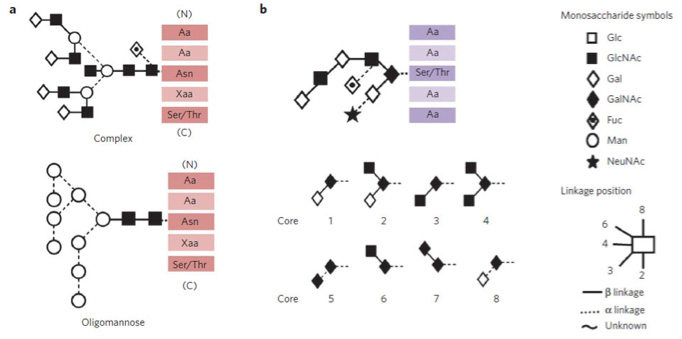
Usually it is not necessary to determine the glycosidic linkages because they rarely cause effect on the function of glycoproteins. However, due to the high complexity of glycan structures of glycoprotein, assessing the configuration and position of glycosidic linkages of the N-glycan is helpful. Up to now, there are five common N-glycan linkages known. Among these five linkages, N-acetylglucosamine to asparagine (GlcNAcβ1-Asn) is the most common one. Other types of linkages to Asn include: glucose in laminin of both mammals and Archaea, N-acetylgalactosamine (GalNAc) in Archaea, rhamnosein bacteria. And a report shows that glucose linkage to argninein a sweet corn.
N-glycans are released from glycoproteins by PNGase F reaction. The released N-glycans are labeled with 2-aminobenzamide (2-AB), followed by digestion with multiple exoglycosidases. Finally, glycans are analyzed by the Hydrophilic Interaction Chromatography (HILIC) with a fluorescence and mass spectrometric detection. Several steps are involved in N-Glycan Linkage analysis:
- Sample preparation
- N-Glycans release
- 2-aminobenzamide (2-AB) labeling and the labeled glycans are reacted sequentially with a series of exoglycosidases.
- UPLC-HILIC-FLD based analysis;
- Additionally, confirmation the detailed glycan structures by MS/MS fragmentation can be done if required.
*What does Endoglycosidase used for?
Endoglycosidase releases oligosaccharides from glycoproteins or glycolipids. Normally it does not release oligosaccharides from conjugated protein and lipid molecules. Endoglycosidase functions by breaking the glycosidic bonds between two sugar monomer in the polymer.
Endoglycosidase H: release oligo mannose and hybrid N-glycans
Endoglycosidase F: release simple biantennary N-glycans
How to place an order:

*If your organization requires signing of a confidentiality agreement, please contact us by email
As one of the leading companies in providing omics analysis service, Creative Proteomics provides reliable glycomics analysis service for our customers. Contact us for more detailed information!





