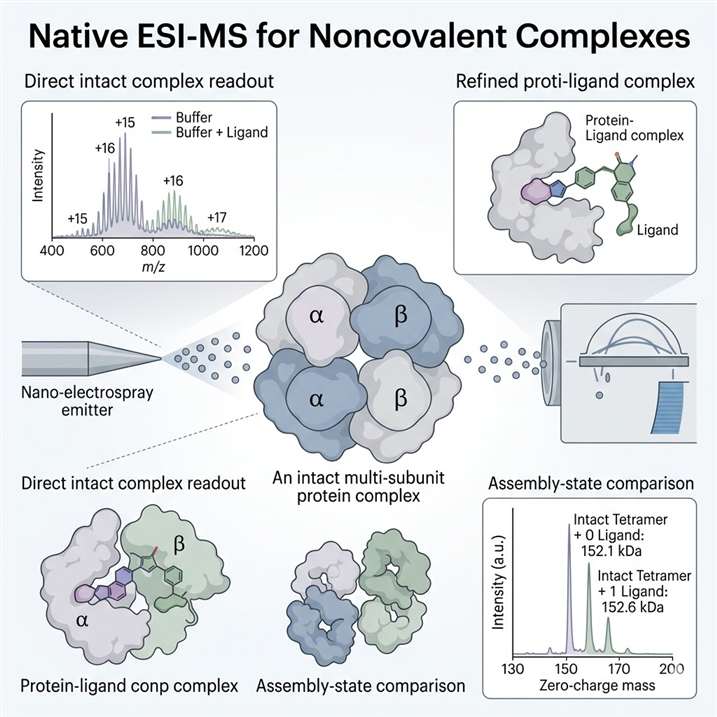
Native ESI-MS for Noncovalent Complexes
Direct analysis of intact noncovalent complexes under native-like conditions.
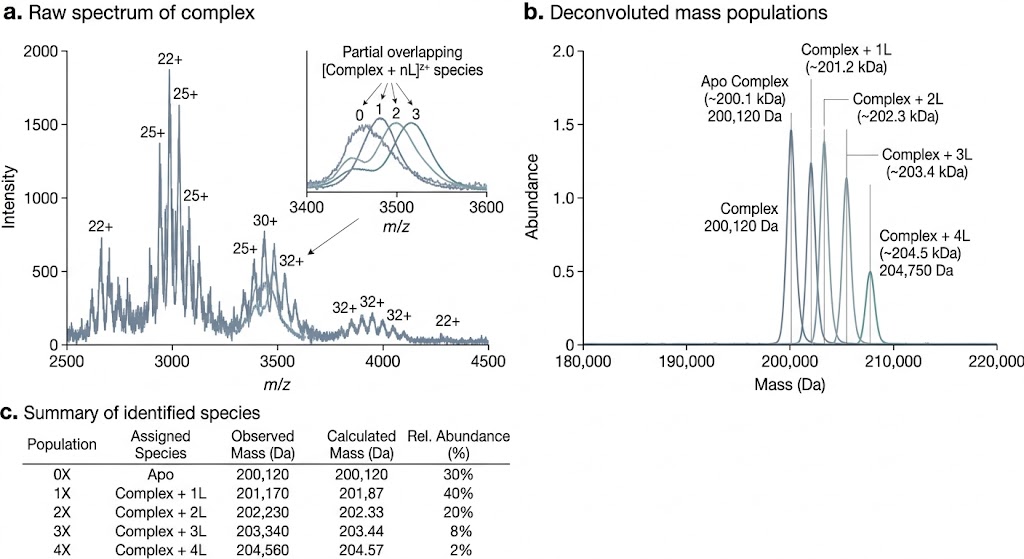
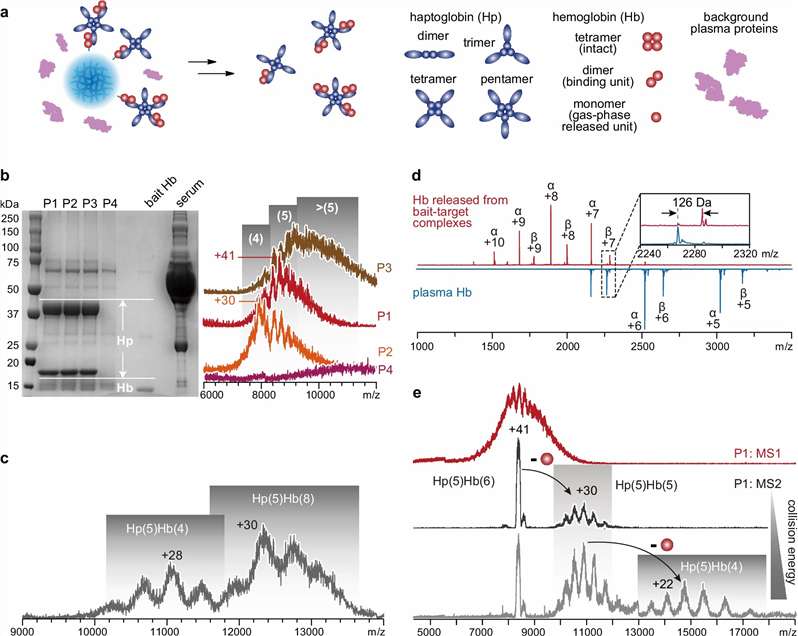
Native ESI-MS helps us analyze intact noncovalent complexes under native-like conditions and turn complex mass spectra into clear project answers. We use this approach to confirm complex formation, assess stoichiometry and occupancy, and compare assembly-state changes across ligands, constructs, and experimental conditions.
Key Advantages:
- Direct readout of intact noncovalent complexes.
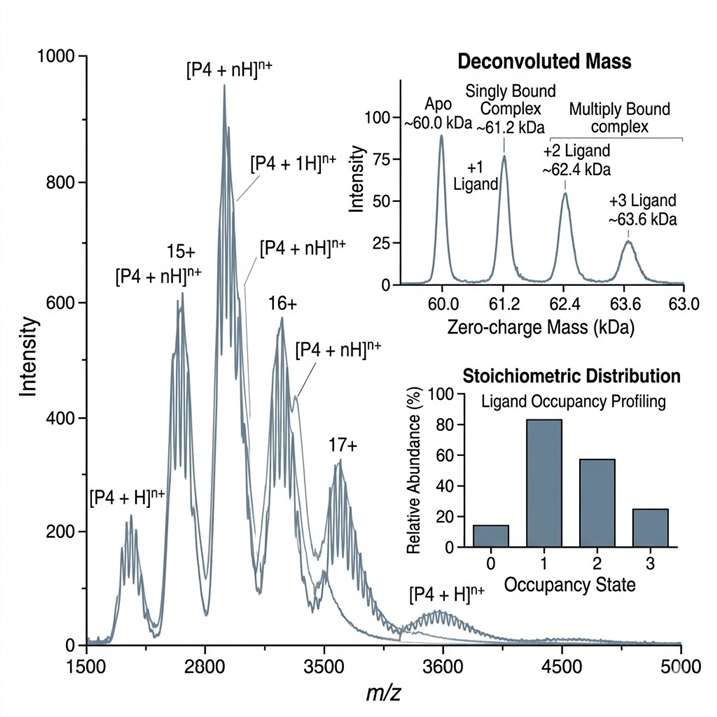
- Stoichiometry and occupancy assessment.
- Support for protein-ligand and protein-protein studies.
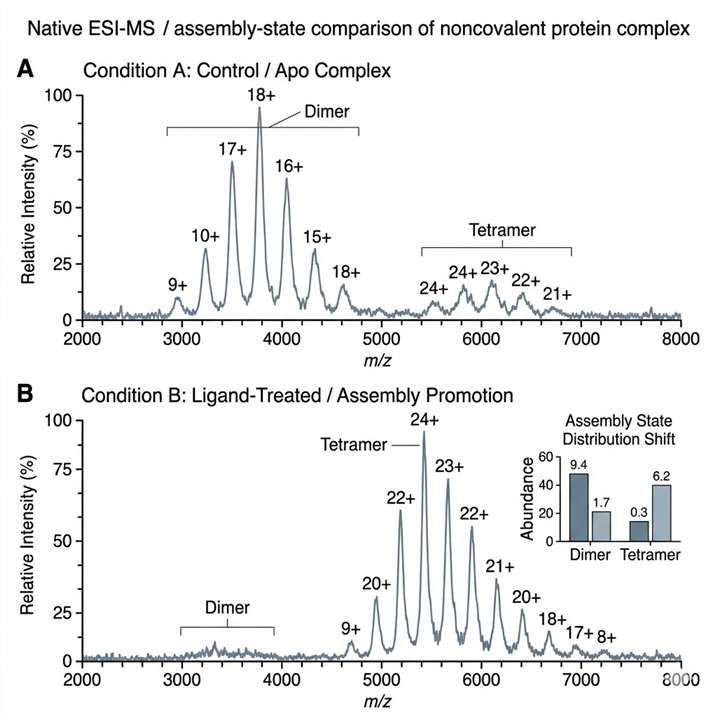
- Comparison of assembly states across conditions.
- Clear project-fit review before analysis.