Enzyme Activity & Reaction Mechanism by Mass Spectrometry
Label-free, direct detection of substrates, products, and intermediates — kinetic characterization and mechanistic elucidation for any enzyme class using native substrates.
Mass spectrometry-based enzyme activity analysis offers a fundamental advantage over fluorescence, absorbance, or radiometric assays: it detects the actual substrate and product molecules by their mass-to-charge ratio. There is no need for fluorogenic substrate analogues, radiolabeled tracers, or coupled enzyme systems. The result is direct, unambiguous measurement of enzyme turnover — in native conditions, with native substrates, at native concentrations.
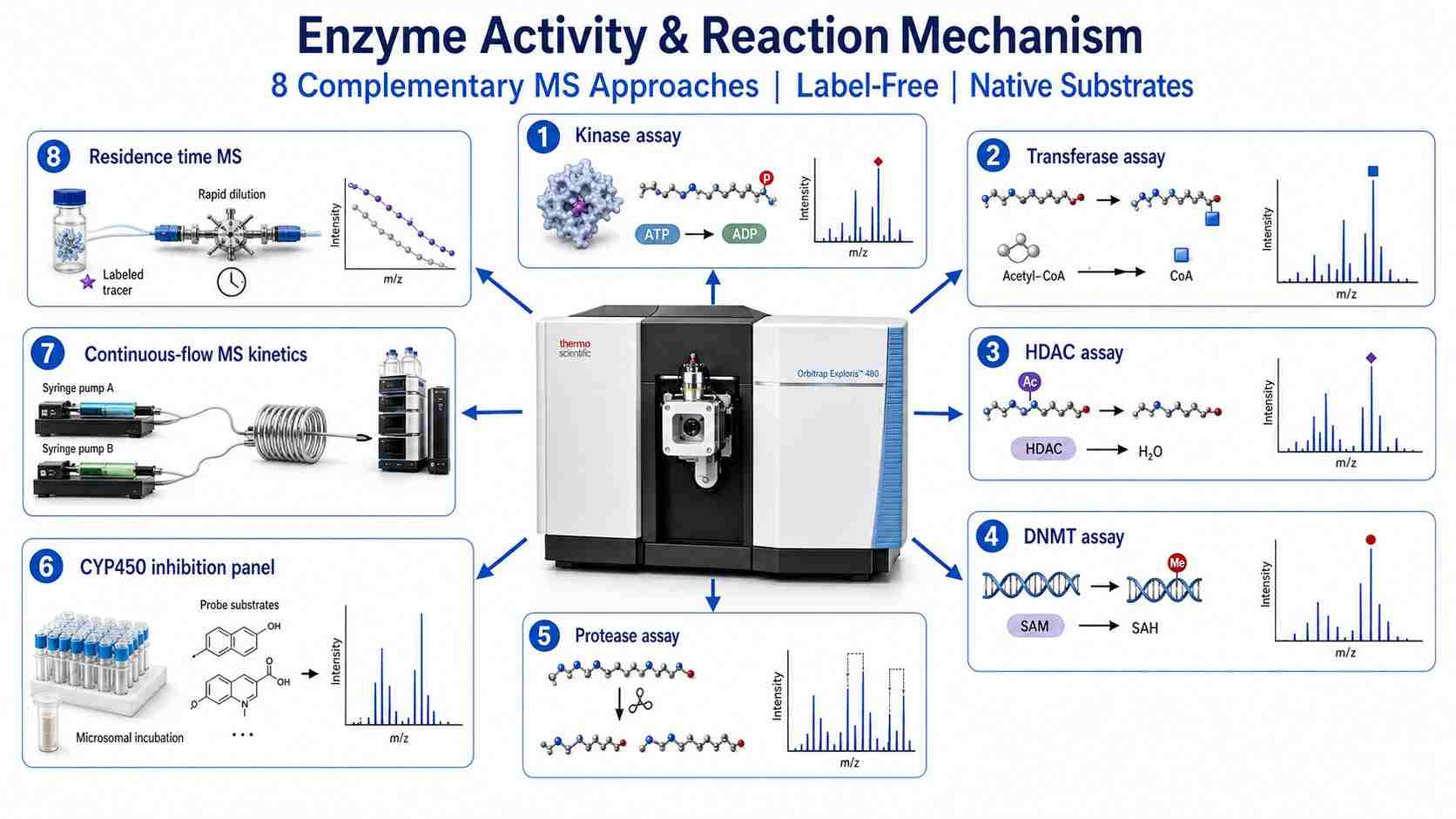
Creative Proteomics' MassTarget™ platform deploys a comprehensive suite of MS-based enzyme activity and mechanism services, covering eight complementary approaches from single-point activity screening to real-time continuous-flow kinetics and drug-target residence time determination. Whether you need to characterize a single enzyme target, profile inhibitor selectivity across a panel of isoforms, or elucidate a multi-step reaction mechanism, we provide a tailored MS strategy supported by rigorous data analysis and regulatory-compliant documentation.
Our MS-Based Enzyme Activity Platform:
- 8 complementary MS approaches covering the full spectrum of enzyme activity analysis
- Native substrate detection — any ionizable molecule, no labeling required
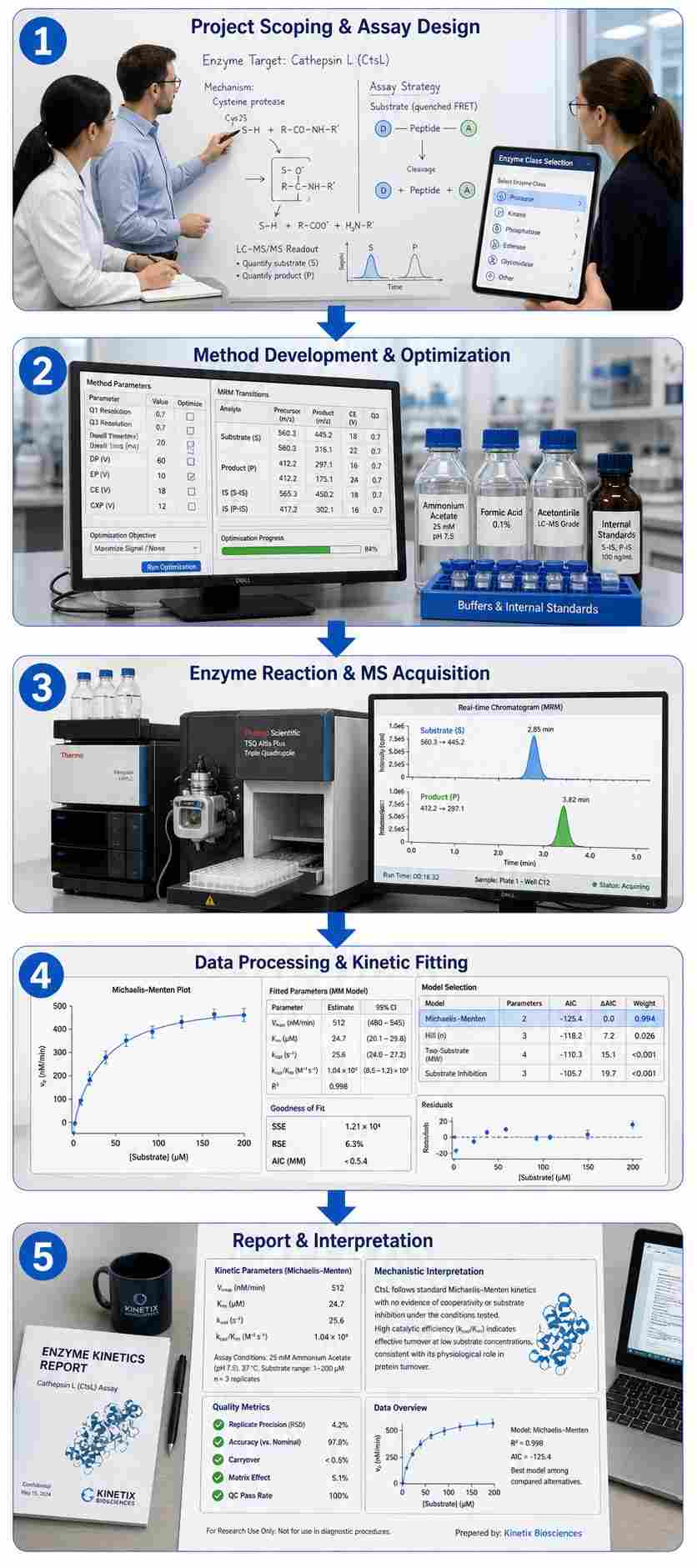
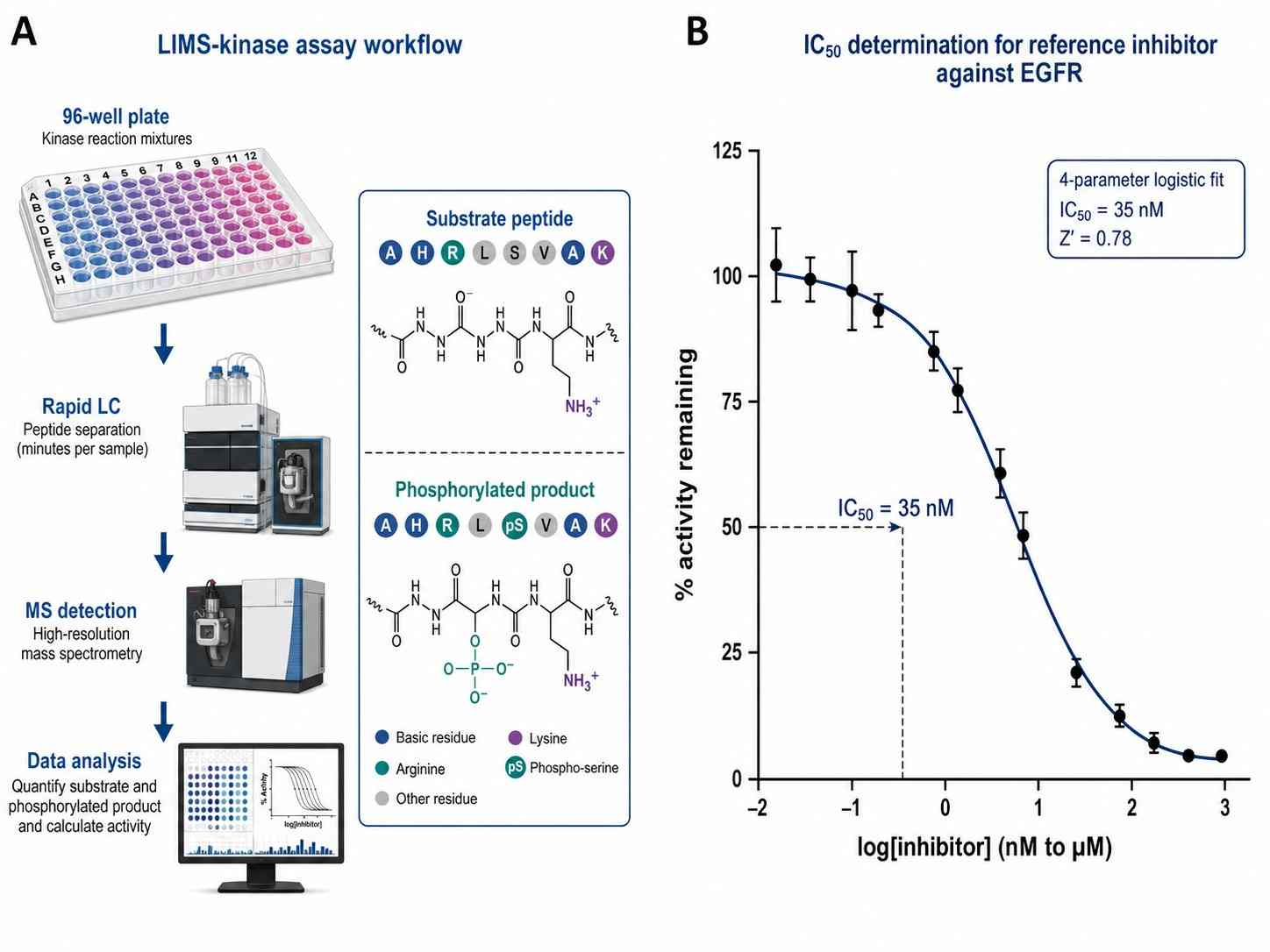
- From single-point screening to full kinetic characterization (Km, Vmax, kcat, IC50, Ki)
- FDA/ICH-compliant documentation for regulated drug metabolism studies
- Microgram-scale enzyme consumption — conserve precious reagents
- Rapid turnaround — assay development in days, not weeks