Overcoming the SuFEx MS Profiling Bottleneck
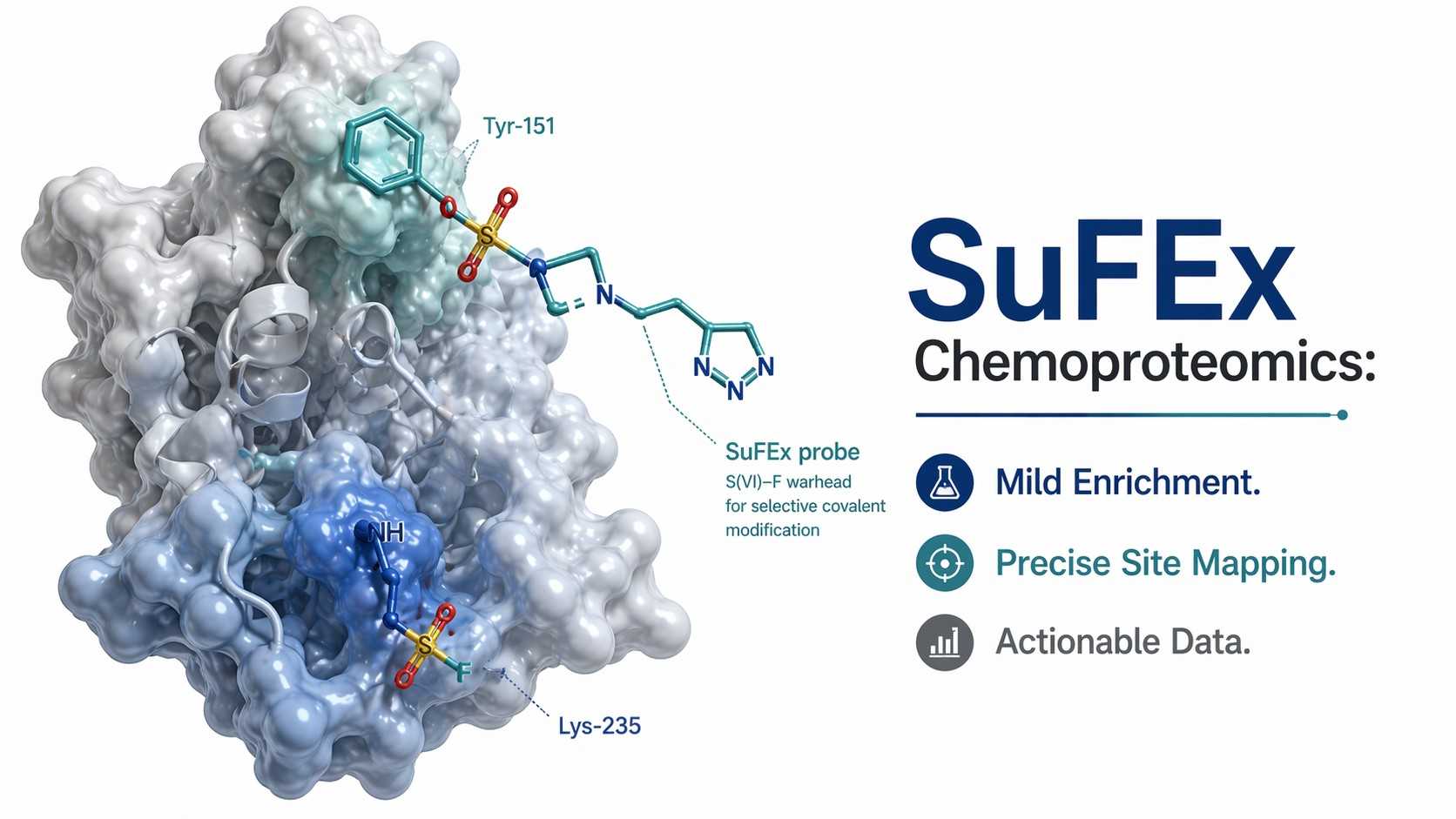
In recent years, the discovery of Sulfur(VI) fluoride exchange (SuFEx) chemistry has revolutionized the way we design covalent drugs. By utilizing sulfonyl fluoride (SVI-F) warheads, researchers can now target nucleophilic amino acids beyond traditional cysteine—such as tyrosine, lysine, histidine, and serine. This opens up massive new opportunities for targeting previously "undruggable" proteins and developing novel molecular glues.
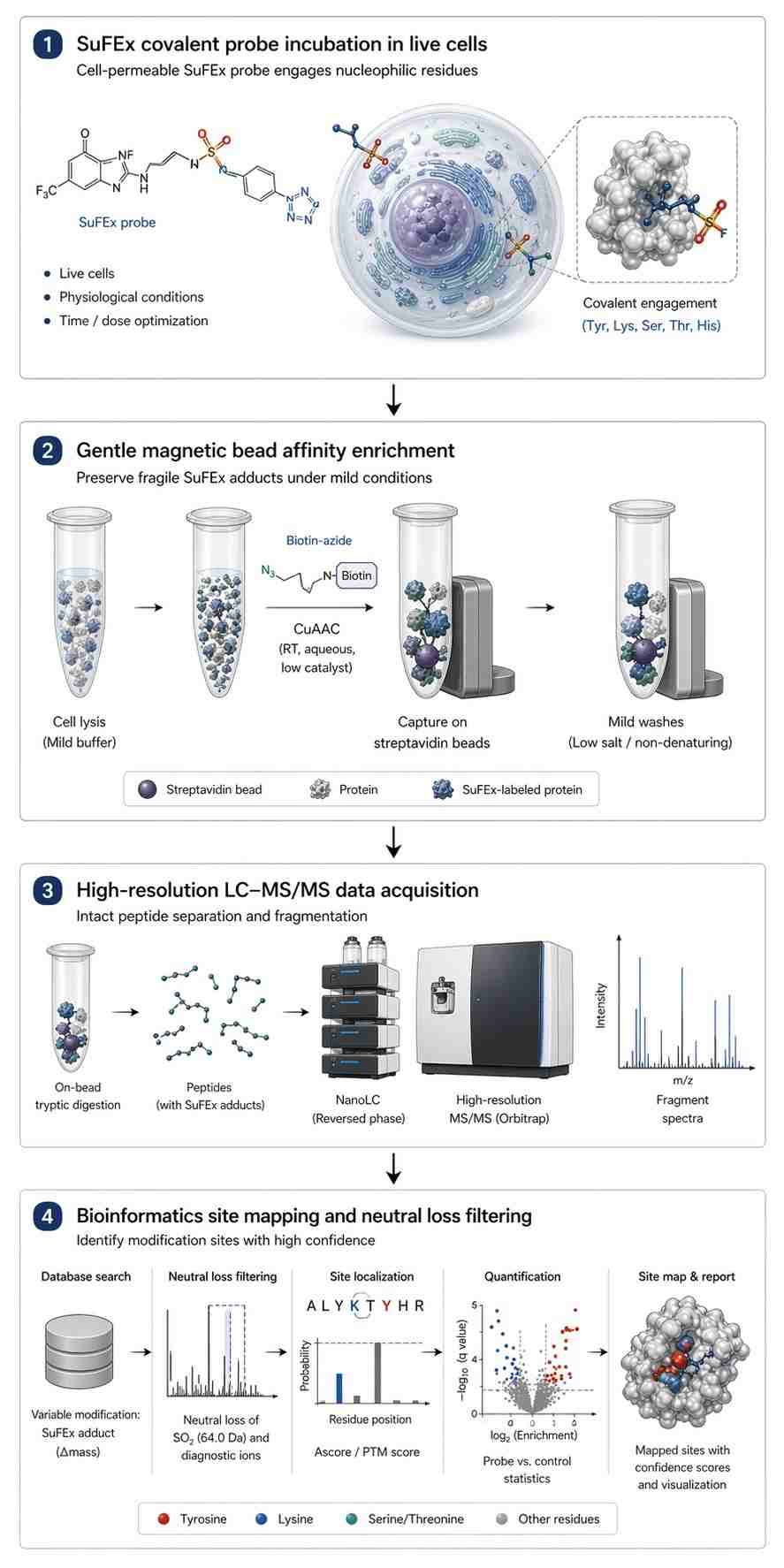
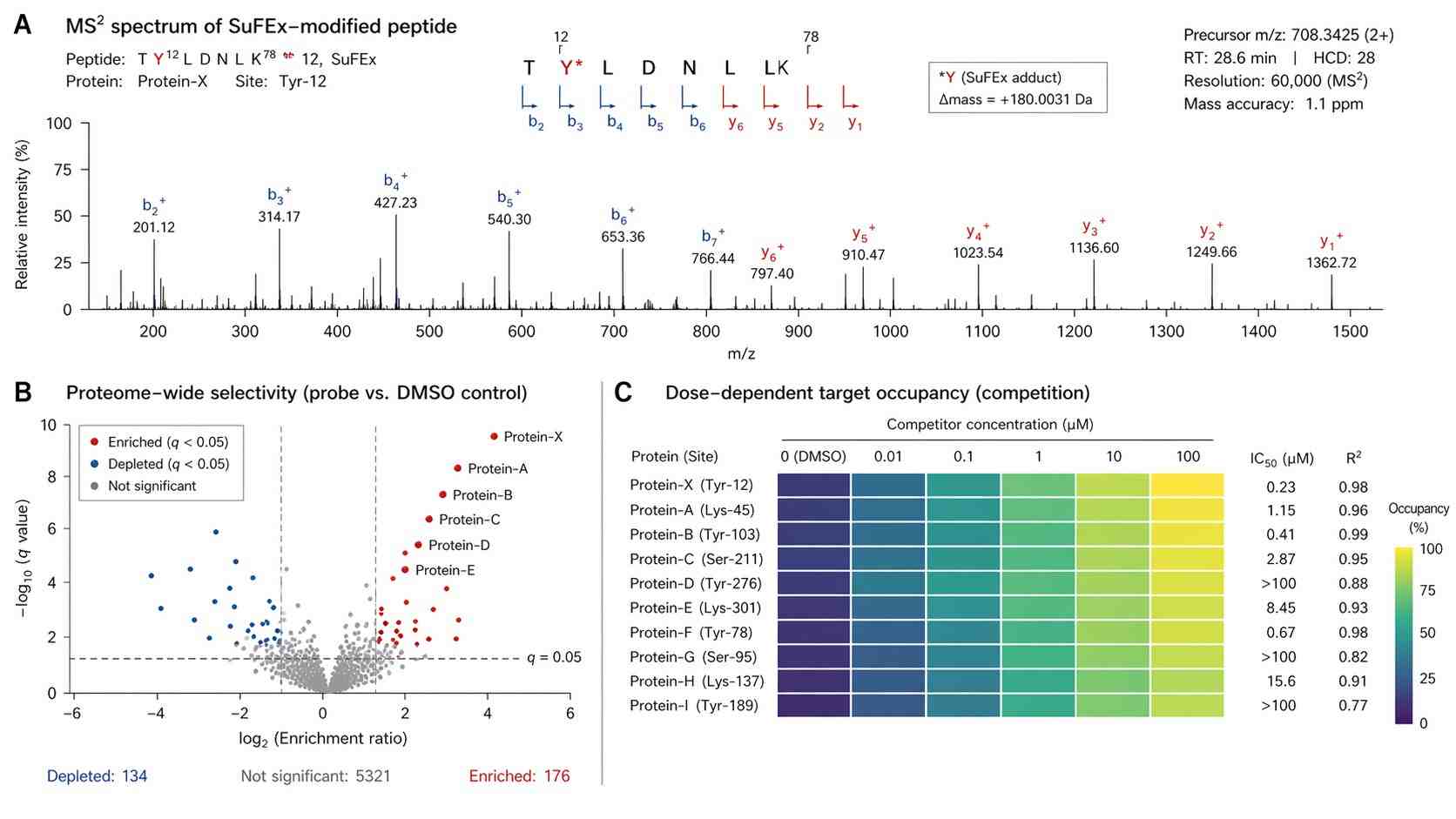
However, moving from a successful chemical synthesis to actual biological target identification presents a major technical bottleneck. When standard mass spectrometry (MS) sample preparation protocols are used, the delicate SVI-F covalent bonds often break down (hydrolyze) due to harsh pH levels or high temperatures. Even if the sample reaches the mass spectrometer intact, SuFEx adducts tend to exhibit "neutral loss"—meaning the probe breaks apart unexpectedly inside the machine before the protein sequence can be read. This leads to missing data, high background noise, and overwhelming false-positive rates.
Our platform is built specifically to address these exact problems. By combining carefully controlled, mild sample preparation with custom-tuned bioinformatics algorithms, we successfully capture and identify these fragile modifications, translating your complex chemistry into clear, actionable biological data. We prioritize the stability of the covalent linkage, ensuring that the critical evidence of target engagement is preserved from the moment of lysis to the final spectral readout.