Patch-Clamp MS Coupling for Ion Channel Drug Binding Analysis
Integrating gold-standard electrophysiology with high-resolution mass spectrometry for simultaneous functional and molecular drug-target engagement evidence.
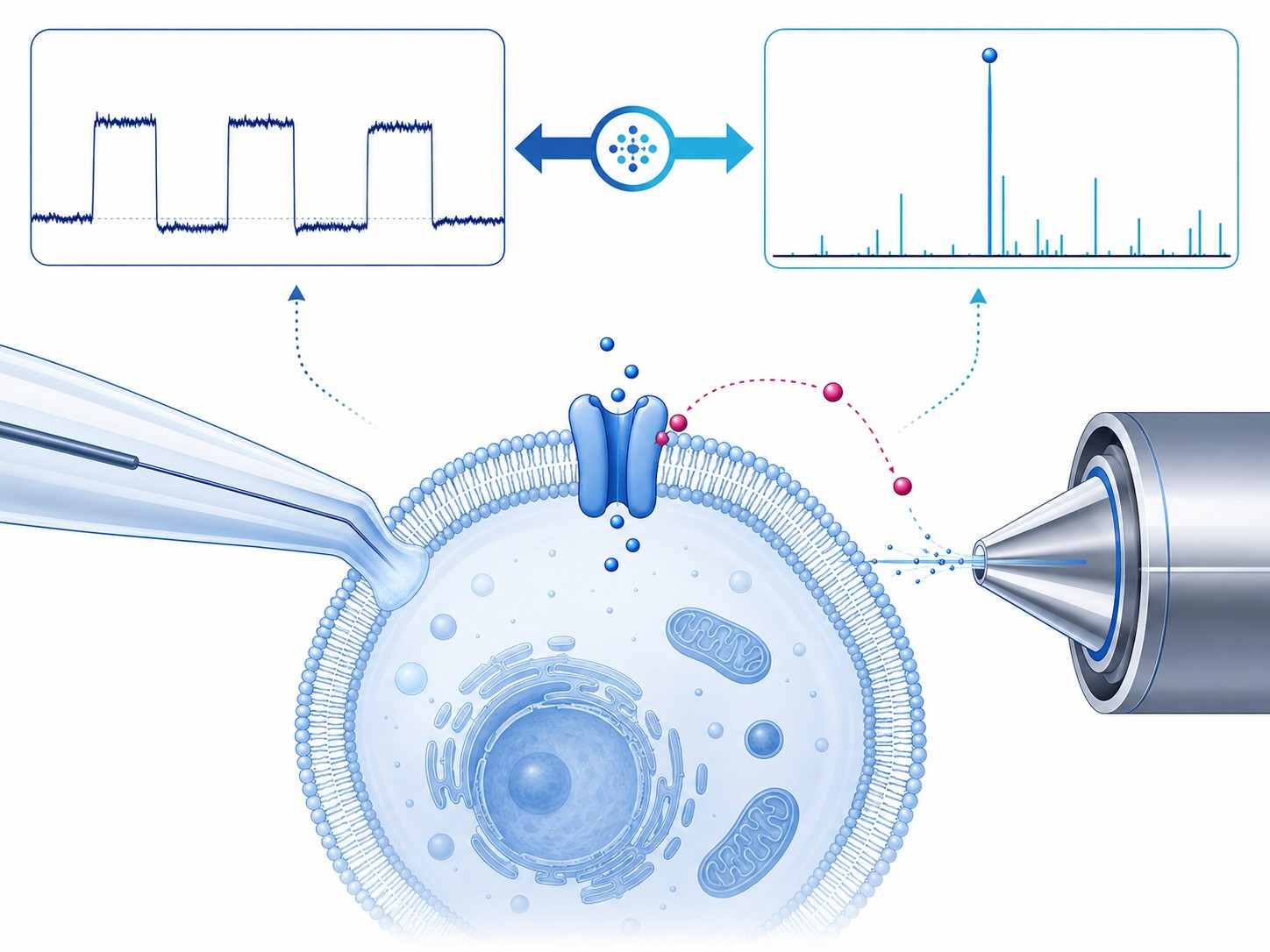
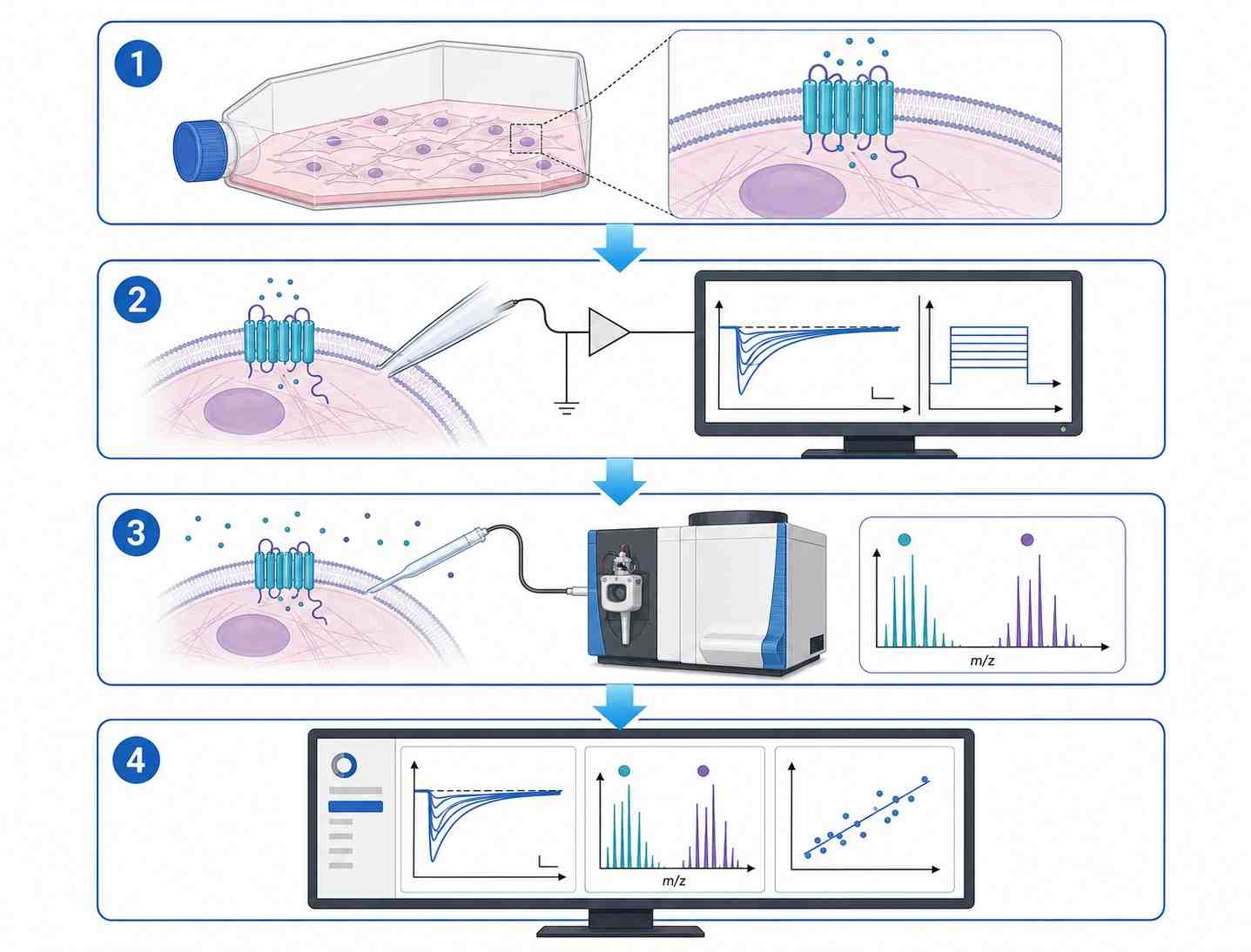
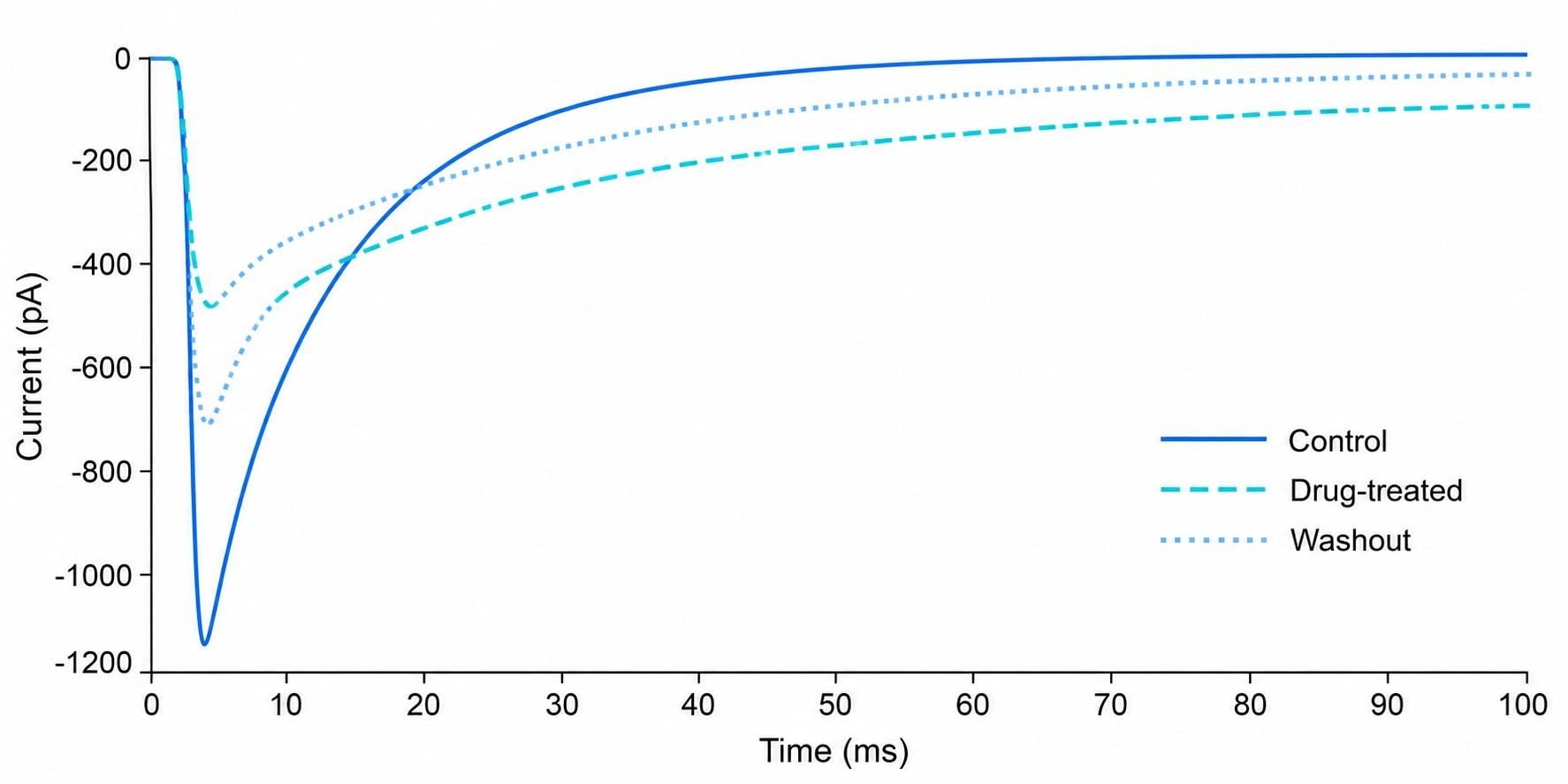
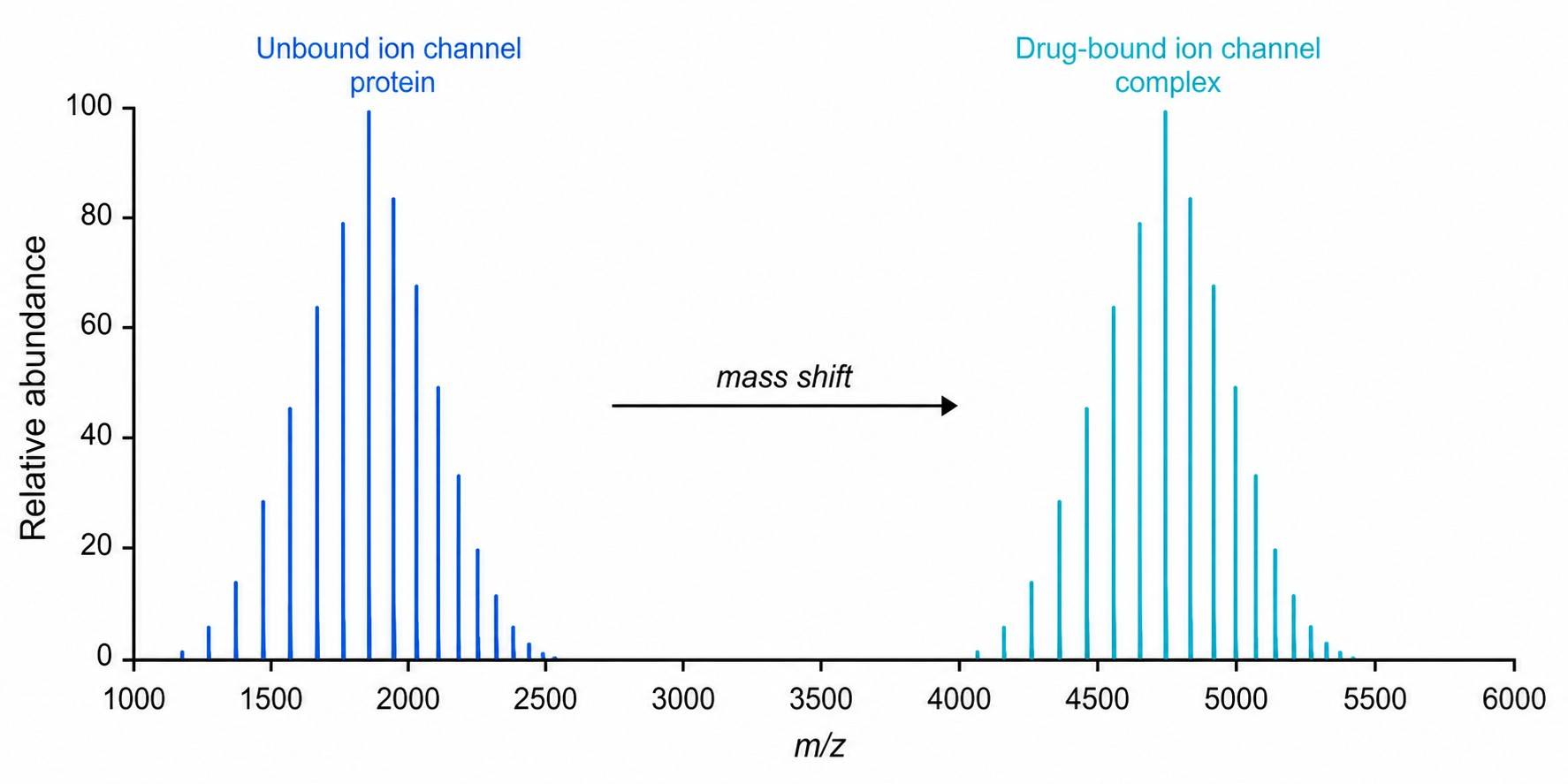
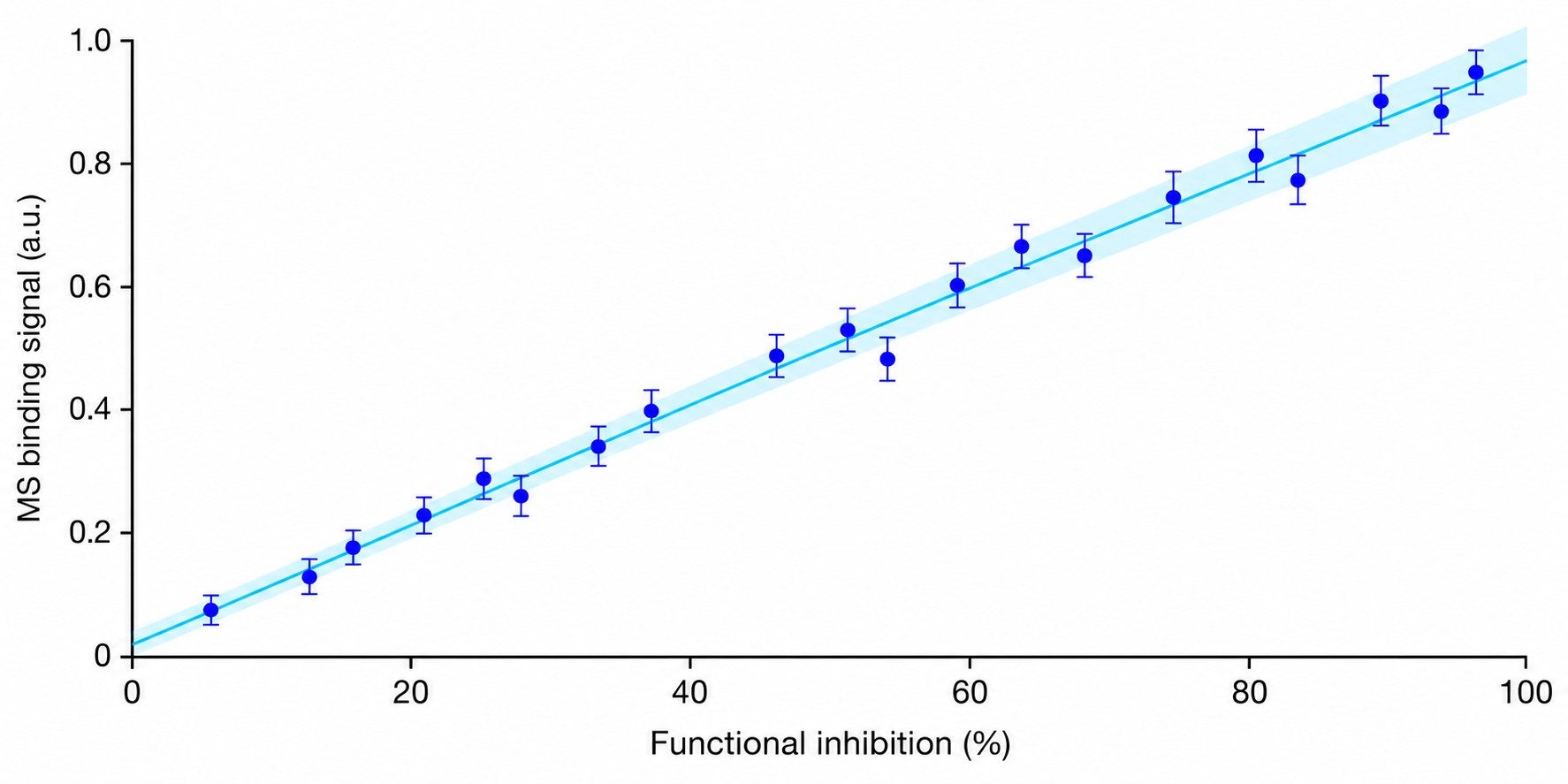
Patch-clamp MS coupling combines whole-cell patch-clamp electrophysiology with mass spectrometry-based detection to deliver concurrent functional activity and direct drug binding confirmation for ion channel targets. This integrated approach addresses a critical gap in ion channel drug discovery: traditional patch-clamp provides exquisite functional data but no molecular binding evidence, while standard binding assays lack functional context. By correlating channel activity with drug binding stoichiometry and chemical identity in a single experiment, researchers can make more informed decisions during lead optimization and candidate selection.
At Creative Proteomics, our Patch-Clamp MS Coupling service is designed to support ion channel drug discovery programs across pharmaceutical, biotech, and academic research settings. We provide end-to-end support from assay design through data integration, delivering orthogonal evidence that accelerates development timelines and reduces late-stage attrition.
Key Advantages:
- Simultaneous electrophysiology and MS readout from the same cellular preparation.
- Direct drug binding confirmation in native cell membrane environments.
- Reduced false positives from functional-only screening approaches.
- Orthogonal evidence for medicinal chemistry optimization.
- Compatible with voltage-gated, ligand-gated, and other ion channel classes.
- Custom assay design tailored to your specific ion channel target.