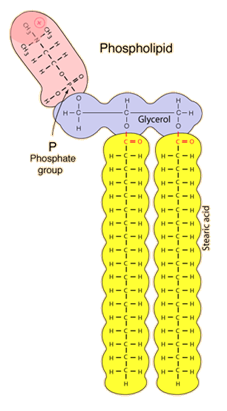
Neutral lipids are the triglycerides which consist of three fatty acids attached to a glycerol molecule. Cholesterol is also a neutral lipid. Neutral lipids such as cholesterol esters and triglycerides are insoluble in water and essentially lack detergent properties. Phospholipids are a class of lipids that are a major component of all cell membranes. Phospholipids belong to the lipid family of biological polymers. A phospholipid is composed of two fatty acids, a glycerol unit, a phosphate group and a polar molecule. There are no simple methods available for analysis of phospholipids since the close range of polarity between different phospholipid species makes detection difficult.
Scientist at Creative Proteomics ultilize GC methods to Profile Neutral or Phospho-Lipids. Lipids are extracted from plasma, tissue or cells using a modified Folch method, and analyzed as either total lipid content, or separated on a TLC plate for analysis of each individual lipid class. Lipids are hydrolyzed prior to GC analysis. GC Analysis provides quantitation of lipids in each class by fatty acid chain length, but does not provide individual molecular structures. Researchers can choose how many total lipid classes to analyze, and should consult the Core Director for pricing based on the number of lipid classes requested. This unique method covers more than 1,100 molecular species across 16 lipid classes, including principle phospholipid, sphingolipid and neutral lipid classes.
Technology Platform:
Gas Chromatography
Sample Type:
Plasma, Tissue, Cells
Minimum sample quantity
- Normal Volume: 200 uL plasma, 20 mg tissue, 1e7 cells
- Minimal Volume: 50uL, 5 mg tissue, 6 e6 cells
| Lipid Classes could monitored in This Service | ||
|---|---|---|
| Total Lipids TL-FA | Phospholipids PL-FA COMP | Neutral Lipids FA CompMG |
| 14:0 | 14:0 | 14:0 |
| 14:1 | 14:1 | 14:1 |
| 16:0 | 16:0 | 16:0 |
| 16:1 | 16:1 | 16:1 |
| 18:0 | 18:0 | 18:0 |
| 18:1 (n-7) | 18:1 (n-7) | 18:1 (n-7) |
| 18:1 (n-9) | 18:1 (n-9) | 18:1 (n-9) |
| 18:2 | 18:2 | 18:2 |
| 18:3 | 18:3 | 18:3 |
| 20:0 | 20:0 | 20:0 |
| 20:1 | 20:1 | 20:1 |
| 20:2 | 20:2 | 20:2 |
| 20:3 | 20:3 | 20:3 |
| 20:4 | 20:4 | 20:4 |
| 20:5 | 20:5 | 20:5 |
| 22:0 | 22:0 | 22:0 |
| 22:1 | 22:1 | 22:1 |
| 22:4 | 22:4 | 22:4 |
| 22:5 | 22:5 | 22:5 |
| 22:6 | 22:6 | 22:6 |
| 24:0 | 24:0 | 24:0 |
| 24:1 | 24:1 | 24:1 |
Ordering Procedure:

With integrated set of separation, characterization, identification and quantification systems featured with excellent robustness & reproducibility, high and ultra-sensitivity, Creative Proteomics provides reliable, rapid and cost-effective Lipidomics services.





