Background
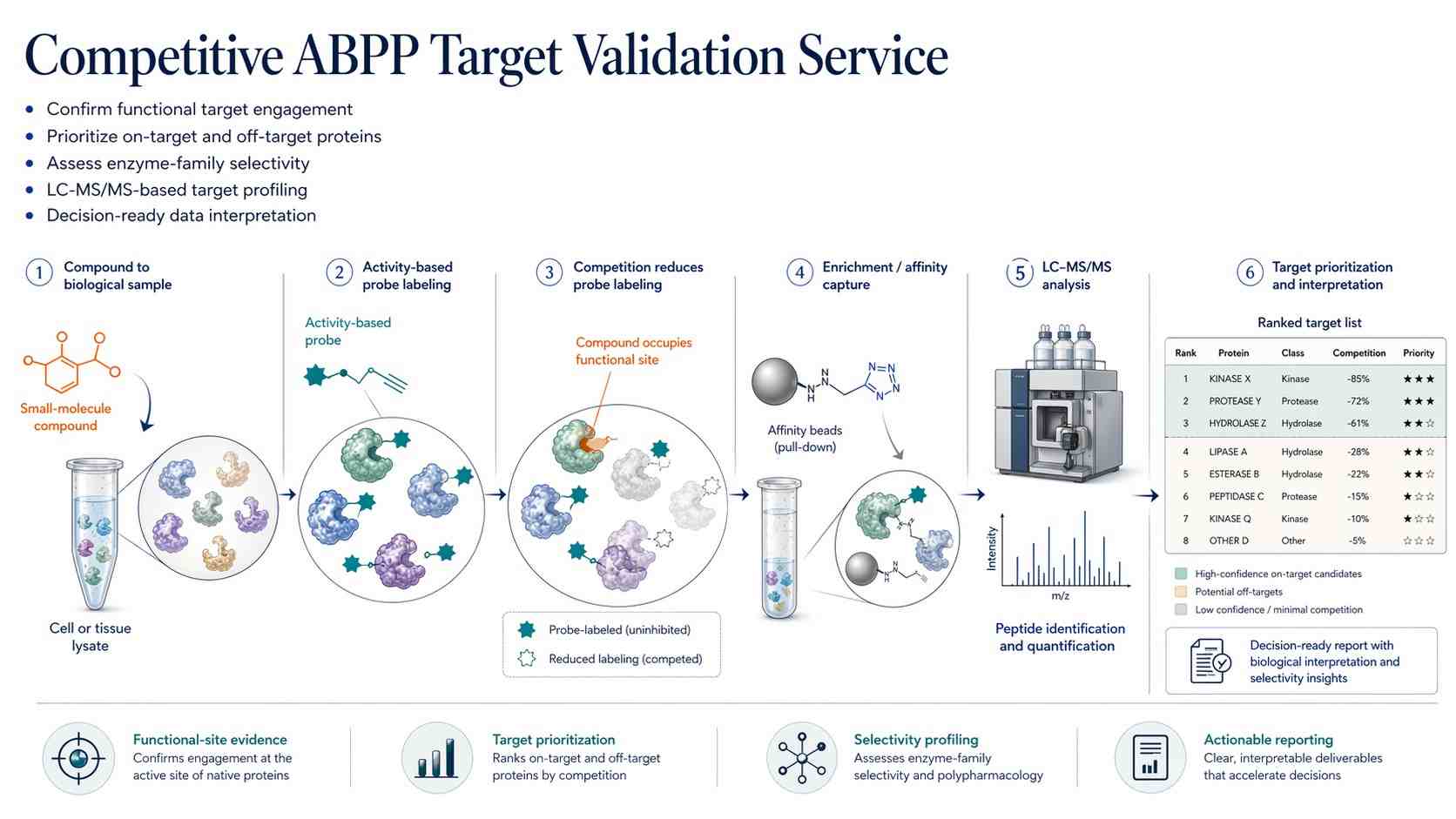
Target identification can become difficult when a small molecule affects several proteins in a complex proteome. In a published Cell Chemical Biology study, researchers used competitive activity-based protein profiling to study 1,2,3-triazole ureas in Mycobacterium tuberculosis research.
The challenge was not only to detect compound-engaged proteins. The researchers also needed to understand which targets were more likely to explain the compound series' biological activity.
Source: Identification of cell wall synthesis inhibitors active against Mycobacterium tuberculosis by competitive activity-based protein profiling.
Methods
The researchers screened a library of serine hydrolase inhibitors and selected four compounds for deeper comparison: AA691, AA692, AA701, and AA702. They then used competitive ABPP to profile serine hydrolase targets affected by these compounds in the M. tuberculosis proteome.
The study combined several layers of evidence:
- Phenotypic screening to identify active compounds
- Competitive ABPP to profile inhibited serine hydrolase targets
- Biochemical enzyme assays to test individual target inhibition
- Computational docking to compare predicted binding modes
- Morphological profiling to connect target activity with cell wall-related effects
This multi-step design helped the researchers avoid treating every detected protein as equally important.
Results
The study screened about 200 compounds and focused on four compounds that formed a structure-activity series. Competitive ABPP helped prioritize serine hydrolase targets associated with the active compounds.
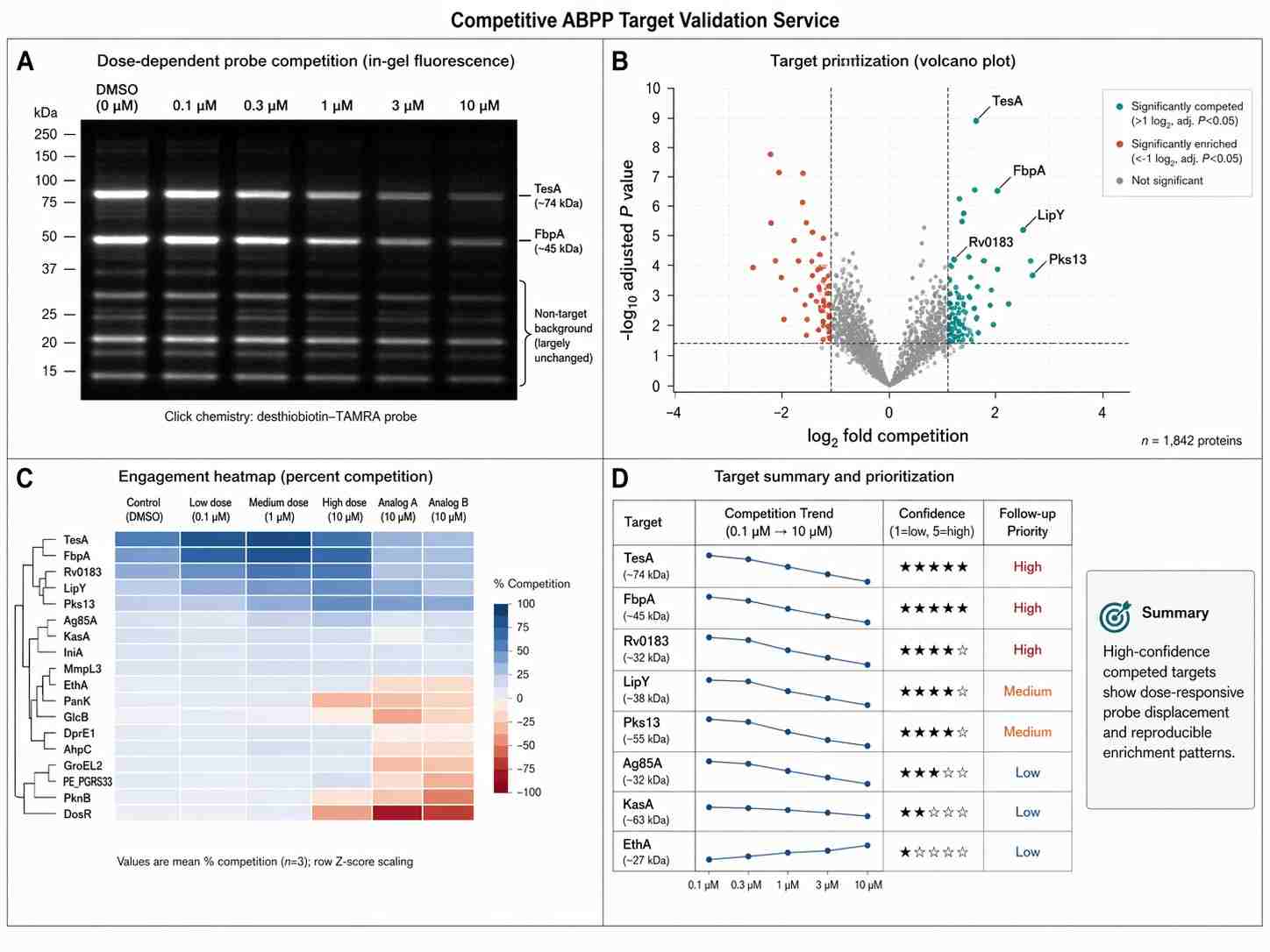
In Figure 3, the researchers measured residual activity of individual serine hydrolases after incubation with the selected compounds. The targets included TesA, FbpA, Rv0183, and Fas.
Several observations are especially relevant for target validation:
- AA691 and AA692 inhibited TesA in close stoichiometry, while AA701 and AA702 showed weaker relative inhibition.
- The competitive ABP assay showed a similar inhibition pattern for FbpA.
- Rv0183 activity was strongly impaired by all tested compounds, which helped separate broadly inhibited targets from targets that better tracked with the active compound pattern.
- Fas was not significantly inhibited under the tested competitive ABP assay conditions, helping narrow the follow-up focus.
These observations show why Competitive ABPP is useful for target prioritization. A detected target is only the starting point. The more important question is whether target inhibition follows the compound activity pattern and supports a biologically meaningful mechanism.
Conclusion
This case shows how Competitive ABPP can support target prioritization in a complex proteome. By combining probe competition with enzyme-level validation and functional follow-up, the study moved from a broad target profile to a more useful interpretation of which targets were most relevant to the compound series.
For our customers, this is the practical value of Competitive ABPP: it helps turn target engagement data into a follow-up-ready decision framework.