LC-HRMS/MS Dereplication Service for Natural Product Identification
Accelerate natural product discovery by rapidly identifying known compounds in complex extracts.
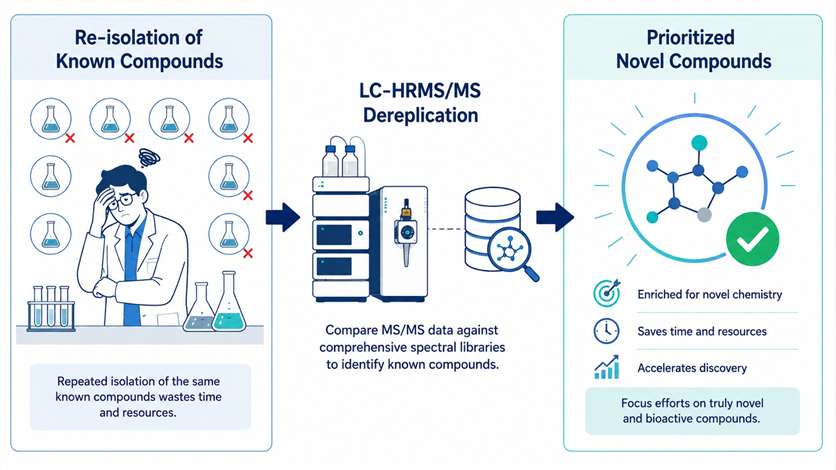
LC-HRMS/MS dereplication is a targeted analytical strategy that identifies known natural products in complex extracts by matching high-resolution mass spectrometry data against comprehensive compound databases. For natural product chemists and drug discovery teams, this service eliminates the costly and time-consuming re-isolation of previously characterized molecules, allowing you to focus resources on truly novel or bioactive compounds.
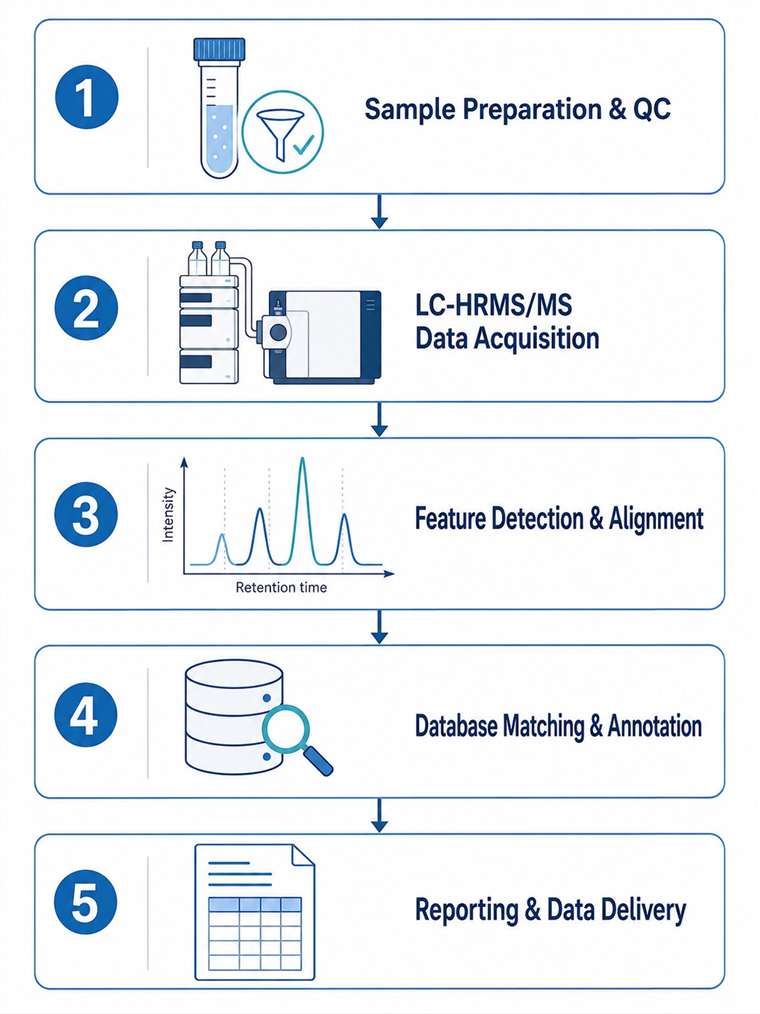
At Creative Proteomics, our LC-HRMS/MS dereplication service integrates high-resolution mass spectrometry, extensive natural product databases, and expert-led annotation workflows to deliver confident compound identifications from plant, microbial, marine, and fungal extracts.
Key Advantages:
- Comprehensive database coverage including Dictionary of Natural Products (DNP), GNPS, PubChem, and AntiBase.
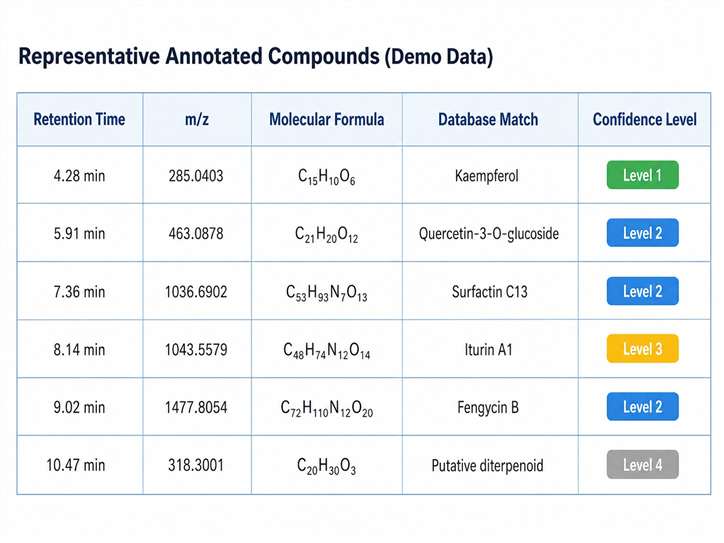
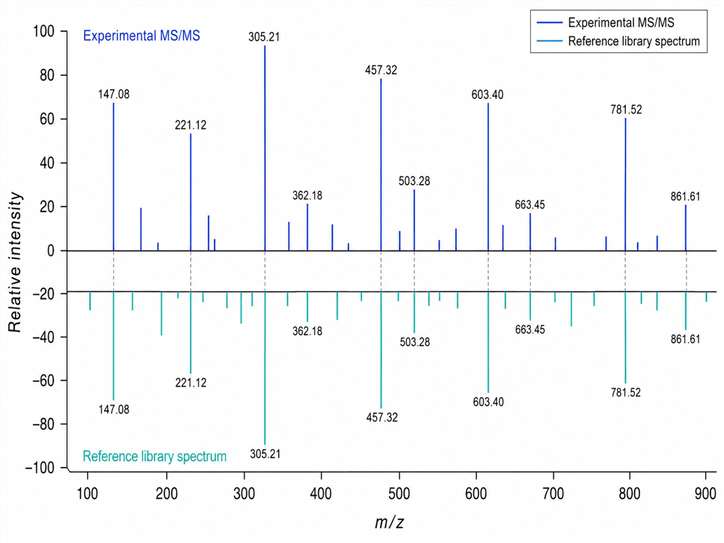
- High-confidence annotations with MS/MS spectral matching, retention time alignment, and expert manual verification.
- Fast turnaround — standard dereplication reports delivered within 2–3 weeks.
- Flexible sample acceptance — plant extracts, microbial broths, marine organisms, and purified fractions.
- Integrated molecular networking (GNPS-compatible) to visualize known vs. novel compound clusters.