Targeting Reactive Lysines in Live Cells
Reference link: https://pmc.ncbi.nlm.nih.gov/articles/PMC10988577/
Background
While cysteine profiling is standard, identifying reactive lysines globally in live cells remains difficult due to lysine's lower physiological nucleophilicity and high proteome abundance. Mapping these sites is crucial for discovering new allosteric pockets and molecular glue degraders.
Methods
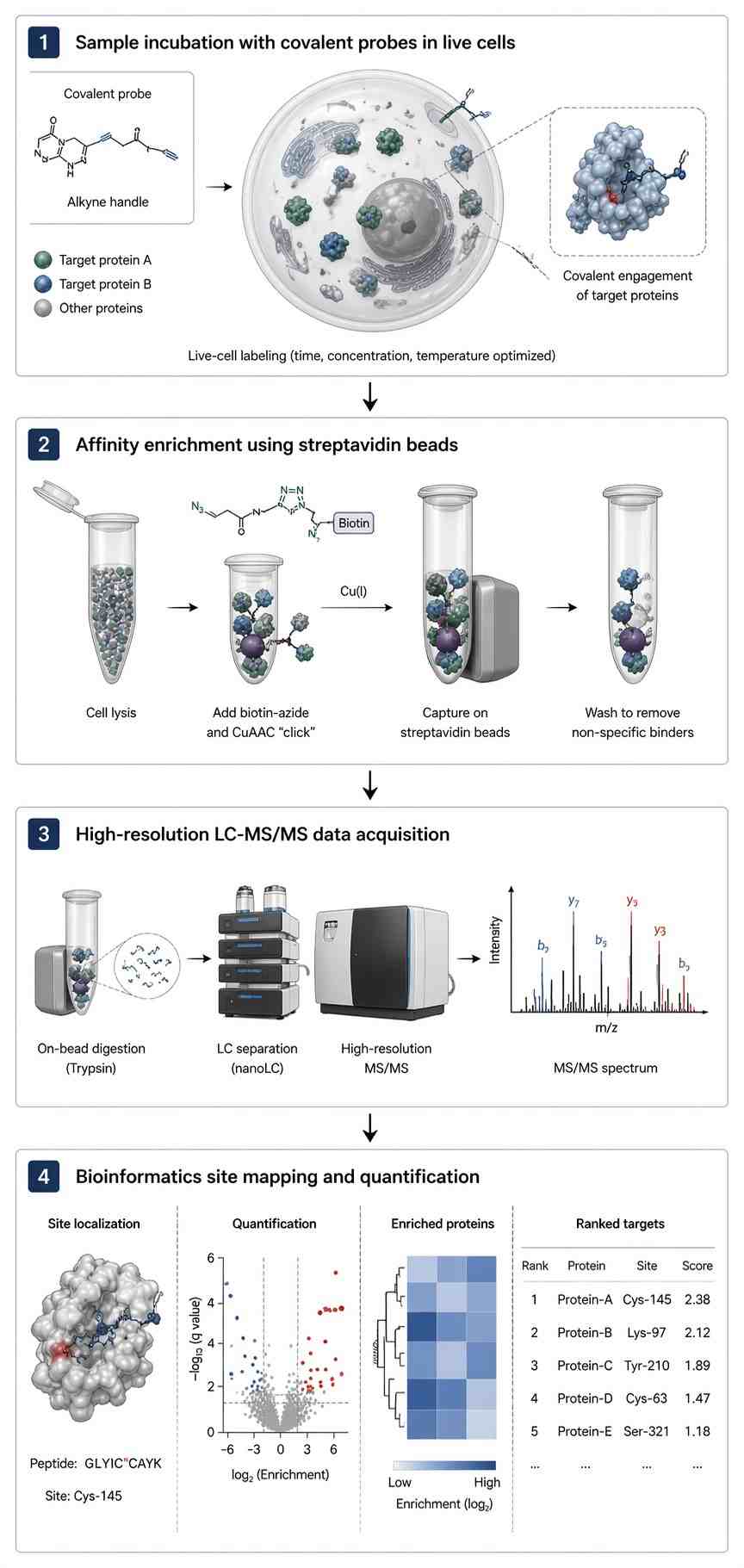
Researchers developed a platform utilizing a specific pyridinium-based probe for the selective modification of lysine residues. The workflow involved incubating the covalent probe directly in live cells, cell lysis, streptavidin enrichment, and rigorous on-bead tryptic digestion. Peptides were analyzed using high-resolution LC-MS/MS with heavily customized bioinformatic pipelines.
Results
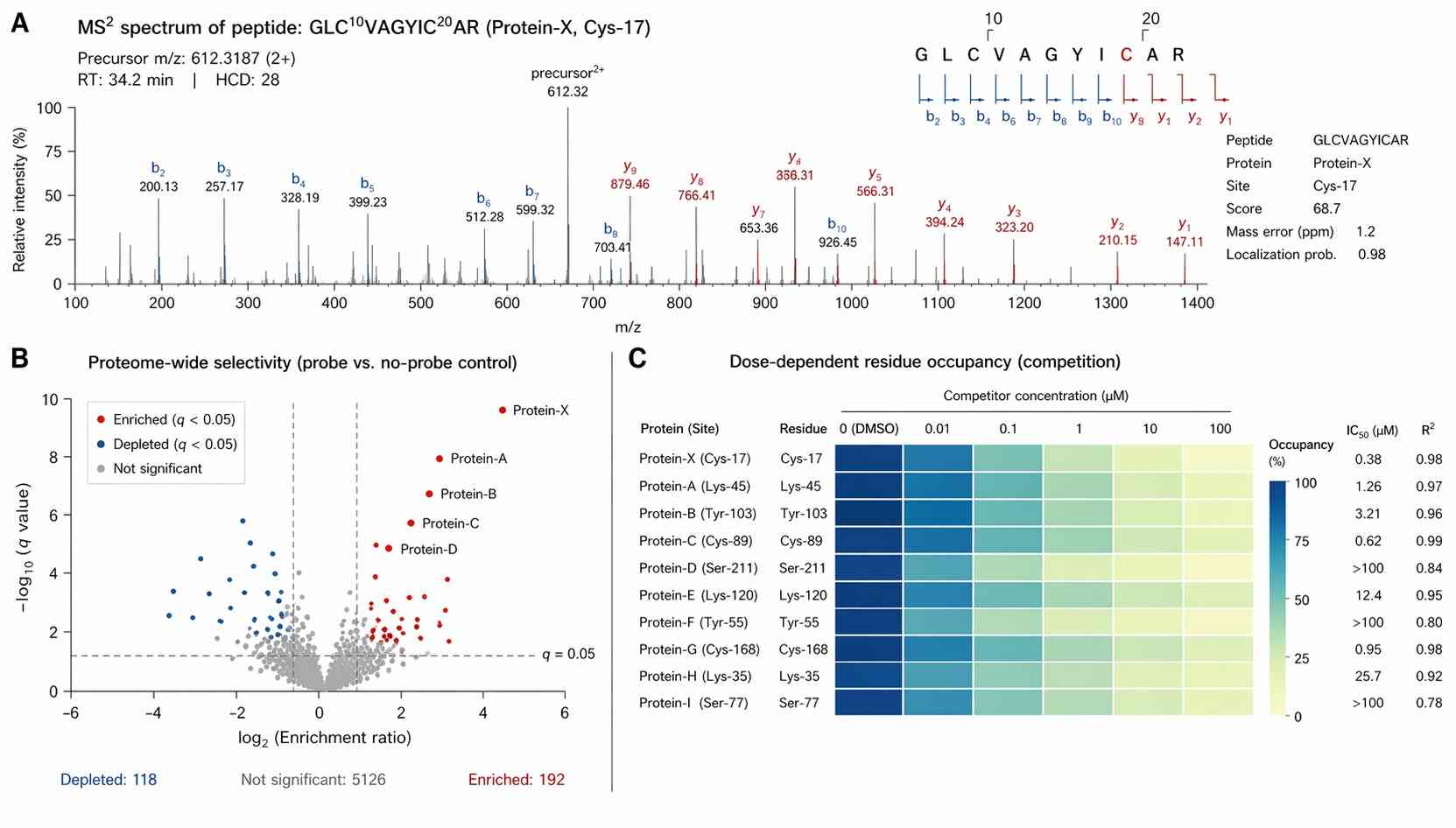
As demonstrated in Figure 1 of the referenced study, the methodology successfully mapped hundreds of highly reactive, ligandable lysine sites across the human proteome directly in a live-cell environment. The MS2 spectra provided clear, unambiguous b and y ion series flanking the modified residues.
Conclusion
This application definitively proves that with the right combination of probe chemistry, tailored enrichment buffers, and precise HRMS parameters, it is entirely feasible to achieve deep, site-specific mapping of challenging non-Cys residues.