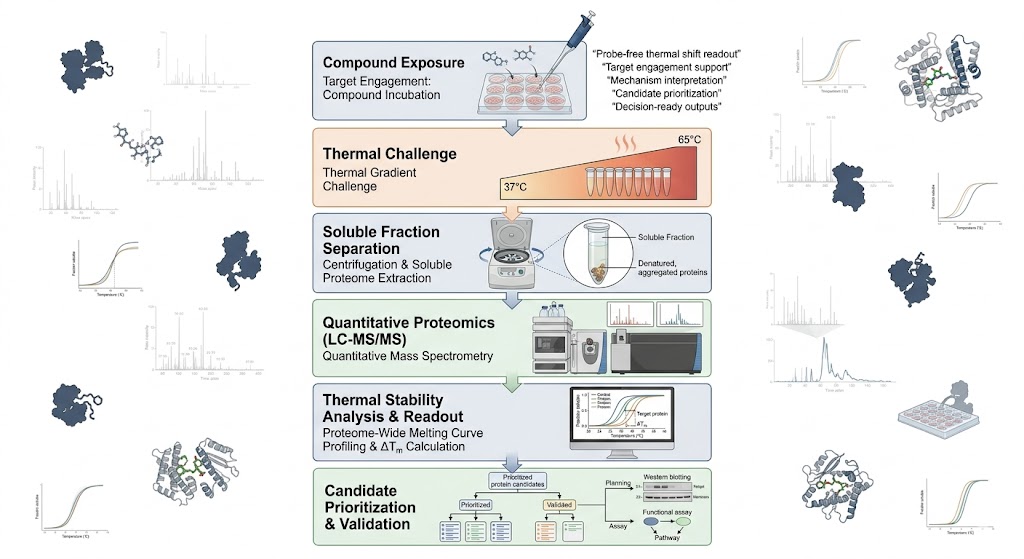
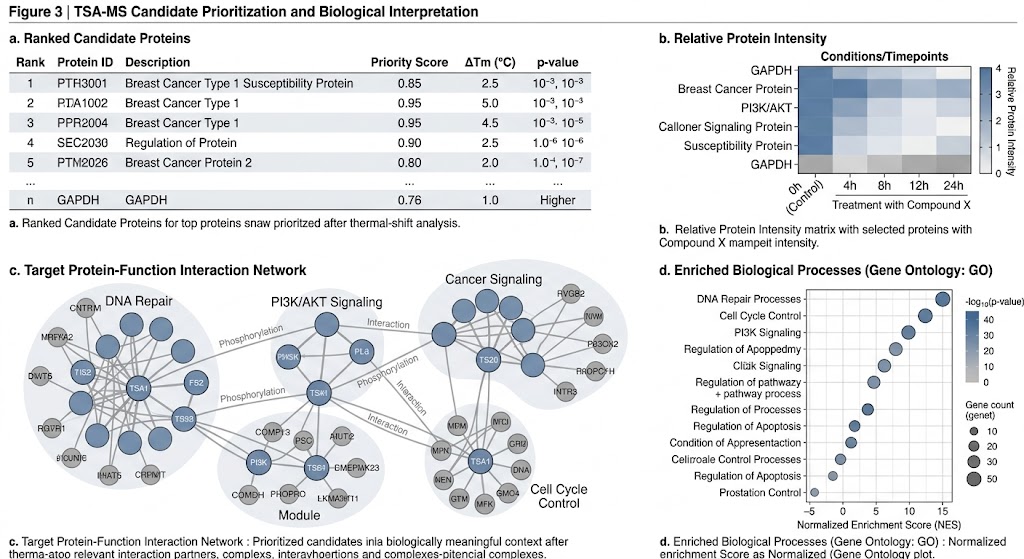
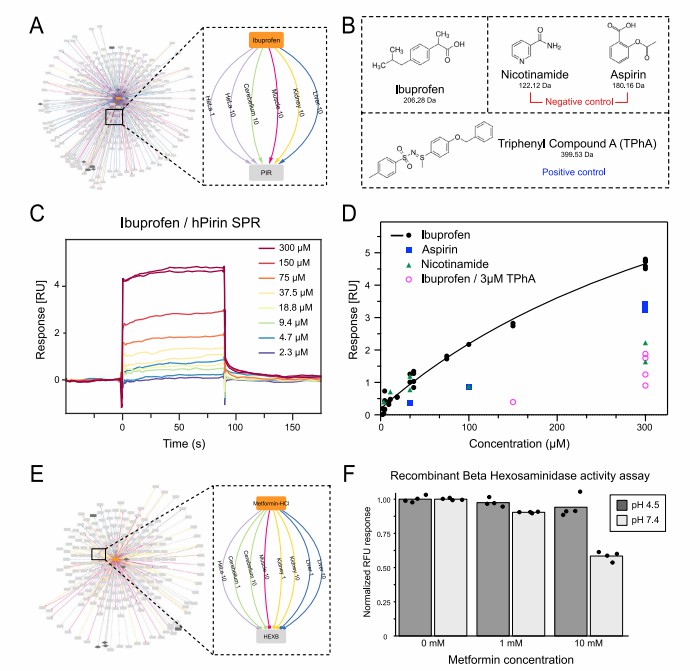
Service Overview: Study Design, Workflow, Deliverables, and Bioinformatics
The same TSA-MS platform should not be run the same way for every project. We shape the design around what you need the data to answer, whether that is target engagement, mechanism-of-action interpretation, target deconvolution, or comparative review across multiple compounds or exposure conditions.
For target engagement-focused work, we emphasize matched exposed and control groups, temperature challenge design, replicate consistency, and a result structure that makes it easier to move from global shifts to shortlisted proteins. For mechanism or deconvolution work, we put more attention on biological context, candidate prioritization logic, and interpretation layers that go beyond a simple ranked list.
Before experimental kickoff, we typically align on sample type and biological context, compound groups and controls, replicate structure, thermal challenge design, and the kind of output that will be most useful for your next experiment. If you are also considering neighboring methods, you may want to compare TSA-MS with MS-based Proteome-wide Thermal Stability Profiling, Thermal Proteome Profiling (TPP), PISA, or Limited Proteolysis-MS (LiP-MS) before locking the final design.
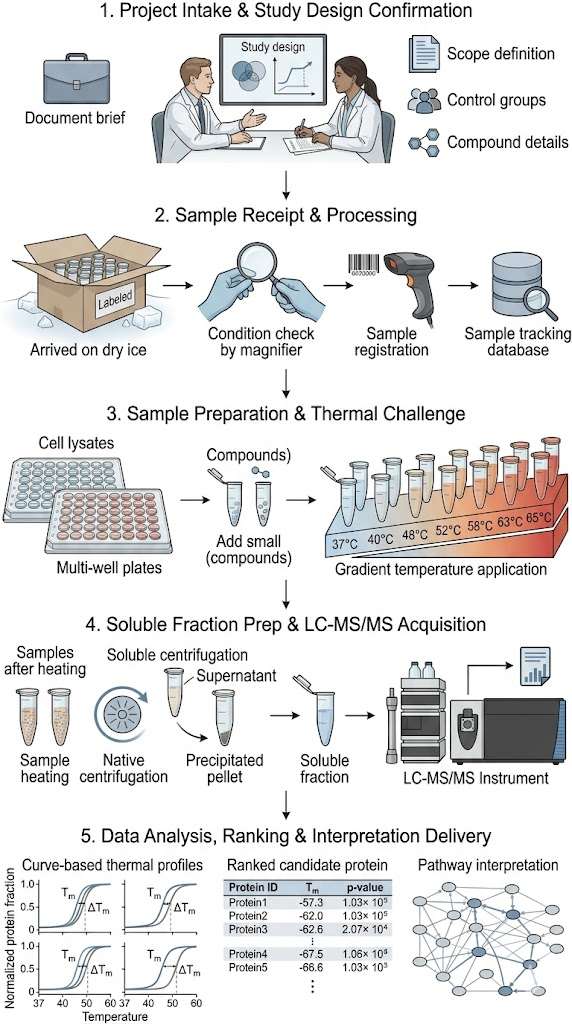
STEP 1Project intake and study design confirmation
We review your study objective, sample matrix, compound grouping, controls, and replicate structure, then confirm whether the project is best framed for target engagement, mechanism analysis, or comparative profiling.
STEP 2Sample receipt, condition check, and registration
When samples arrive, we verify labeling, grouping, replicate completeness, storage condition, and container suitability because pre-analytical variation can directly affect stability-shift interpretation.
STEP 3Sample preparation and thermal challenge
Samples proceed into compound incubation or condition setup, thermal challenge across defined points, separation of soluble fractions, and preparation for MS-compatible analysis.
STEP 4LC-MS/MS acquisition and data processing
Prepared fractions are analyzed by mass spectrometry, followed by protein identification, quantification, normalization, and thermal-shift assessment, with attention to signal consistency and group comparability.
STEP 5Interpretation delivery for follow-up review
We organize summary observations, ranked candidates, representative thermal profile views, and interpretation notes into a structure that supports internal review and follow-up planning.
QCCheckpoints throughout the workflow
Key checkpoints include sample labeling and group integrity, replicate completeness, sample condition at receipt, extraction consistency, usable MS signal quality, and fit between output structure and study objective.
What you receive
- Protein and peptide identification and quantification tables
- Normalized abundance matrix across thermal conditions
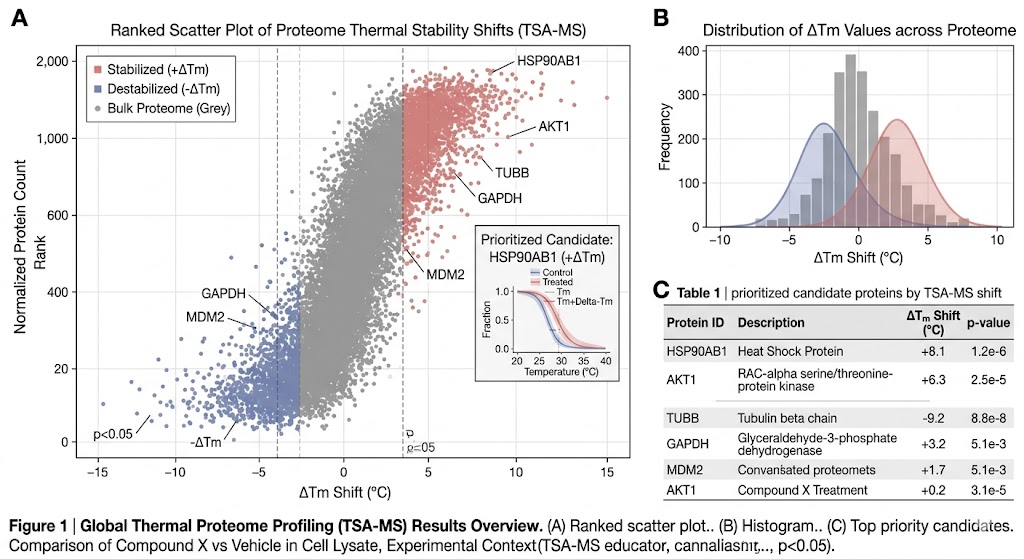
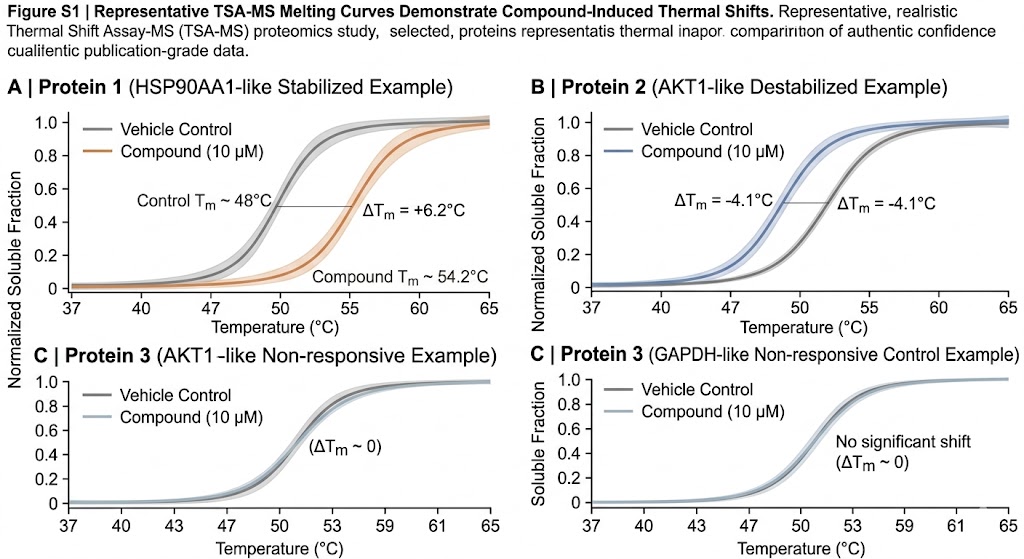
- Thermal profile fitting and shift assessment
- Ranked candidate table
- Functional annotation and pathway context
- Project summary report with interpretation notes
Bioinformatics analysis
The analytical layer is often where TSA-MS becomes truly useful. Our baseline analysis scope can include protein identification and quantification, matrix normalization, thermal profile assessment, candidate ranking, annotation of prioritized proteins, and pathway-oriented interpretation summary. If your study needs a more decision-focused readout, we can expand the analysis toward replicate-aware advanced statistics, target-family-focused interpretation, comparison across multiple compounds or exposure conditions, mapping against hypotheses from CETSA-MS, LiP-MS, or ABPP-MS, and support for orthogonal validation planning.