Kurzawa et al. (2020) “A computational method for detection of ligand-binding proteins from dose range thermal proteome profiles.” Nature Communications 11, 5783. https://doi.org/10.1038/s41467-020-19529-8
Background
Detecting precise off-target interactions of epigenetic drugs inside living cells is absolutely critical for understanding clinical drug toxicity. Traditional methods often struggle with untargeted, proteome-wide profiling without permanently modifying the drug structure, which can introduce massive artifacts. In a highly cited, landmark study, researchers utilized 2D-TPP to seamlessly map the full proteomic interactome of the HDAC8 inhibitor PCI-34051.
Methods
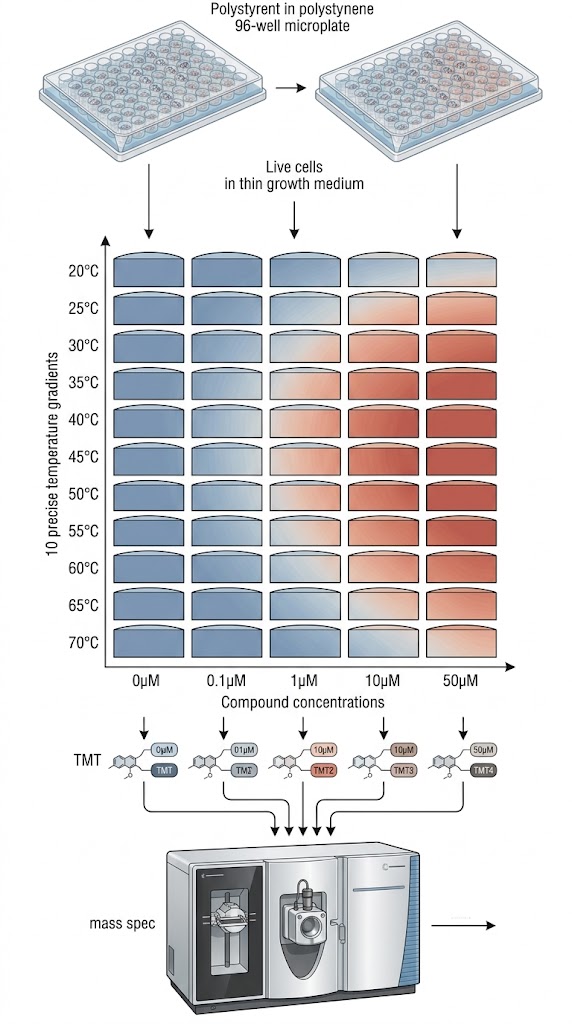
Live cells were treated with a wide concentration gradient of the inhibitor PCI-34051 across an equally expansive gradient of temperatures. The soluble protein fractions were subsequently labeled with multiplexed TMT reagents and analyzed via quantitative LC-MS/MS to construct a highly detailed, comprehensive dose-range thermal profile matrix.
Results
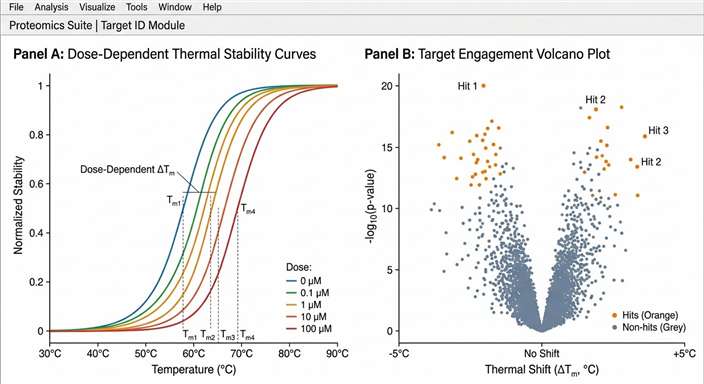
Using rigorous, non-linear curve-fitting algorithms designed to process the 2D matrix, the researchers successfully verified engagement with the primary known target (HDAC8) with a distinct dose-response stabilization. More importantly, as clearly demonstrated in Figure 4 of the referenced study, the dose-range thermal profiling confidently identified leucine aminopeptidase 3 (LAP3) as a direct, previously entirely unknown off-target of the drug. The data showed distinct, concentration-dependent thermal stabilization curves for LAP3 achieving a calculated EC50 of stabilization.
Conclusions
2D-TPP successfully deconvoluted the complex off-target landscape of the inhibitor in a completely label-free, physiologically relevant manner. This elegantly demonstrates the immense power of 2D-TPP in safety profiling, toxicity prediction, and uncovering hidden mechanisms of action without the need for synthetic chemistry interventions.