AirID In Vivo Proximity Labeling CRO Service
Cleaner in vivo interactomes start with lower background. AirID delivers them.
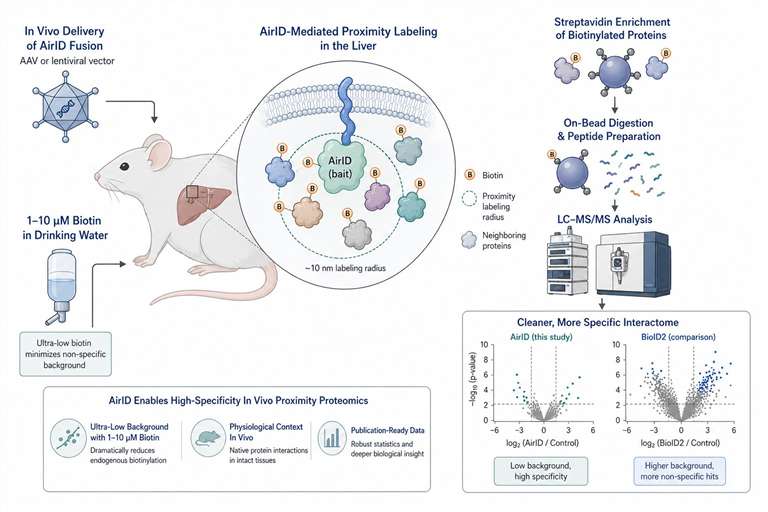
Our AirID proximity labeling service provides ultra-low-background proteomics optimized for animal models—from viral delivery to publication-ready interaction networks, with less noise and fewer false positives.
You've run the BioID2 experiment in vivo. You've validated the hits. And too many turned out to be noise—endogenous biotinylated proteins, metabolic artifacts, background that built up over long labeling windows. The real interactors are in there somewhere, buried under false positives.
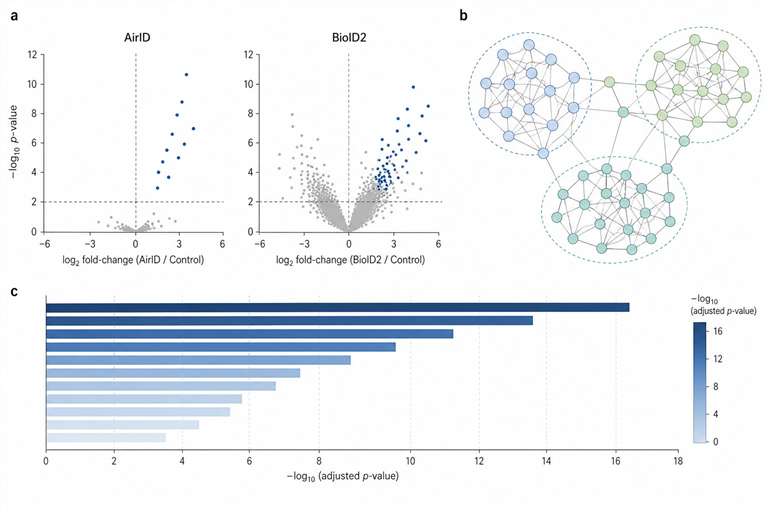
AirID changes that. It's an engineered biotin ligase that produces dramatically less background than BioID2, works with far lower biotin concentrations, and labels for up to 48 hours without accumulating a haze of non-specific signal. For in vivo work—where you can't just wash away background and you can't risk losing your animal model to toxicity—that's the difference between a candidate list you trust and one you chase.
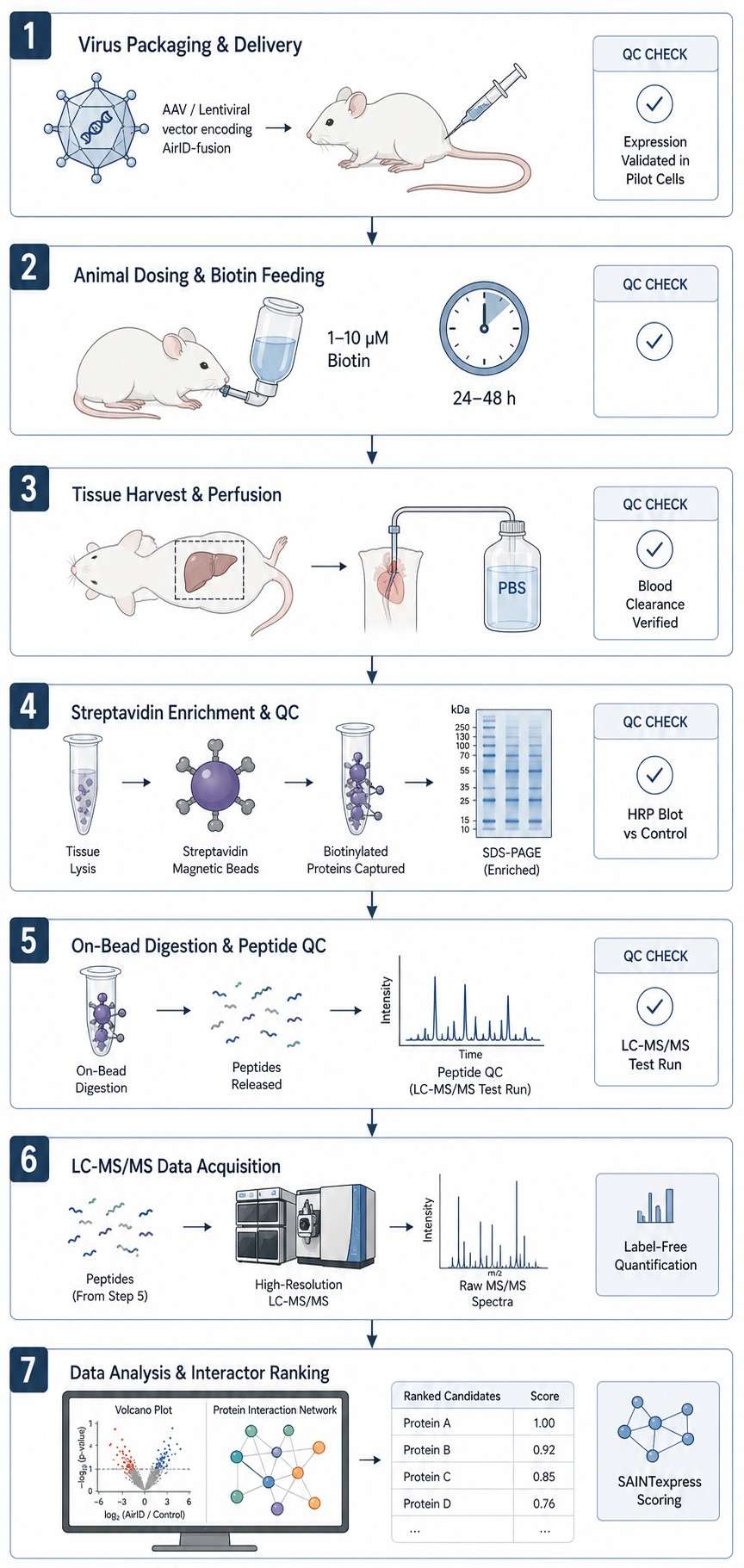
We've built a complete in vivo AirID workflow around this advantage. Viral packaging. Animal dosing. Tissue harvest. Mass spectrometry. All connected, all optimized for low background from the start. What you get is a cleaner shortlist of interactors you can take straight into validation.
Key Advantages:
- Ultra-low background, ideal for in vivo
- Full workflow: virus, animal, tissue, MS
- Optimized for low biotin concentrations
- Publication-ready, traceable data