Photo-Proximity Labeling Service: Spatial Proteomics with Light-Activated Precision
Some questions can only be answered in space and time. Capture them with light.
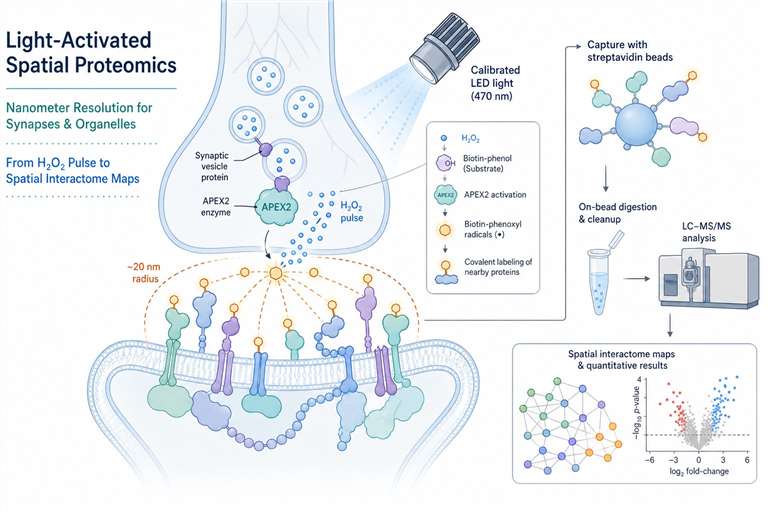
Our photo-proximity labeling service uses APEX/APEX2 enzymes for sub-second, nanometer-resolution spatial proteomics—ideal for synapses, organelle contact sites, and GPCR microdomains.
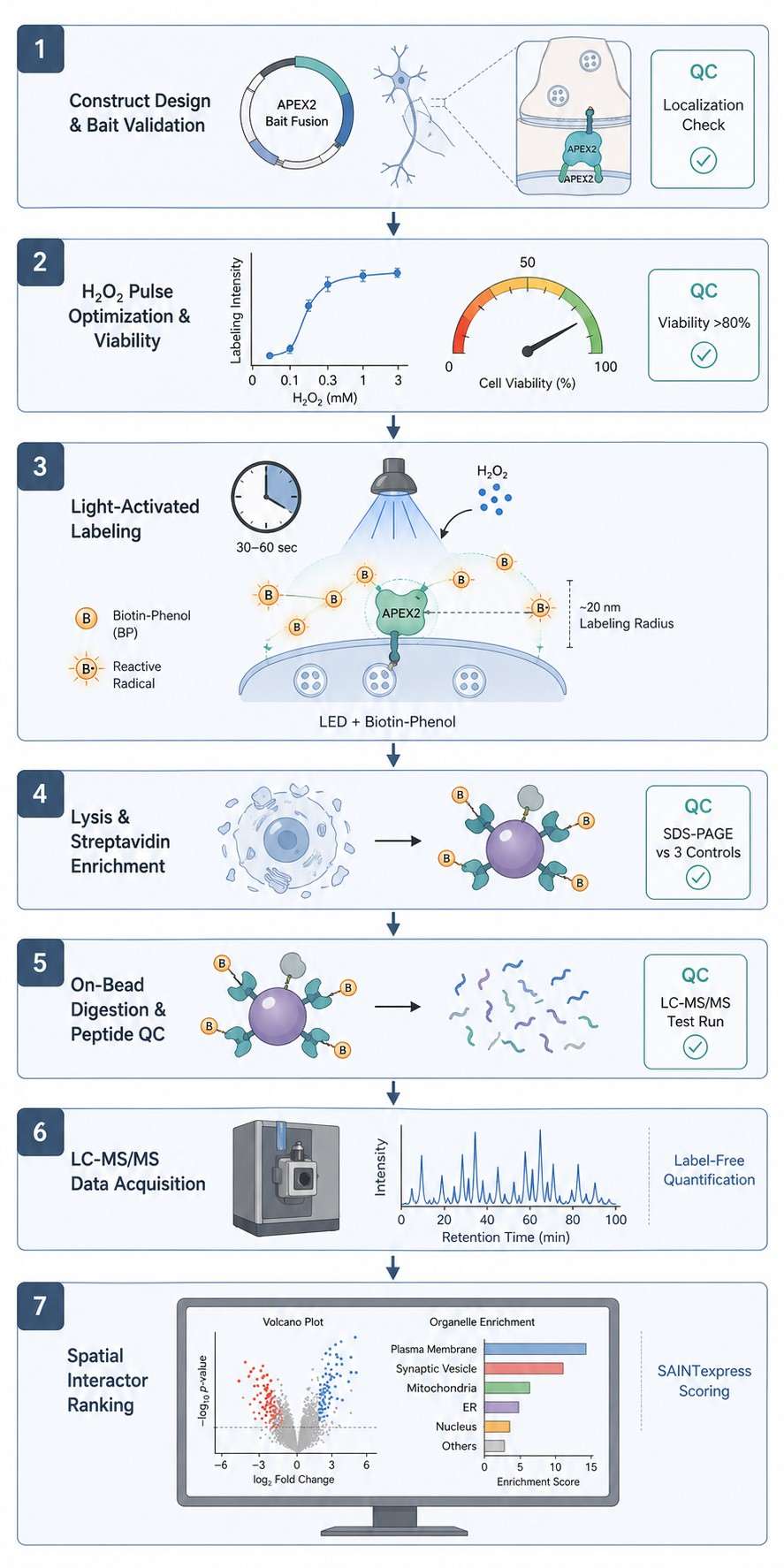
Integrated light activation, H₂O₂ pulse control, viability monitoring, and high-resolution mass spectrometry deliver publication-ready spatial interactome maps from your most precious samples.
Some questions can only be answered in space and time. Which proteins sit at the synaptic cleft when a vesicle fuses? What changes at the mitochondrial outer membrane during apoptosis? How does a GPCR rewire its neighborhood in the seconds after ligand binding?
If your labeling window is hours—or even minutes—those answers blur. Photo-proximity labeling, powered by APEX and APEX2 enzymes, captures them in under a second, at nanometer resolution. We've built an integrated platform around this. Light source, H₂O₂ delivery, viability monitoring, mass spectrometry—all connected, all dialed in. What you get isn't just any list of proteins. It's a spatially resolved snapshot of your subcellular compartment, frozen at the exact moment you chose.
Key Advantages:
- Sub-second, light-activated labeling precision
- Nanometer spatial resolution for subcellular compartments
- Integrated H₂O₂ pulse control and viability QC
- Dedicated APEX/APEX2 platform and protocols