This literature example summarizes the findings and conclusions from Verhelst, Bonger, and Willems' review article, Bioorthogonal Reactions in Activity-Based Protein Profiling. It is not a Creative Proteomics customer case.
Background
The review explains that activity-based protein profiling can label and detect active enzyme species in cell lysates, cells, and whole animals. It also describes how ABPP has been used to study biological processes, identify drug targets, evaluate drug selectivity, and image probe targets.
A key problem discussed in the paper is that directly attaching large detection or affinity tags to an activity-based probe can change probe behavior. Large tags may reduce cell uptake, create steric clashes with the protein of interest, or affect probe localization. The authors present bioorthogonal chemistry as a way to use smaller mini-tags, such as alkyne or azide handles, during biological treatment and then attach larger detection or enrichment tags at a later stage.
Methods
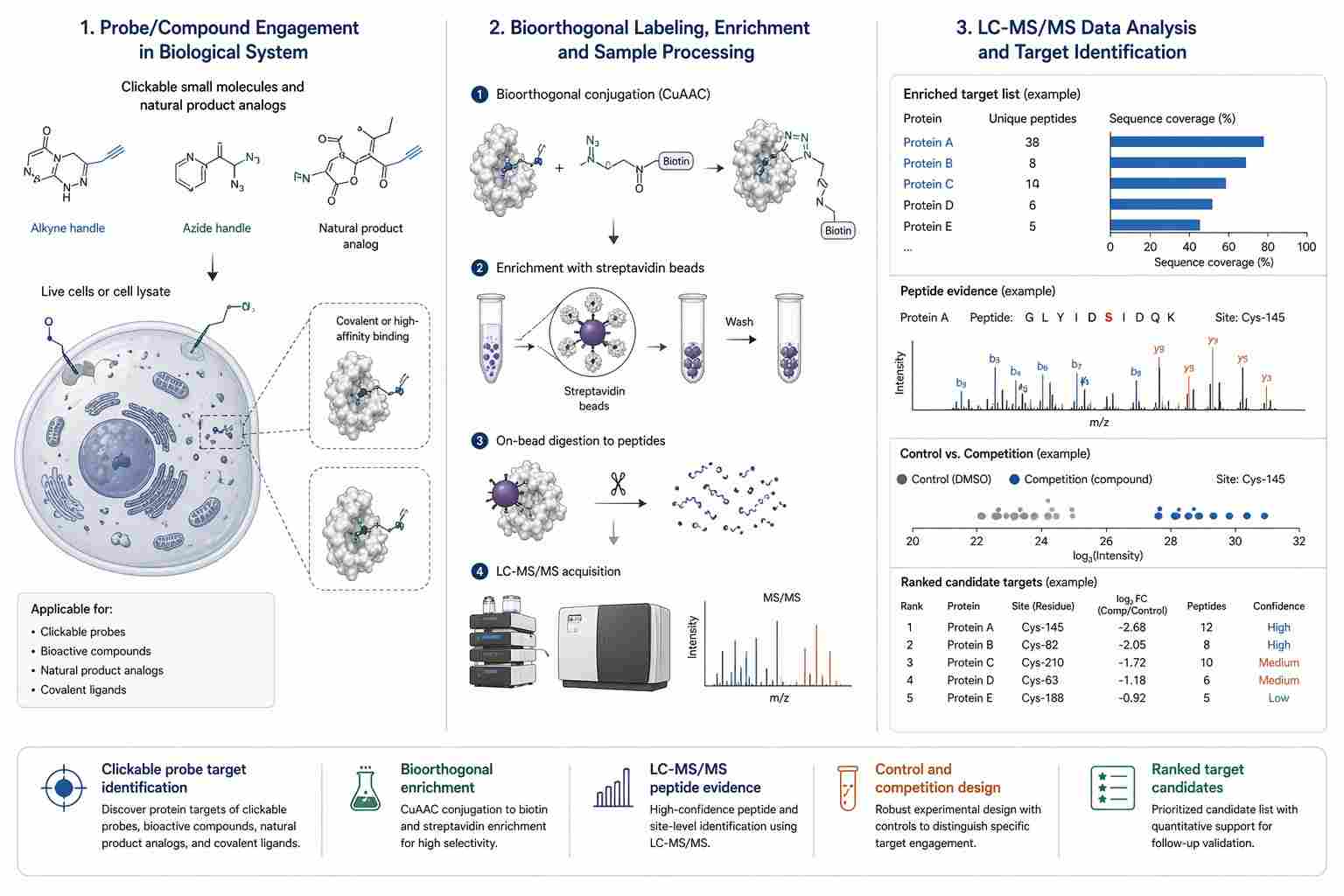
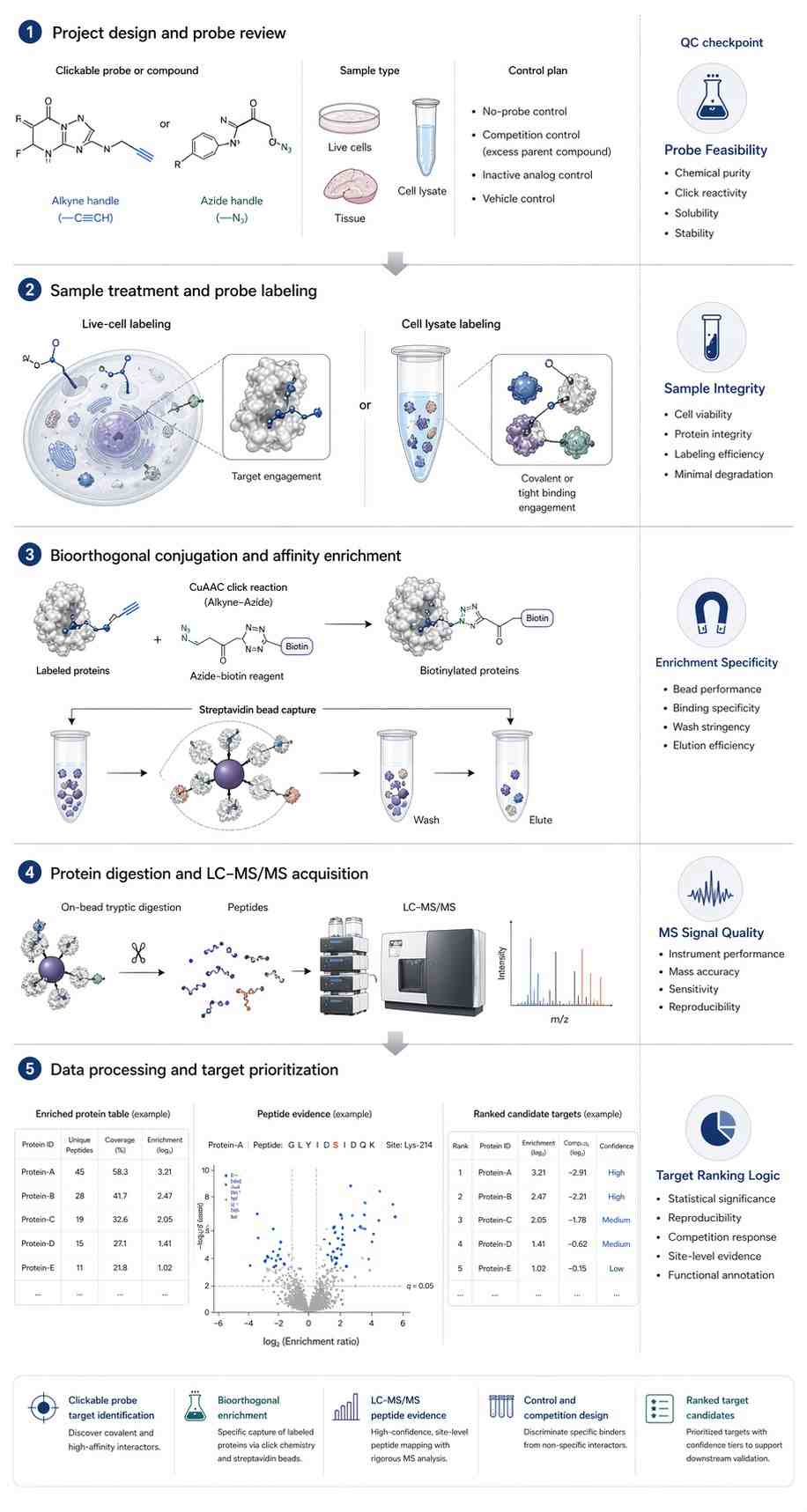
Figure 7, titled "Target identification in ABPP by making use of bioorthogonal chemistry," presents the target identification workflow.
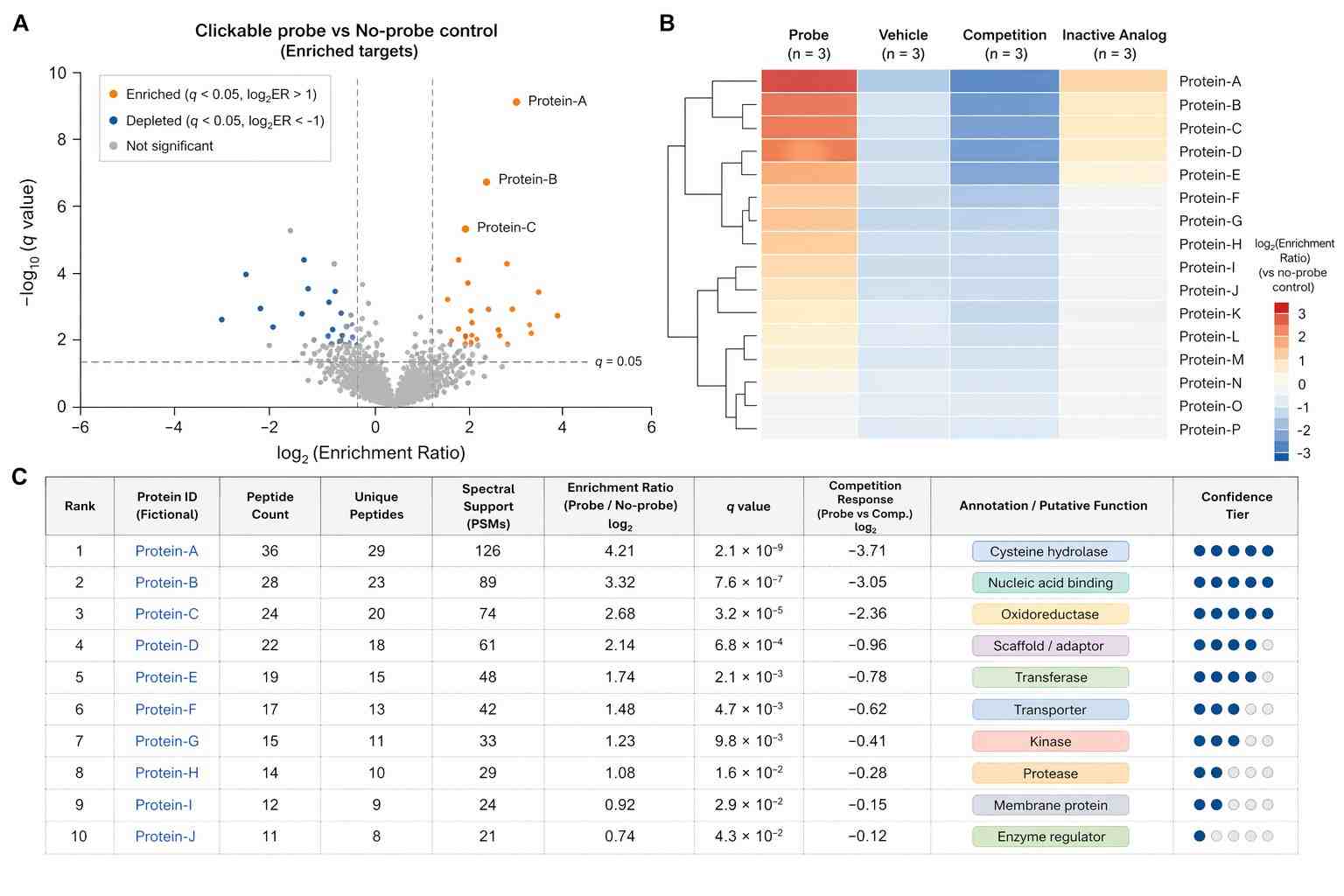
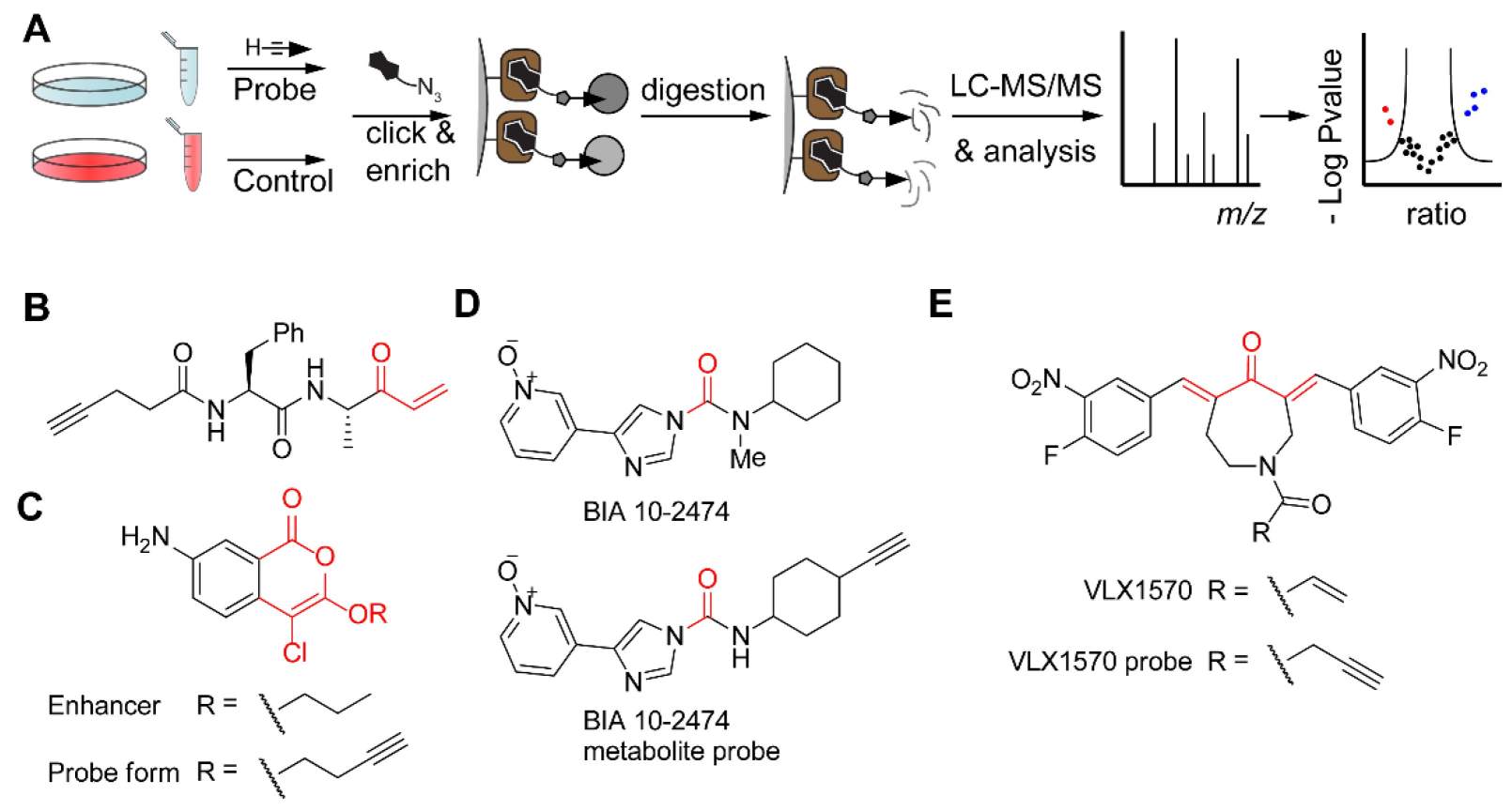
In Figure 7A, live cells or a cell lysate are treated with a clickable probe. The paper recommends a control experiment without a probe or with a competitive inhibitor. After cell lysis, CuAAC is used with a biotin-tagged reagent. Labeled proteins are enriched on immobilized streptavidin, followed by tryptic digestion. The resulting peptides are analyzed by LC-MS/MS and processed, for example, by a volcano plot.
The figure also notes that the control experiment can be isotopically labeled, such as by SILAC, or that tryptic peptides can be labeled with heavy and light isotope labels before mixing. Label-free quantification is also described as an alternative. The figure further notes that proteins may be eluted before digestion to identify modified peptides, or that modified peptides may be released after on-bead digestion by using a cleavable linker.
Results
Figure 7 does not present a new numeric experimental dataset. Instead, it reports a target-identification workflow and several concrete literature examples of probes used in ABPP target and off-target studies.
The figure includes an alkyne-tagged activity-based probe based on an inhibitor of attachment and invasion of the Toxoplasma parasite. It also shows an enhancer of Toxoplasma invasion into host cells and an alkynylated derivative. In addition, the figure includes a FAAH inhibitor, BIA 10-2474, and a probe derived from a de-methylated metabolite. It also shows the DUB inhibitor VLX1570 and an alkyne-tagged probe form.
The review explains that enrichment and tandem MS-based ABPP workflows have been used to identify targets and off-targets of drugs or drug candidates. The authors group these studies into two main strategies: identifying targets of inhibitors discovered through phenotypic screening, and identifying off-targets of drugs that already have a known target. The review also states that exact binding-site identification can provide information about an enzyme's binding pocket or mechanism of action.
Conclusion
The authors conclude that bioorthogonal chemistry has been advantageous in ABPP because it allows late-stage introduction of experimental readout tags. This helps avoid problems that may occur when large tags are installed directly onto a probe, such as reduced cell uptake, steric clashes, or mislocalization.
The review also concludes that a two-step labeling approach gives researchers flexibility. A biological sample treated with an activity-based probe can be divided into aliquots, with one portion labeled for visualization and another portion conjugated to an affinity tag for target protein isolation.
The authors further state that, despite major advances, new bioorthogonal reactions are still needed to improve reaction kinetics, compatibility, and future applications. Their overall conclusion is that ABPP will continue to benefit from future developments in bioorthogonal chemistry.
For our clients, this literature example shows why probe design, late-stage enrichment tagging, control conditions, LC-MS/MS quality, and target ranking all need to be considered together when planning a bioorthogonal labeling-based target identification project.