Live-Cell MS Profiling Service — Cellular Drug Target Engagement & Multi-Omics Analysis
Characterize drug-target interactions and cellular responses directly in living cells with integrated mass spectrometry-based proteomics, metabolomics, and lipidomics — from target engagement to multi-omics phenotypic profiling.
You've identified a promising compound. But does it actually engage its intended target inside living cells? And what else happens in the cell as a result? Traditional biochemical and lysate-based assays, while valuable for initial screening, cannot answer these questions — they miss the complexity of drug-target interactions in their native cellular environment, where protein complexes, post-translational modifications, subcellular compartmentalization, and metabolic context all influence pharmacology. Without direct evidence of drug binding and cellular response in living cells, researchers risk advancing compounds with poor target engagement, unanticipated off-target effects, or inadequate intracellular penetration.
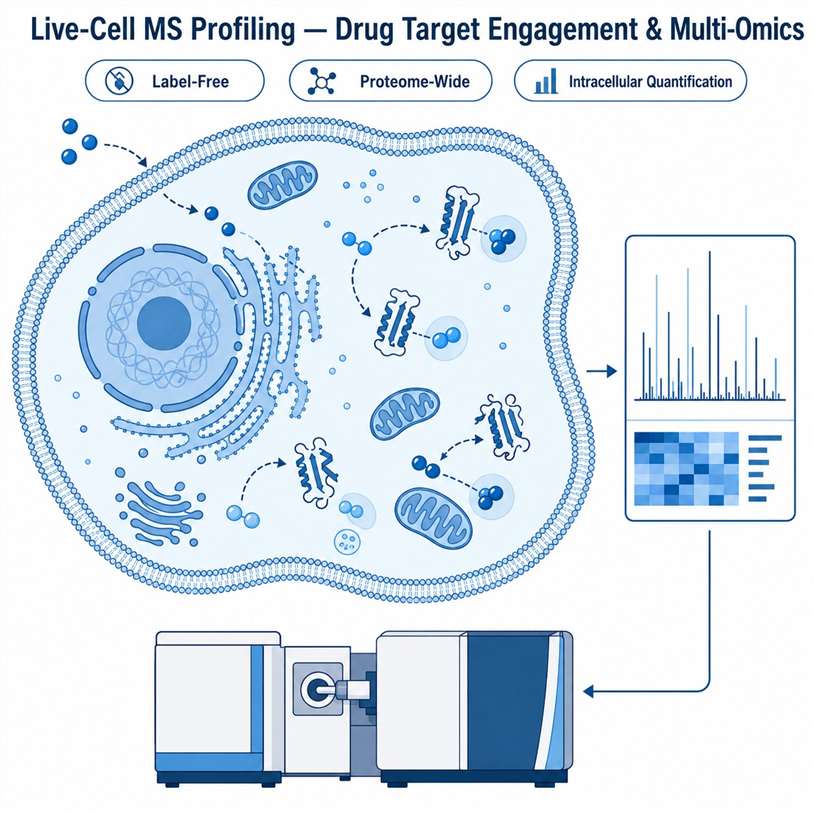
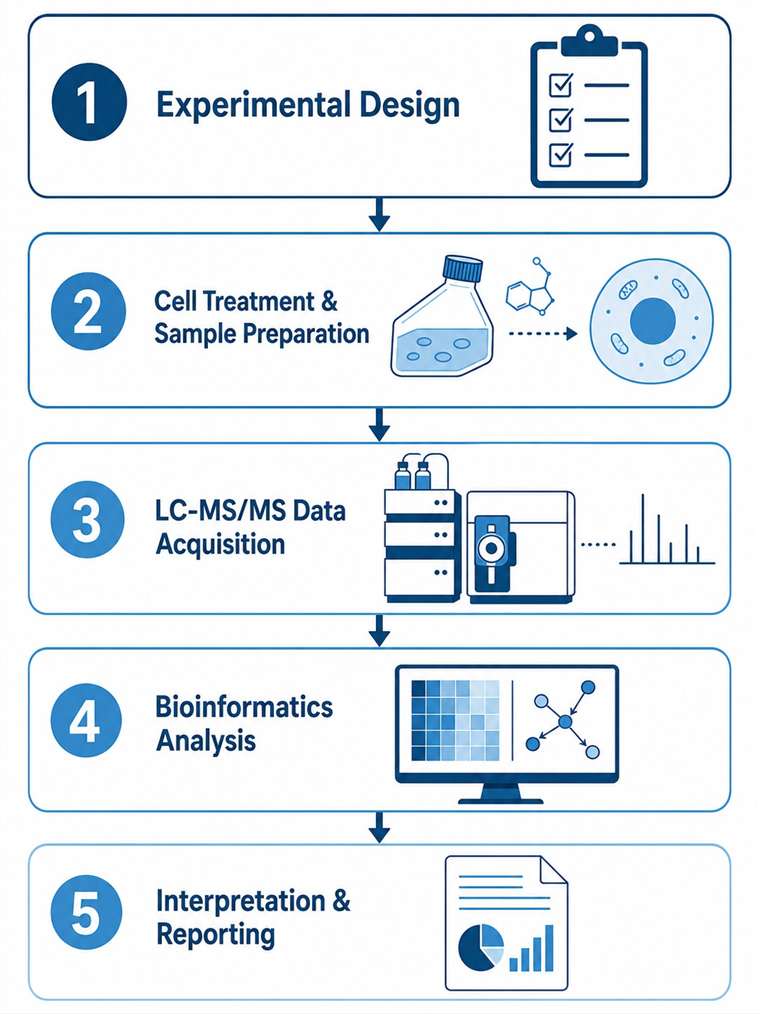
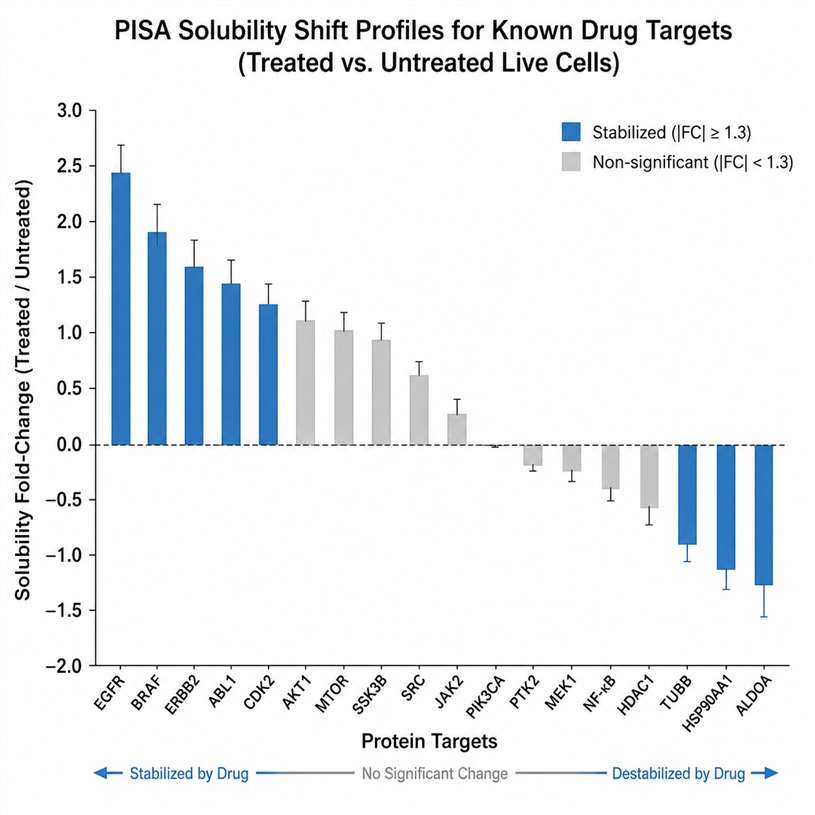
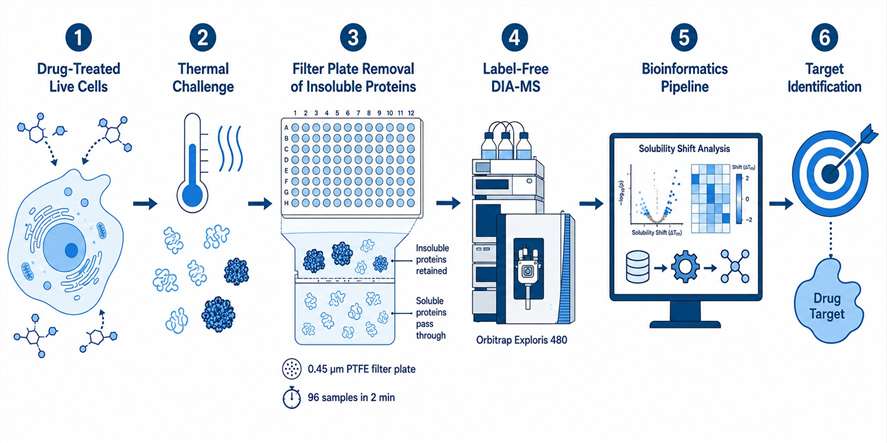
MassTarget™ offers a comprehensive Live-Cell MS Profiling Service that combines multiple mass spectrometry-based modalities — including thermal stabilization assay (PISA-based), Proteome Integral Solubility Alteration (PISA), intracellular drug quantification by LC-MS/MS, and cellular multi-omics profiling — to provide an integrated view of drug behavior directly in live cells. Our service covers the full spectrum from target engagement validation to proteome-wide target deconvolution, intracellular drug concentration measurement, and downstream cellular response profiling through proteomics, metabolomics, and lipidomics.
By consolidating these complementary capabilities under one roof, we eliminate the need to coordinate across multiple vendors and provide a single, integrated workflow for live-cell drug profiling — enabling you to make confident, data-driven decisions earlier in the drug discovery process.