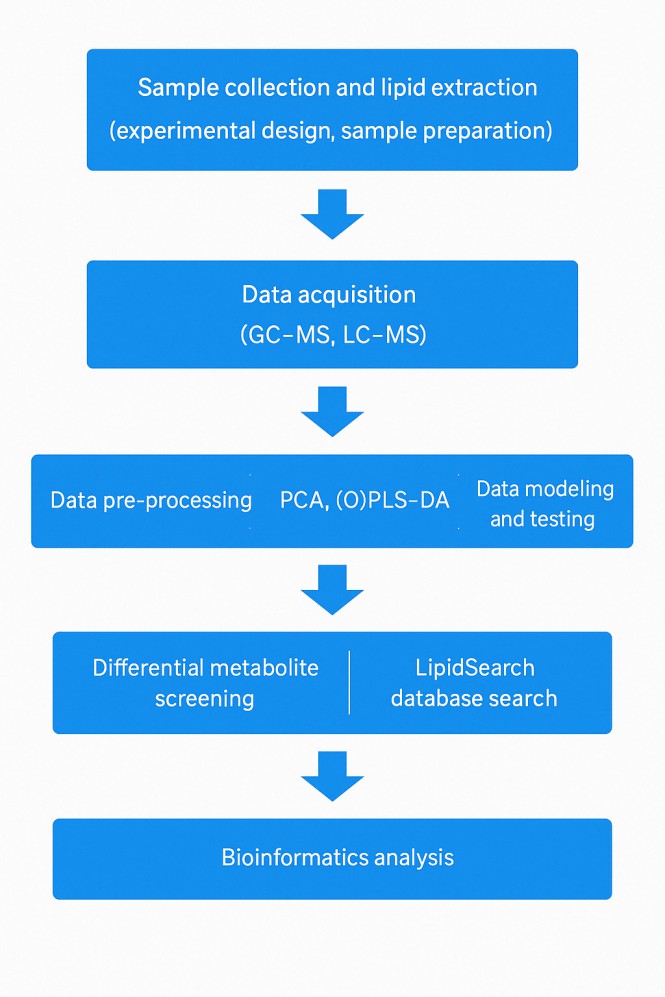
Lipidomics Technology Platform: Revolutionizing Lipid Research
Lipidomics, the study of lipid profiles within biological systems, involves several advanced techniques, including lipid extraction, separation, analysis, and identification. At Creative Proteomics, we leverage cutting-edge biological mass spectrometry technology to offer comprehensive lipidomic analysis services. Our approach integrates two primary strategies: LC-MS and shotgun lipidomics.
LC-MS
In our LC-MS analysis, we employ various lipid extraction methods tailored to different lipid classes. We categorise lipids based on their polarity using normal phase chromatography, sorting them into distinct species. Once separated, reverse-phase chromatography is used to further isolate lipid molecules, allowing for in-depth qualitative and quantitative analysis through mass spectrometry.
Shotgun Lipidomics
Shotgun lipidomics takes a more direct approach. In this method, lipid extracts are analyzed without prior chromatographic separation, providing rapid and high-throughput analysis of lipid profiles.
- State-of-the-Art Mass Spectrometry Platforms We utilize industry-leading platforms, including:
- AB Triple TOF 6600 (coupled with Agilent 1290 UHPLC)
- Scientific Orbitrap Fusion Lumos Tribrid Triple UHPLC-Mass Spectrometer System
- Orbitrap Mass Spectrometer
These systems deliver precise and reliable lipidomic data, enabling comprehensive insights into lipid composition and their biological implications.
Features of Our High-Quality Lipidomics Service
In lipidomics research, the accuracy and depth of your data can make or break a study. That's why choosing the right service provider is essential. At Creative Proteomics, our lipidomics platform is built to support researchers across pharma, biotech, and academia with high-resolution, high-confidence lipid data. Here's what makes our offering stand out:
1. Cutting-Edge Instrumentation
We deploy advanced LC-MS and GC-MS platforms that combine:
- Ultra-high resolution for precise lipid profiling
- Fast scanning speeds and high-throughput capacity
- Exceptional sensitivity to detect low-abundance lipids
These technologies ensure you get comprehensive lipid coverage, no matter how complex your sample is.
2. Expert Bioinformatics for Actionable Insights
Raw data alone won't drive decisions. Our in-house bioinformatics team provides:
- Tailored data analysis aligned with your study goals
- Interactive reports with pathway mapping and statistical interpretation
- Ongoing support for troubleshooting and custom data mining
This ensures your lipidomics data translates into meaningful biological insights.
3. Rigid Quality Control from Start to Finish
We maintain stringent quality control protocols at every stage:
- Standardised sample handling to minimise variability
- Routine instrument calibration and performance checks
- Internal QC samples run in parallel for benchmarking
These practices ensure your results are both reproducible and publication-ready.
4. Versatile Sample Handling
We've successfully processed an extensive range of biological materials, including:
- Human samples: plasma, serum, urine, feces, saliva
- Animal models: tissues, culture fluids
- Microbial and plant extracts
Whatever your sample type, we have the protocols to preserve lipid integrity and maximise recovery.
5. In-House Lipidomics Database
Creative Proteomics offers one of the industry's most robust lipid databases:
- 34 lipid categories
- 70,000+ lipid species
- Over 100,000 curated MS/MS spectra
This proprietary resource enables precise identification and quantification across thousands of lipid molecules.
6. Integrated Multi-Omics Capability
We support multi-omics integration, combining lipid data with transcriptomics, proteomics, and metabolomics to reveal systems-level insights. This holistic approach helps uncover novel biomarkers and regulatory networks in complex biological systems.
7. All-in-One Workflow
Our platform covers every step of the lipidomics pipeline:
- Sample prep → Metabolite ID → Quantification → Pathway analysis
- Fully customisable depending on your research question
- One-stop service for efficiency and consistency
With Creative Proteomics, you get more than just data—you get clarity.
Applications of Lipidomics Analysis
Drug Discovery and Development: Lipidomics plays a crucial role in drug development by tracking how drugs affect lipid metabolism. It helps identify lipid-related drug targets, aiding in the design of more effective and targeted therapies.
Lipid Function and Metabolic Regulation: Lipidomics enables detailed research on the roles of lipids in cellular functions and metabolic regulation, enhancing our understanding of how lipids influence disease and health.
Lipid Metabolic Pathways and Networks: By studying lipid metabolic pathways and their interactions, lipidomics provides deeper insights into the biochemical networks that regulate cell function and disease progression.
Lipid Biomarker Discovery: Lipidomics facilitates the discovery of novel lipid biomarkers, which can be used for early disease detection, monitoring treatment response, and predicting disease outcomes.
Sample Requirements
| Sample type | Recommended sample size | Pre-treatment and storage |
|---|---|---|
| Tissue | 100-200 mg | Snap freezing in liquid nitrogen, stored at -80℃. |
| Urine | 200-500 μL | 5000×g 4℃ Centrifuge for 30-60min, remove supernatant, store at -80℃. |
| Serum/plasma | >100 μL | Collected serum/plasma, snap freezing in liquid nitrogen, stored at -80℃. |
| Cerebrospinal fluid, amniotic fluid, bile and other body fluids | >200 μL | 4℃ Centrifuge for 10min, (or filter using 0.22μm membrane), remove supernatant and store at -80℃. |
| Suspension cells | >1*107 | Centrifuge and collect cells after liquid nitrogen snap freezing and store at -80℃. |
| Walled cells | >1*10 | Cultured walled cells are stored in 1.5ml centrifuge tubes, snap freezing in liquid nitrogen and stored at -80℃. |
| Cell supernatant | >2 mL | centrifuge at 4℃ for 3 minutes, take the supernatant and store at -80℃. |
Case Study: Client Success Stories

Unveiling the Immunomodulatory Role of SCFAs Through Lipidomics
B cell-intrinsic epigenetic modulation of antibody responses by dietary fiber-derived short-chain fatty acids
(Nature Communications, 2020)
- Background
- Technical Approach
- Key Findings
- Conclusion
Short-chain fatty acids (SCFAs), including butyrate and propionate, are metabolic byproducts of dietary fiber fermentation by gut microbiota. Their roles in modulating immune responses—particularly B cell functions—have gained research interest due to potential applications in autoimmunity and inflammation control.
In a landmark study led by Dr. Paolo Casali's group, researchers investigated how SCFAs influence antibody production and epigenetic regulation in B cells. To dissect these complex metabolic pathways, they partnered with Creative Proteomics, leveraging our advanced lipidomics platform to quantify SCFA levels and map their impact on immune regulation.
To trace the biological activity of dietary fiber-derived SCFAs, the research team employed a multi-omics strategy. Here's how Creative Proteomics contributed:
- Lipidomics Profiling by LC-MS/MS
Creative Proteomics performed targeted lipidomics analysis using LC-MS/MS to accurately quantify butyrate and propionate levels across multiple tissues—including feces, colon, spleen, and serum. These data were essential to correlate SCFA exposure with downstream immune modulation. - Comprehensive Data Integration
Lipid profiles were mapped alongside immunological and transcriptomic markers. This included: - Flow cytometry for B cell subtype identification.
- ELISA for antibody titers (IgG1, IgA, IgE).
- RT-PCR and luciferase assays to track AID (Aicda) and Blimp1 (Prdm1) gene regulation.
- Somatic hypermutation (SHM) analysis of antibody sequences.
- Precise SCFA Quantification Revealed a Dose-Response Relationship
With the help of Creative Proteomics, the team demonstrated that elevated SCFA levels suppress class-switch recombination (CSR) and plasma cell differentiation in a dose-dependent manner. This was attributed to epigenetic inhibition of AID and Blimp1 expression in B cells. - Lipidomics Enabled Tissue-Level Insights
The quantification of SCFAs across fecal, lymphatic, and systemic compartments revealed clear metabolic signatures associated with dietary fiber intake and supplementation. These insights would not have been possible without high-sensitivity lipidomics data. - Immune Impact Validated in Multiple Mouse Models
The study spanned wild-type, T-cell-deficient, and autoimmune-prone (lupus) mice. In all models, SCFA-induced suppression of antibody responses correlated with the lipidomic fingerprint generated by Creative Proteomics.
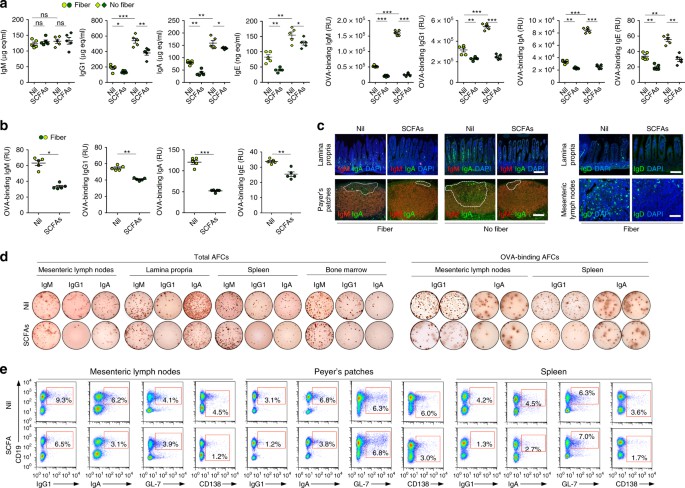 Figure 1: Dietary fibers and SCFAs dampen CSR, plasma cell differentiation and class-switched antibody responses.
Figure 1: Dietary fibers and SCFAs dampen CSR, plasma cell differentiation and class-switched antibody responses.
1a & 1b of the publication clearly illustrate the inverse relationship between SCFA concentrations and IgG1/IgA/IgE titers.
This study showcases how Creative Proteomics' lipidomics services empower immunology researchers to draw mechanistic connections between metabolites and immune regulation. By precisely mapping SCFA levels across biological compartments, we helped the client uncover a B cell-intrinsic pathway for immune suppression—offering new therapeutic avenues for autoimmunity and allergy.
Whether you're exploring microbiome-derived metabolites or systemic lipid responses, our LC-MS/MS lipidomics platform delivers the precision and reliability you need to publish high-impact findings.
People Also Ask: Common Questions About Lipidomics Services
What sample types can be analyzed?
We can analyze a variety of biological samples, including blood, tissue, cells, microbial samples, and exosomes.
How long does a typical lipidomics analysis take?
The analysis usually takes about 2–3 weeks from receiving the sample to delivering the final report.
What do clients receive?
Clients receive comprehensive quantitative data, including Principal Component Analysis (PCA), volcano plots, visualizations, and in-depth statistical analysis.
How is data quality ensured?
We ensure data quality through rigorous pre-testing, sample replicates, the use of top-tier instruments, and statistical validation to confirm the accuracy and reliability of results.
Is fatty acid testing referring to free fatty acids in biological samples?
No, fatty acid testing typically refers to fatty acids within various lipid forms, such as triglycerides and phospholipids. Free fatty acids are present in low amounts. During analysis, all lipids are hydrolyzed into fatty acids, which are then methylated (esterified). As a result, the test reflects the total fatty acid content in all lipids. However, this method does not specify the source of the fatty acids, as the analysis focuses on fatty acid methyl esters (FAMEs) using gas chromatography.
Should fatty acid composition be measured using dried or fresh samples?
Dried samples are preferred for fatty acid composition analysis. Dried samples are easier to transport and avoid the interference of water, which can affect the methylation process when using methanol-sulfuric acid as a reagent.
Regarding Quantification
Gas chromatography, coupled with a flame ionization detector (FID), is commonly used for fatty acid quantification. The FID responds to nearly all volatile organic compounds, with a consistent response for hydrocarbon compounds (carbon number ≥3) and their isomers. In quantification, the ratio of fatty acid compound concentration to the internal standard is directly proportional to the ratio of their peak areas.
When a mass spectrometer is used as a detector, the ionization efficiency of compounds varies, which must be accounted for in the analysis.
Learn about other Q&A about proteomics technology.
Our Lipidomics Review
Here are some publications in lipidomics research from our clients:

- White matter lipid alterations during aging in the rhesus monkey brain. Dimovasili, C., Vitantonio, A. T., Conner, B., Vaughan, K. L., Mattison, J. A., & Rosene, D. L. 2024. https://doi.org/10.1007/s11357-024-01353-3
- Annexin A2 Controls Lipid Composition Necessary for Junctional Stability in Response to Sphingosine 1-Phosphate (Doctoral dissertation). Sveeggen, T. 2022. https://oaktrust.library.tamu.edu/handle/1969.1/198138
- Characterising Chinese Hamster Ovary cell extracellular vesicle production in biopharmaceutical manufacturing (Doctoral dissertation, University of Sheffield). O'Donnell, F. 2022. https://etheses.whiterose.ac.uk/33062/
- Multi‐omics identify xanthine as a pro‐survival metabolite for nematodes with mitochondrial dysfunction. Anna Gioran et al. 2019. https://doi.org/10.15252/embj.201899558
References
- Rivier, C., Camès, B., Lamouroux, C., Casanova, F., Daubizit, M., Neulat, C., Organista, M., Pochon, P., & Roudil, D. (2012). Development and Validation of Methods for the Analysis of Reprocessing Solvents: Role of CETAMA Working Group 24. Procedia Chemistry. https://doi.org/10.1016/j.proche.2012.10.107
- Gregor, A., Malleier, M., Auñon-Lopez, A., Auernigg-Haselmaier, S., König, J., & Pignitter, M. (2025). Glutathione Contributes to Caloric Restriction-Triggered Shift in Taurine Homeostasis. Nutrients, 17(5), 777. DOI: 10.3390/nu17050777
- Shi, F., Li, L., Wu, X., Wang, Y., Niu, R., & Niu, R. (2024). Building Vulnerability to Landslides: Broad-Scale Assessment in Xinxing County, China. Sensors, 24(13), 4366. https://doi.org/10.3390/s24134366
- Sales-Dias, J., Silva, G., Lamy, M., Ferreira, A., & Barbas, A. (2019). The Notch ligand DLL1 exerts carcinogenic features in human breast cancer cells. PLoS One, 14(5), e0217002. DOI: 10.1371/journal.pone.0217002















