Hydroxy-Eicosatetraenoic Acids (HETEs) Analysis Service
Creative Proteomics provides targeted hydroxy-eicosatetraenoic acids (HETEs) analysis using high-resolution Orbitrap MS and UPLC technology to deliver ultra-sensitive, quantitative profiling of over 50 lipid mediators. Our service helps researchers decode lipid signaling pathways, monitor oxidative stress responses, and support mechanistic studies in inflammation and metabolism.
Submit Your Request Now
×- What We Provide
- Advantage
- Workflow
- Technology Platforms
- Sample Requirements
- FAQ
- Publication
What are Hydroxy-Eicosatetraenoic Acids (HETEs)?
Hydroxy-eicosatetraenoic acids (HETEs) are oxygenated metabolites derived from arachidonic acid (AA) through enzymatic pathways involving cytochrome P450 (CYP450), lipoxygenases (LOX), and cyclooxygenases (COX). These bioactive lipids play critical roles in inflammation, vascular regulation, oxidative stress, and cellular proliferation. For instance, 20-HETE is a key CYP450-derived metabolite implicated in blood pressure regulation and cardiovascular pathologies, while 12(S)-HETE and 5-HETE are linked to inflammatory responses. Accurate analysis of HETEs is essential for understanding their physiological and pathological mechanisms in diseases such as hypertension, cancer, and myocardial injury.
HETEs Analysis Service Offered by Creative Proteomics
- Targeted Quantification: Absolute quantification of specific HETEs (e.g., 5-HETE, 12(S)-HETE, 20-HETE) using isotope-labeled internal standards (e.g., 12(S)-HETE-d4) to ensure precision.
- Metabolite Profiling: Identification and semi-quantification of over 50 HETEs and related metabolites, including epoxyeicosatrienoic acids (EETs) and leukotrienes.
- Pathway Analysis: Mapping HETEs biosynthesis and downstream signaling cascades (e.g., CYP450/LOX pathways) through integrated bioinformatics tools.
- Custom Lipidomics Services: Tailored lipidomics profiling to analyze the complete spectrum of lipid metabolites, including HETEs, prostaglandins, leukotrienes, and other eicosanoids.
List of Detected HETEs and Related Metabolites
| Analyte Name | Related Metabolites | Associated Pathways |
|---|---|---|
| 5-HETE | 5-Oxo-ETE, 5,6-EET, 5,6-DiHETE | Arachidonic Acid Metabolism (5-LOX Pathway) |
| 8-HETE | 8-Oxo-ETE, 8,9-EET, 8,9-DiHETE | LOX & CYP450 Pathways |
| 11-HETE | 11,12-EET, 11,12-DiHETE | Cytochrome P450 Monooxygenase Pathway |
| 12(S)-HETE | 12-oxo-ETE, 12,13-EET, 12,13-DiHETE | 12-LOX Pathway |
| 12(R)-HETE | R-isomer derivatives | CYP450 Pathway |
| 15(S)-HETE | 15-oxo-ETE, 15,16-EET, 15,16-DiHETE | 15-LOX Pathway |
| 15(R)-HETE | R-isomer derivatives | Aspirin-triggered Lipoxin Pathway |
| 18-HETE | 18-HEPE (from EPA), 18-oxo-HETE | Specialized Pro-resolving Mediator (SPM) Pathway |
| 20-HETE | 20-COOH-AA, 20-Oxo-ETE | CYP4A/4F Pathway |
| 5,15-diHETE | 5,15-DiHETE | Dual LOX Pathway |
| 8,15-diHETE | 8,15-DiHETE | LOX Interaction Pathway |
| Lipoxins (e.g., LXA4) | Aspirin-triggered lipoxins, 15-epi-LXA4 | Lipoxin Synthesis Pathway |
| EETs (Epoxyeicosatrienoic acids) | 5,6-EET, 8,9-EET, 11,12-EET, 14,15-EET | CYP450 Epoxygenase Pathway |
| DiHETEs | 5,6-DiHETE, 8,9-DiHETE, 11,12-DiHETE, 14,15-DiHETE | Soluble Epoxide Hydrolase Pathway |
Advantages of HETEs Assay
- Ultra-High Sensitivity: Detection limits as low as 1 pg/mL, enabling precise quantification of trace-level HETEs in plasma, serum, and tissue samples.
- High-Throughput Quantification: Simultaneous detection and quantification of 20+ HETE-related metabolites in a single run.
- Exceptional Reproducibility: Inter-assay coefficient of variation (CV) maintained below 10%, ensuring consistent and reliable results across batches.
- Wide Dynamic Range: Linear quantification range from 1 pg/mL to 10,000 pg/mL, suitable for both physiological and pathological conditions.
- Advanced Instrumentation: Powered by Thermo Fisher Q Exactive™ Orbitrap MS coupled with Waters ACQUITY UPLC®, providing resolution up to 140,000 FWHM for high-confidence identification.
- Pathway-Level Insight: Data integrated with pathway enrichment analysis covering >10 lipid signaling pathways, including 5-LOX, 12-LOX, 15-LOX, CYP450, and SPM biosynthesis.
Workflow for HETEs Analysis Service
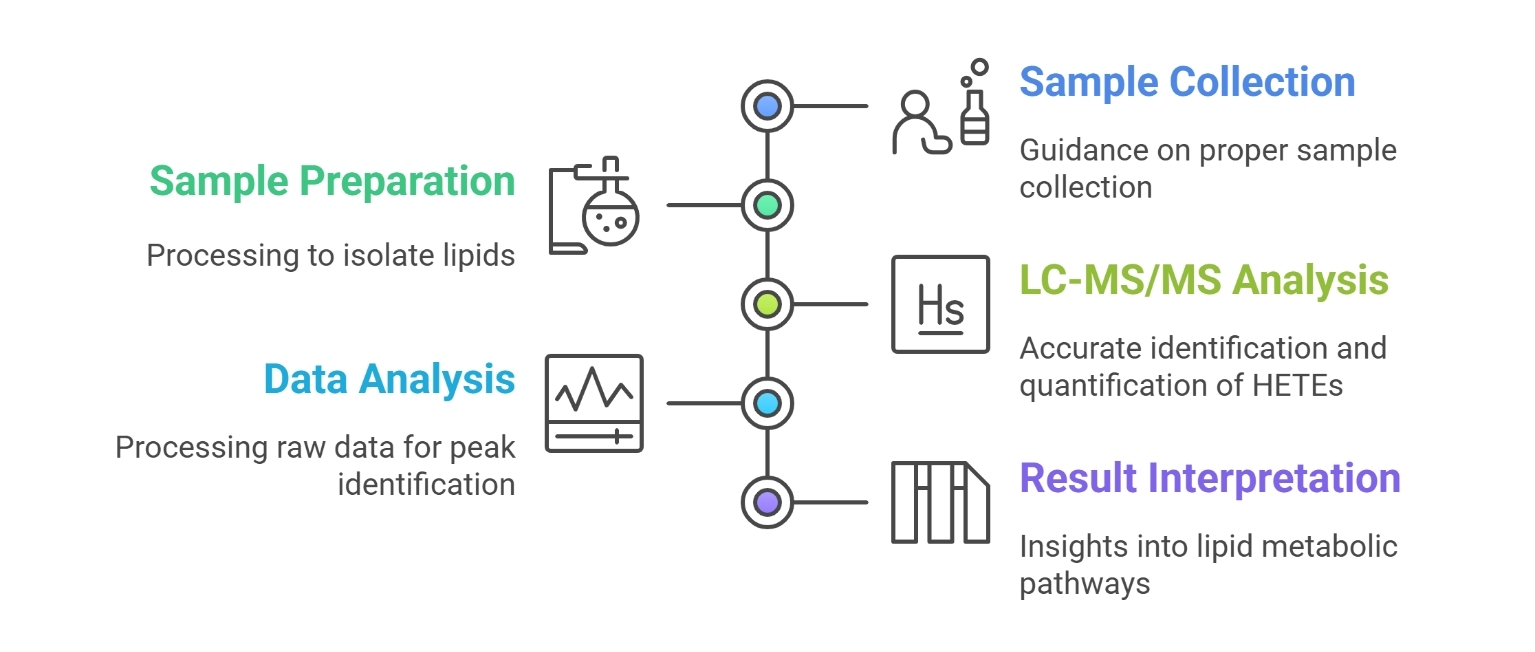
Technology Platform for HETEs Analysis Service
- UPLC System: Waters ACQUITY UPLC®
- Mass Spectrometer: Thermo Scientific™ Q Exactive™ Orbitrap
- Ionization Mode: Negative ESI (Electrospray Ionization)
- Resolution: Up to 140,000 FWHM at m/z 200
- Quantification Mode: Full scan with targeted SIM (Selected Ion Monitoring)
- Software: TraceFinder™, Compound Discoverer™, LipidSearch™

Waters ACQUITY UPLC System (Figure from Waters)

Thermo Fisher Q Exactive (Figure from Thermo Fisher)
Sample Requirements for HETEs Analysis Service
| Sample Type | Required Volume/Amount |
|---|---|
| Plasma / Serum | ≥ 200 μL |
| Tissue (e.g., liver, brain) | ≥ 50 mg (fresh or frozen) |
| Cell Pellet | ≥ 1 × 10⁷ cells |
| Culture Medium / Supernatant | ≥ 500 μL |
| Bronchoalveolar Lavage Fluid (BALF) / Urine | ≥ 500 μL |
| Dried Blood Spot (DBS) | ≥ 3 punches (6 mm diameter) |
Applications of HETEs Assay Service
Lipid Mediator Network Mapping
Analyze the role of HETEs within broader lipid signaling pathways (e.g., LOX, CYP450), supporting systems biology studies.
Preclinical Drug Evaluation
Assess off-target or on-target modulation of HETE metabolism in animal models during early-stage compound testing.
Nutritional and Metabolic Research
Investigate dietary lipid impacts on eicosanoid profiles, including shifts induced by omega-3 or omega-6 fatty acids.
Oxidative and Inflammatory Stress Response
Track HETE accumulation under cellular stress conditions such as hypoxia, oxidative damage, or immune activation.
Comparative Lipidomics Across Conditions
Perform differential profiling of HETEs in control vs. experimental groups (e.g., knockout models, treatment groups).
Environmental Exposure Studies
Quantify HETE fluctuations in response to pollutants, toxins, or xenobiotics in toxicology and exposure science.
Demo

LC-MS-MS detection of HETE standards (Alexanian et al., 2013).
FAQ of HETEs Analysis Service
How should samples be shipped to ensure stability?
Samples should be shipped on dry ice using insulated containers. Avoid thawing during transit.
Can you analyze HETEs from multiple sample types in one project?
Yes, we support multi-matrix analysis with unified data output, enabling cross-comparison.
Are isotope-labeled standards included for quantification?
Yes, we use a panel of internal standards to ensure absolute quantification accuracy.
How is data delivered?
Clients receive raw data, processed result tables, chromatograms, and optional biological interpretation reports.
Can I reanalyze my sample later with a different panel?
If sufficient sample volume remains and is stored properly, we can accommodate follow-up analyses.
Do you provide custom metabolite panels or spike-in validation?
Yes, custom panels and method validation with client-supplied spike-ins are available upon request.
What are the sample rejection criteria?
Samples with hemolysis, visible contamination, or insufficient volume will not be processed.
Is method performance data (LOD/LOQ, CV%) available upon request?
Absolutely, method validation data can be provided for transparency and reproducibility assurance.
What's the minimum number of samples required to start a project?
We accept projects starting from as few as 3 samples, although for statistical robustness, we recommend ≥6 biological replicates per group.
Can you handle samples from multiple species (e.g., mouse, rat, human)?
Yes, our validated platform supports HETE detection across multiple species. Please specify species during submission for accurate calibration.
Can I receive raw data files in vendor format?
Absolutely. We provide Thermo .raw files alongside processed data (e.g., Excel, PDF reports). Open formats (e.g., mzML) are also available on request.
How are data normalized across different sample types?
Depending on the sample, data are normalized by volume (e.g., μL), weight (e.g., mg tissue), or cell number to ensure comparability.
Do you provide support for publication or grant applications?
Yes, we offer detailed method descriptions, raw spectra, and figure generation to support publication or proposal submissions.
What's the best way to avoid sample degradation before shipping?
Use dry ice and insulated containers, avoid repeated freeze-thaw, and if possible, aliquot samples. Ship samples early in the week to avoid weekend delays.
Can this analysis be combined with other lipid panels?
Yes, HETE profiling can be integrated with broader oxylipin, eicosanoid, or SPM (specialized pro-resolving mediator) panels as part of a custom lipidomics solution.
Learn about other Q&A.
Publications
Here are some publications in Lipidomics research from our clients:

- The olfactory receptor Olfr78 promotes differentiation of enterochromaffin cells in the mouse colon. 2024. https://doi.org/10.1038/s44319-023-00013-5
- Microbial dysbiosis associated with impaired intestinal Na+/H+ exchange accelerates and exacerbates colitis in ex-germ free mice. 2018. https://doi.org/10.1038/s41385-018-0035-2
- Summative and ultimate analysis of live leaves from southern US forest plants for use in fire modeling. 2020. https://doi.org/10.1152/ajpgi.00184.2023
- Annexin A2 modulates phospholipid membrane composition upstream of Arp2 to control angiogenic sprout initiation. 2023. https://doi.org/10.1096/fj.202201088R
- Loss of G0/G1 switch gene 2 (G0S2) promotes disease progression and drug resistance in chronic myeloid leukaemia (CML) by disrupting glycerophospholipid metabolism. 2022. https://doi.org/10.1002/ctm2.1146
Reference
- Alexanian, Anna, and Andrey Sorokin. "Targeting 20-HETE producing enzymes in cancer–rationale, pharmacology, and clinical potential." OncoTargets and therapy (2013): 243-255. https://doi.org/10.2147/OTT.S31586







