Yeast Display Ligand Screening and LC-MS Analysis Service
Target-binding peptide, scFv, antibody fragment, and engineered binder discovery with LC-MS/MS characterization.
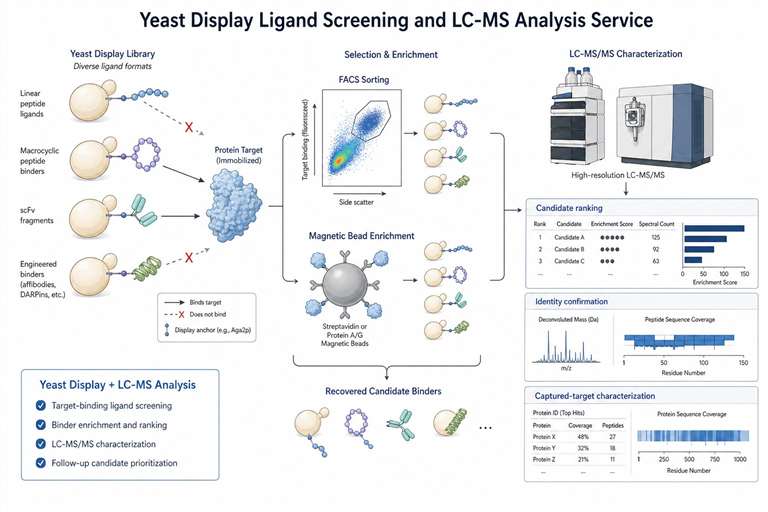
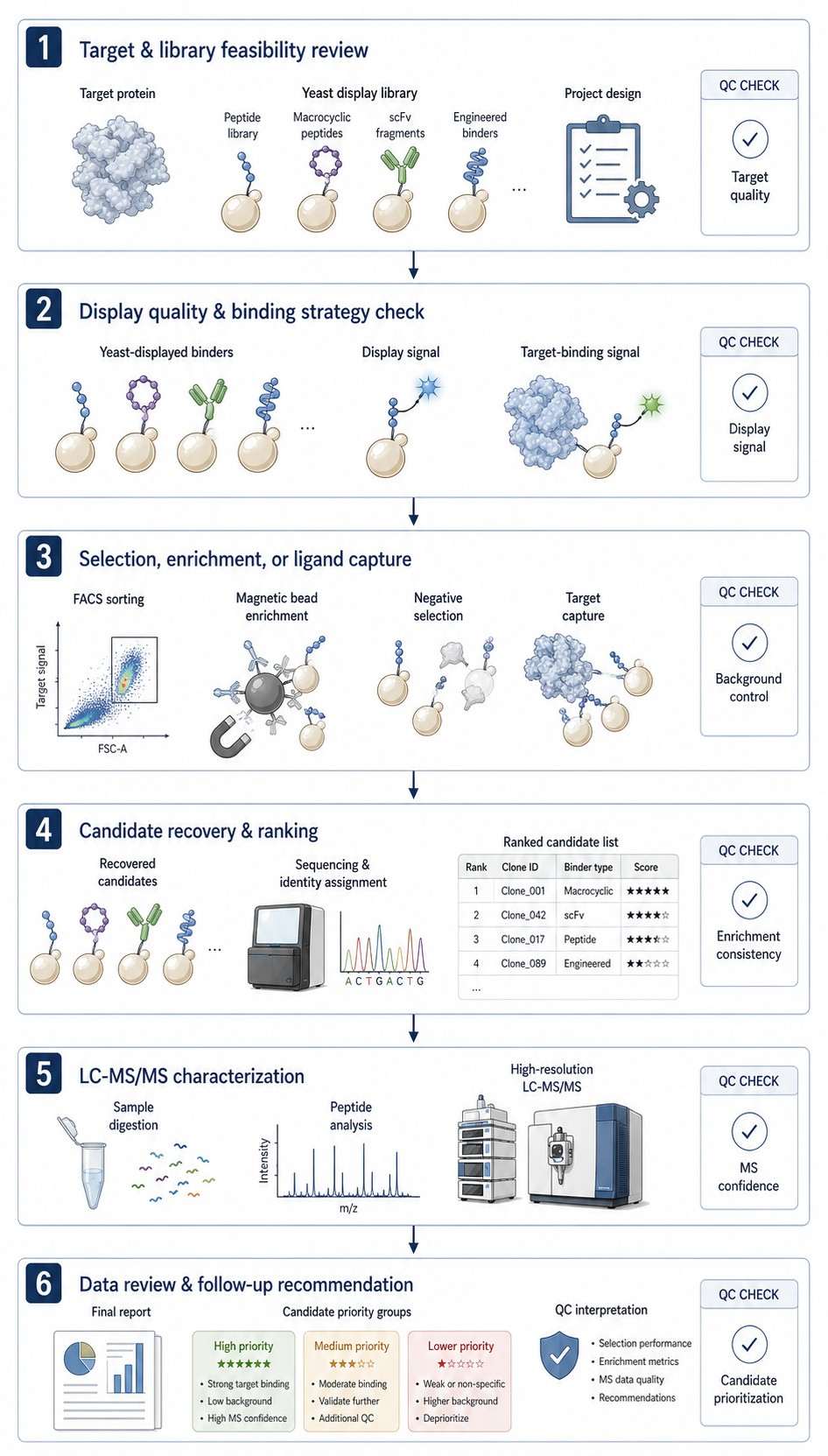
Creative Proteomics provides Yeast Display Ligand Screening and LC-MS Analysis Service for researchers who need to identify, rank, and characterize target-binding peptides, antibody fragments, or engineered binders. We help connect target and library design, display-based enrichment, LC-MS/MS characterization, and candidate prioritization in one practical workflow.
Our service is built for teams working on peptide ligand discovery, macrocyclic peptide screening, scFv or antibody fragment selection, captured-target identification, and protein-ligand interaction research. From the first project discussion to the final data package, we focus on clear controls, interpretable results, and follow-up-ready candidate lists.
Service strengths:
- Screen target-binding ligands and binders.
- Support peptide, scFv, and engineered libraries.
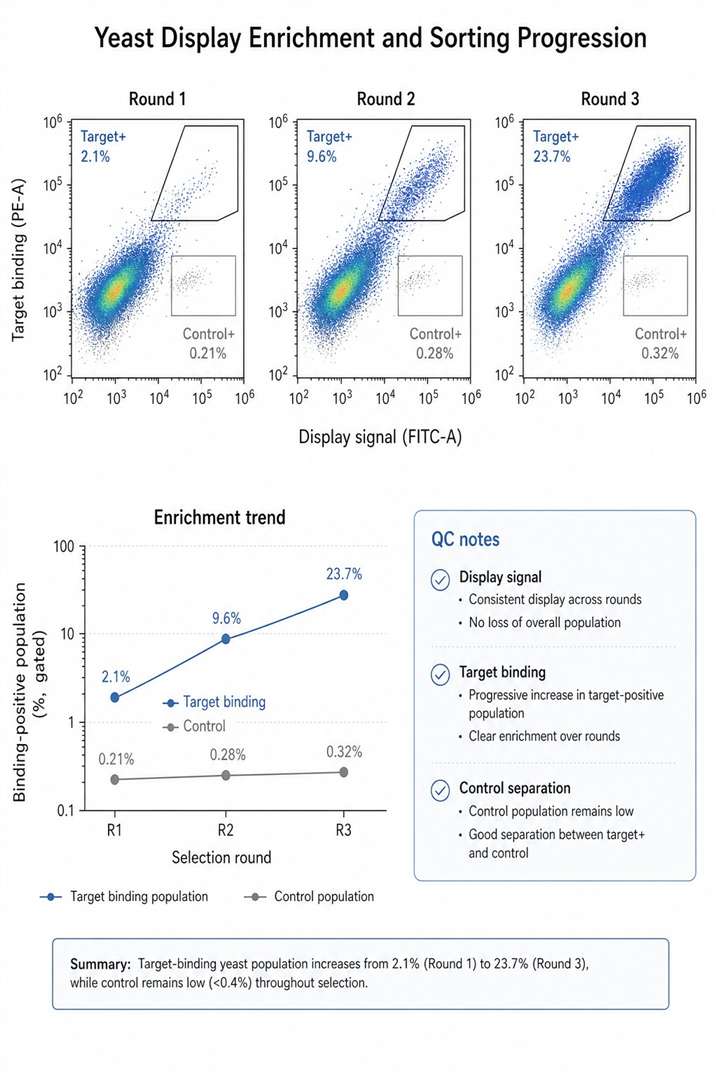
- Combine display selection with LC-MS/MS analysis.
- Evaluate target, library, controls, and capture design.
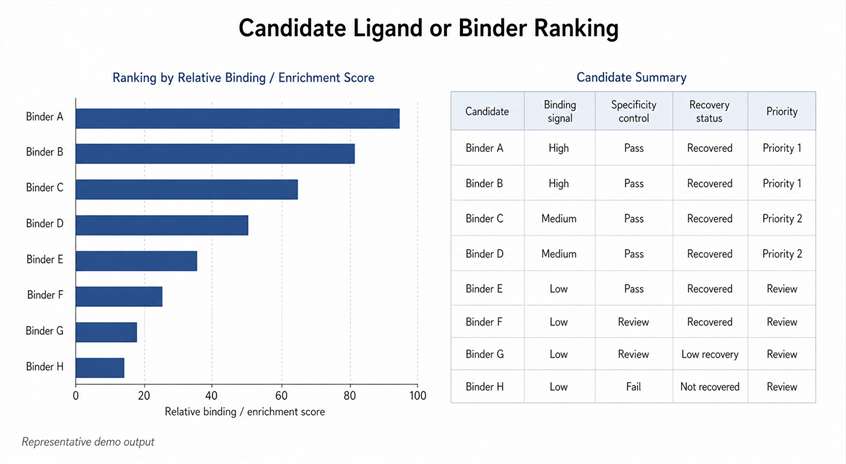
- Deliver ranked candidates and characterization reports.