Multiplex Cell Death Pathway Analysis by Targeted LC-MS/MS
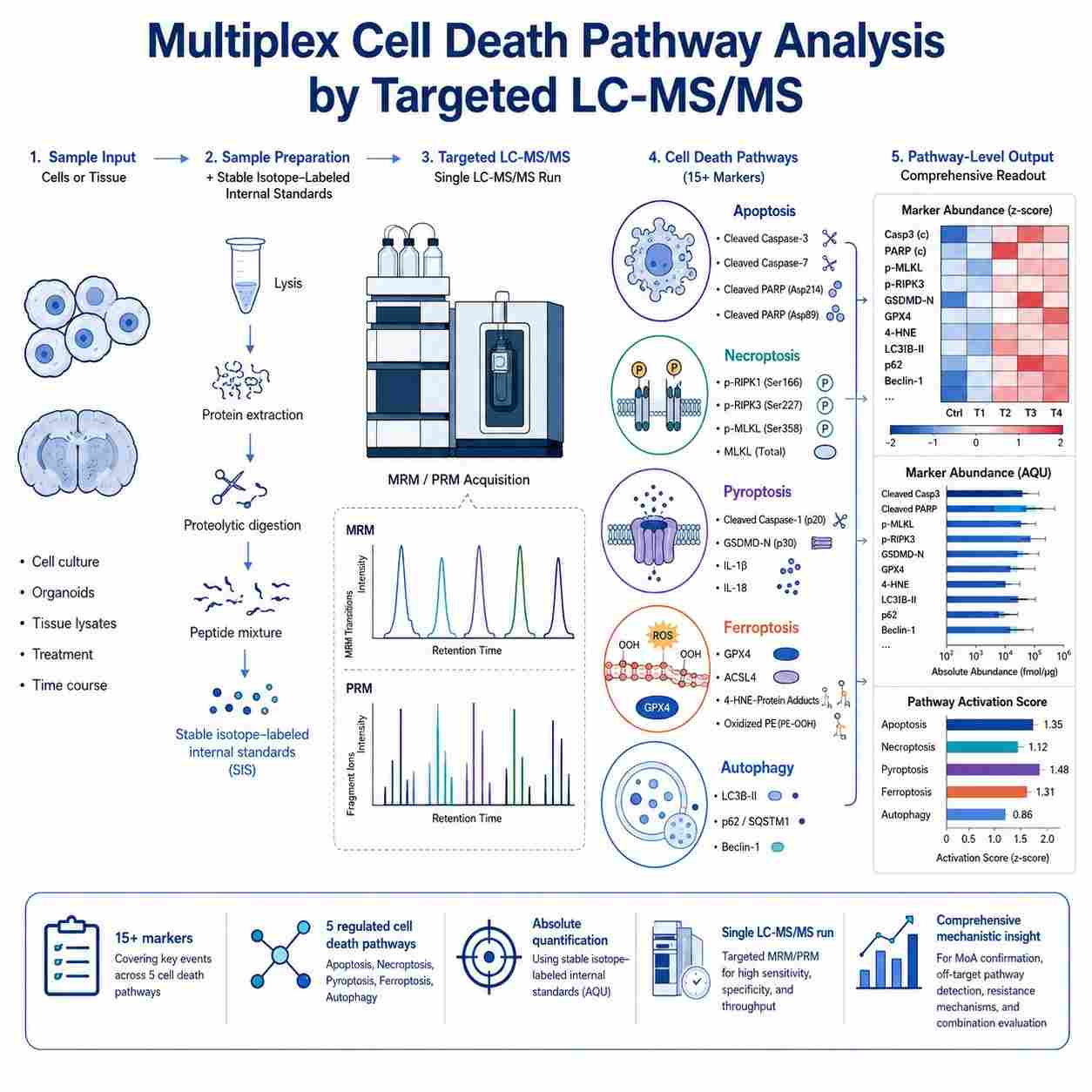
Multiplex quantification of protein and metabolite markers across five major regulated cell death pathways — apoptosis, necroptosis, pyroptosis, ferroptosis, and autophagy — from a single sample injection.
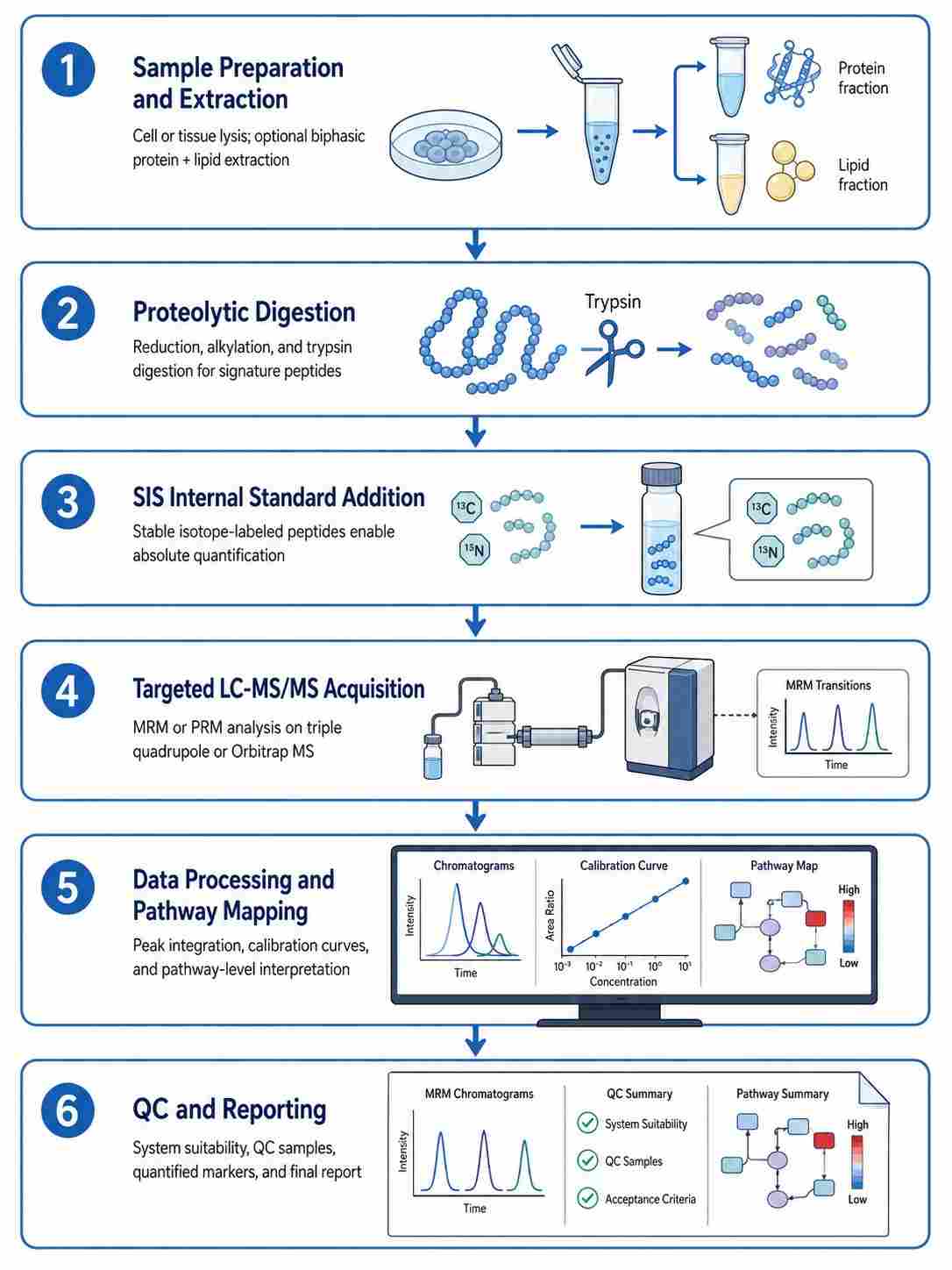
Cell death pathway MS signatures is a targeted mass spectrometry approach that enables simultaneous, absolute quantification of protein and metabolite markers across five major regulated cell death pathways — apoptosis, necroptosis, pyroptosis, ferroptosis, and autophagy — from a single sample injection, providing comprehensive cell death modality profiling with superior specificity and dynamic range compared to conventional antibody-based methods.
Key Advantages:
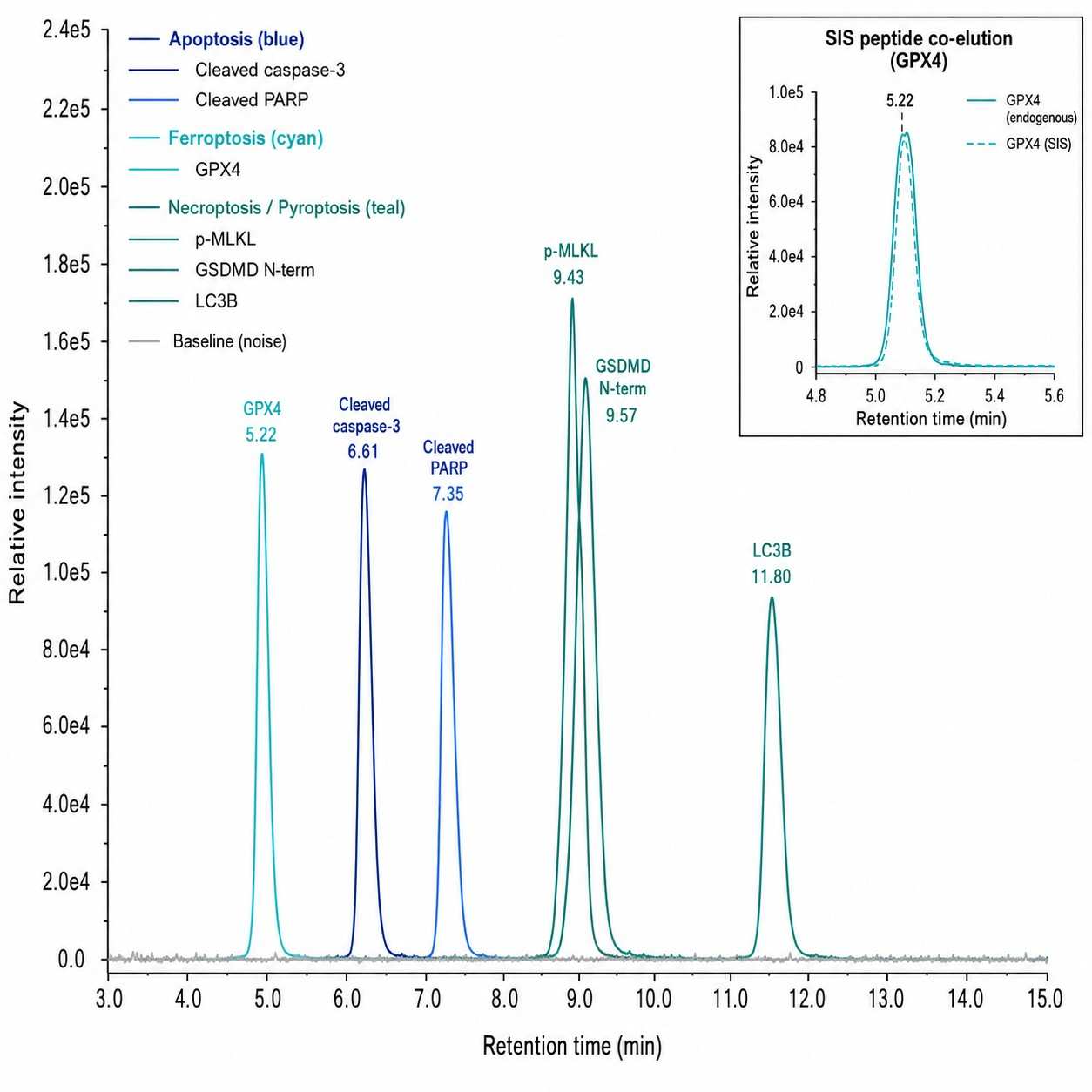
- Multiplex quantification of 15+ markers across 5 cell death pathways in a single LC-MS/MS run
- Absolute quantification with stable isotope-labeled internal standards
- Isoform-specific detection of cleaved vs. full-length caspases, gasdermins, and MLKL
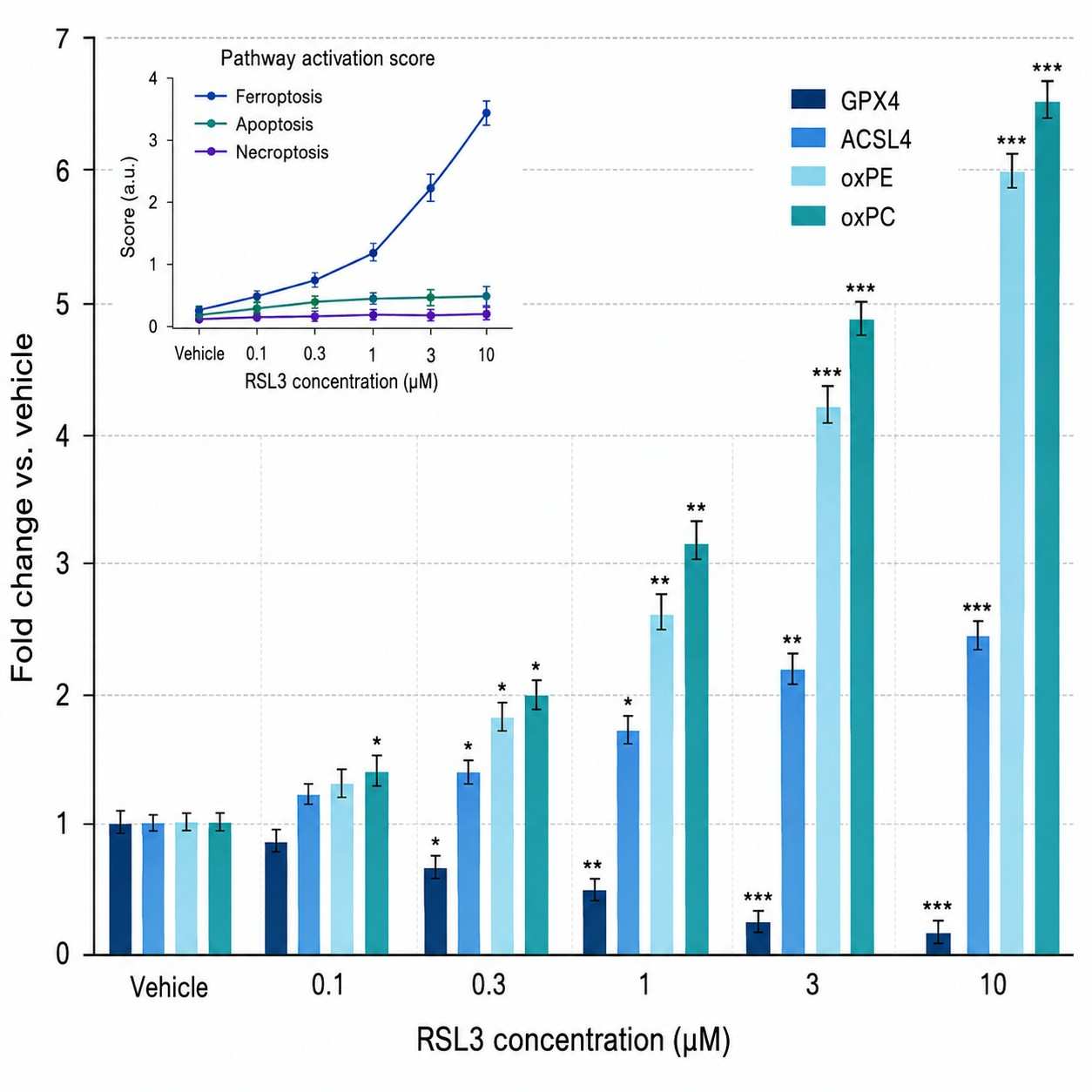
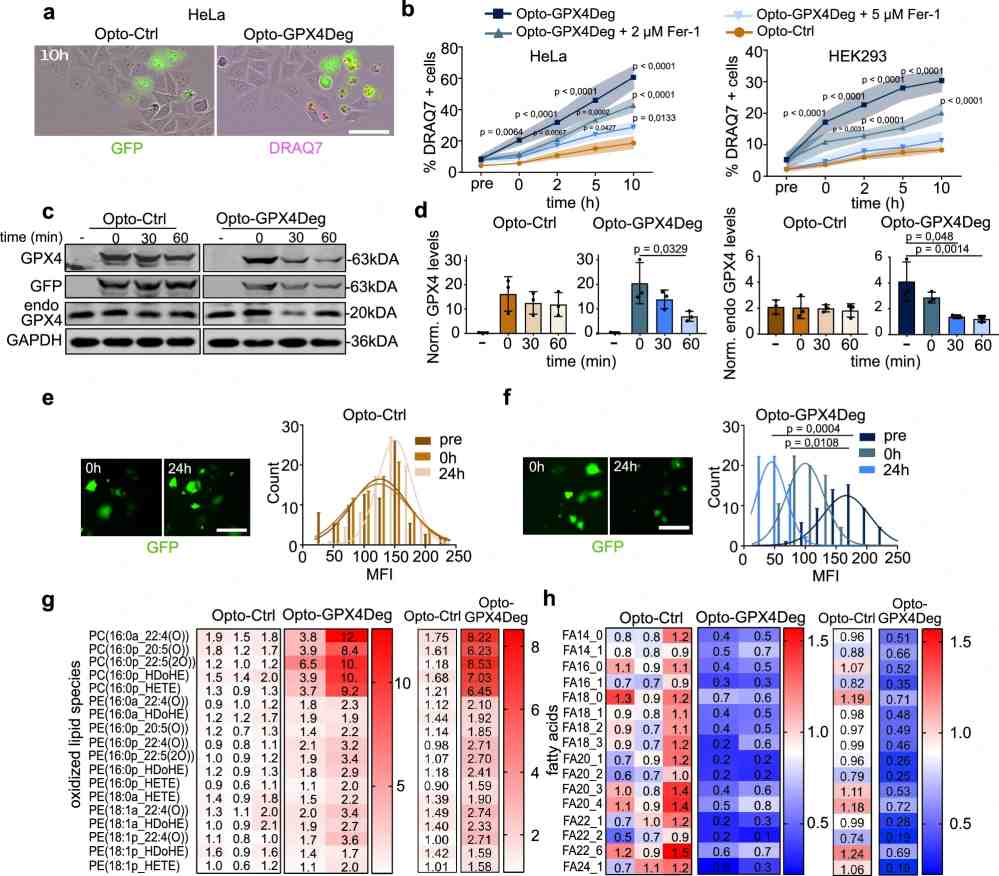
- Integrated proteomics + lipidomics for ferroptosis (GPX4, ACSL4, oxidized phospholipids)
- 50-100 µg total protein sufficient for comprehensive pathway profiling