The primary structure of a glycan is consist of the nature and order of monosaccharide residues, the configuration and position of glycosidic linkages, and the nature and location of non-glycan entities to which they are attached. In addition, different glycans can be attached to different sites on a glycoprotein. Moreover, it is often three-dimensional features, or particular surface distributions of glycans, that are recognized by glycan-binding proteins. The structural characterization of glycans includes monosaccharide residue composition, sequence and linkage types. Characterization of these diverse structural features requires an array of different methods.
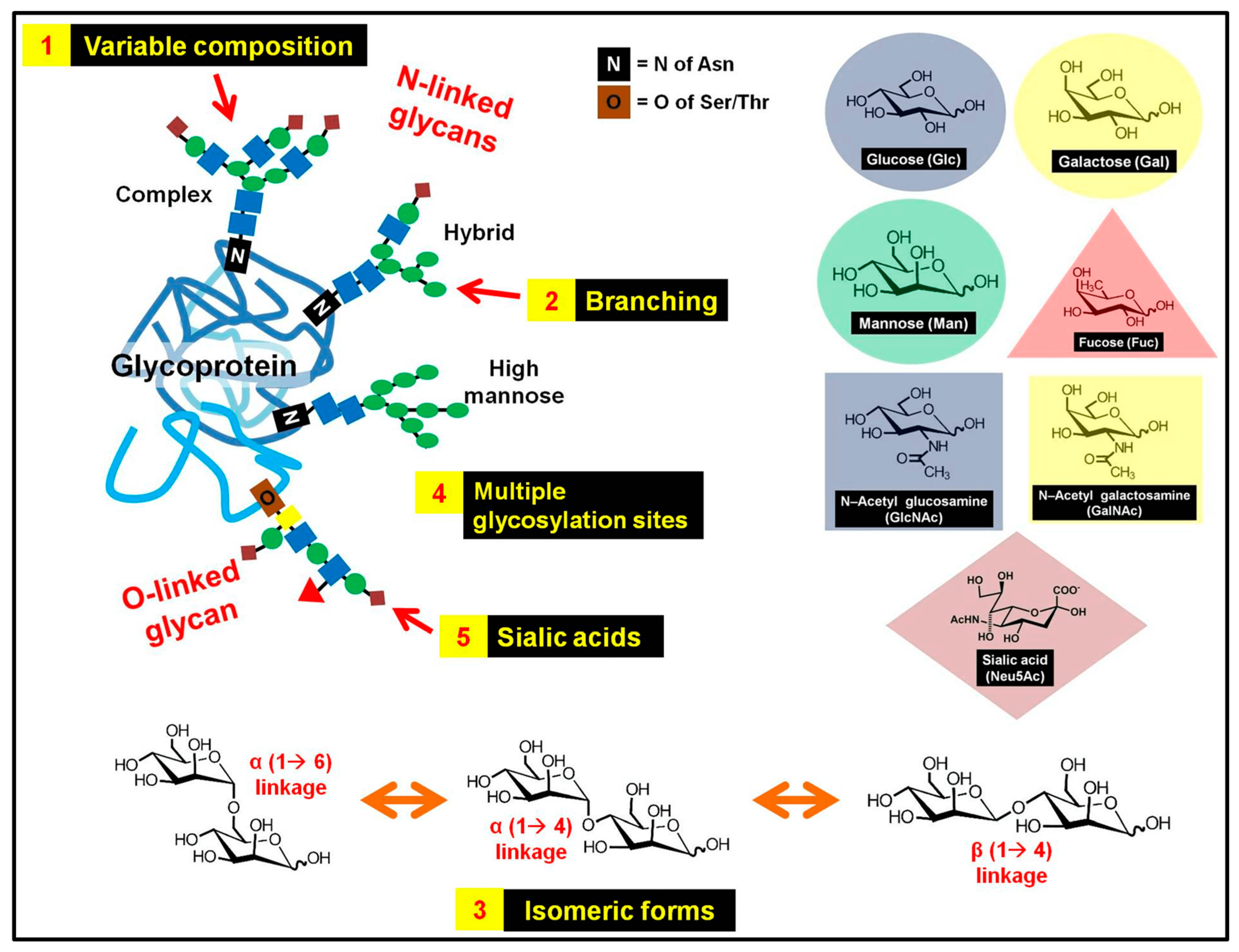 Figure 1. The complex and heterogeneous structure of glycans
Figure 1. The complex and heterogeneous structure of glycans
Monosaccharide composition analysis
There is a convenient and informative approach for the qualitative analysis of monosaccharide composition of a glycan. After hydrolysis of a glycan into monosaccharides, colorimetric reactions can be used to determine the total amount of hexose, hexuronic acid, or hexosamine in the sample. This approach does not require prior release of glycans from the glycoconjugates.
The quantitative monosaccharide analysis provides estimated molar ratios of individual monosaccharide components of glycans. The analysis involves several steps: cleavage of all glycosidic linkages (generally by acid hydrolysis), fractionation of the cleaved monosaccharides, detection by GC-MS or HPLC, and quantification.
Linkage analysis
See N-glycan linkage analysis and O-glycan linkage analysis
Glycan sequence analysis
The highly specific exoglycosidases are usually used to determine the sequence and structure of glycans. This enzymatic analysis is a powerful technique and applies either sequentially or in a matrix array. Exoglycosidases remove terminal sugar groups from the non-reducing end of a glycan, but do not cleave internal bonds between monosaccharides. The linkage and sugar groups of the removed glycan residues can be identified by using various linkage-specific exoglycosidases.
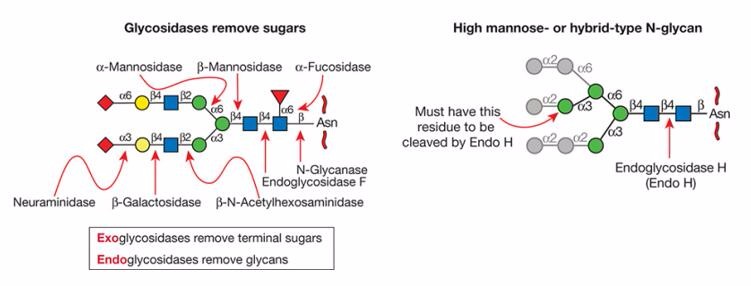 Figure 2. Glycosidases used for structural analysis
Figure 2. Glycosidases used for structural analysis
The exoglycosidase digestion coupled with high-performance separation techniques such as electrophoresis or chromatography is an efficient and fast approach for the structural characterization of oligosaccharides.
Three-dimensional Glycan Structure
Glycans do not possess a well-defined three-dimensional structure in solution due to the inherent flexibility of glycosidic linkages. Crystal structures are available for many mono- and oligosaccharides. Some molecular modeling softwares can be used to generate approximate models of highly complex glycans from the simple sugar structures. The total characterization of glycan conformation and dynamics in solution remains an area of active research and relatively based on NMR spectroscopy result. The initial three-dimensional glycan structure can be built up by using some convenient tools such as carbohydrate builder, and then can be further refined by incorporating the solution NMR data such as NOEs, angular and long range distance constraints from residual dipolar couplings and paramagnetic effects measurements.
As one of the leading companies in the omics field with over years of experience in omics study, Creative Proteomics provides glycomics analysis service customized to your needs. Contact us to discuss your project.
How to place an order

*If your organization requires signing of a confidentiality agreement, please contact us by email.


















