Background
In early drug discovery, potency and selectivity are both critical. A compound may show activity in a biochemical assay, but its behavior in a cellular matrix can differ because of permeability, protein accessibility, pathway context, and off-target engagement.
Jones and colleagues addressed this challenge in ABPP-HT - High-Throughput Activity-Based Profiling of Deubiquitylating Enzyme Inhibitors in a Cellular Context. The study focused on deubiquitylating enzyme inhibitors and evaluated how an accelerated ABPP workflow could support inhibitor selectivity profiling in cellular and tissue-derived biological contexts.
Methods
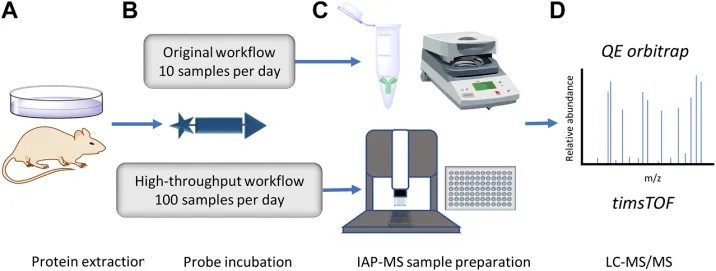
The study developed a high-throughput-compatible activity-based protein profiling workflow using activity-based probe labeling, immunoaffinity purification, sample preparation, LC-MS/MS, and label-free quantification. Figure 1 describes the accelerated DUB inhibitor ABP immunoprecipitation workflow, including protein extraction and inhibitor treatment, HA-Ub-PA probe incubation, anti-HA enrichment, and LC-MS/MS proteomic analysis.
The authors optimized multiple workflow variables, including starting material, probe labeling, immunoaffinity purification, elution, and sample preparation. They also used filtering rules during data analysis, removing DUBs based on no-probe controls, missing values in probe controls, or signals near the lower limit of MS dynamic range. Raw data and MaxQuant search results were deposited to PRIDE under dataset identifier PXD023036.
Results
The study reported that ABPP-HT implemented a semi-automated proteomic sample preparation workflow that increased throughput capabilities by approximately ten times compared with the classical ABPP workflow while preserving enzyme profiling characteristics. The method identified a reduced but representative panel of DUBs, about 15-25 DUBs compared with about 30-40 DUBs in regular ABPP.
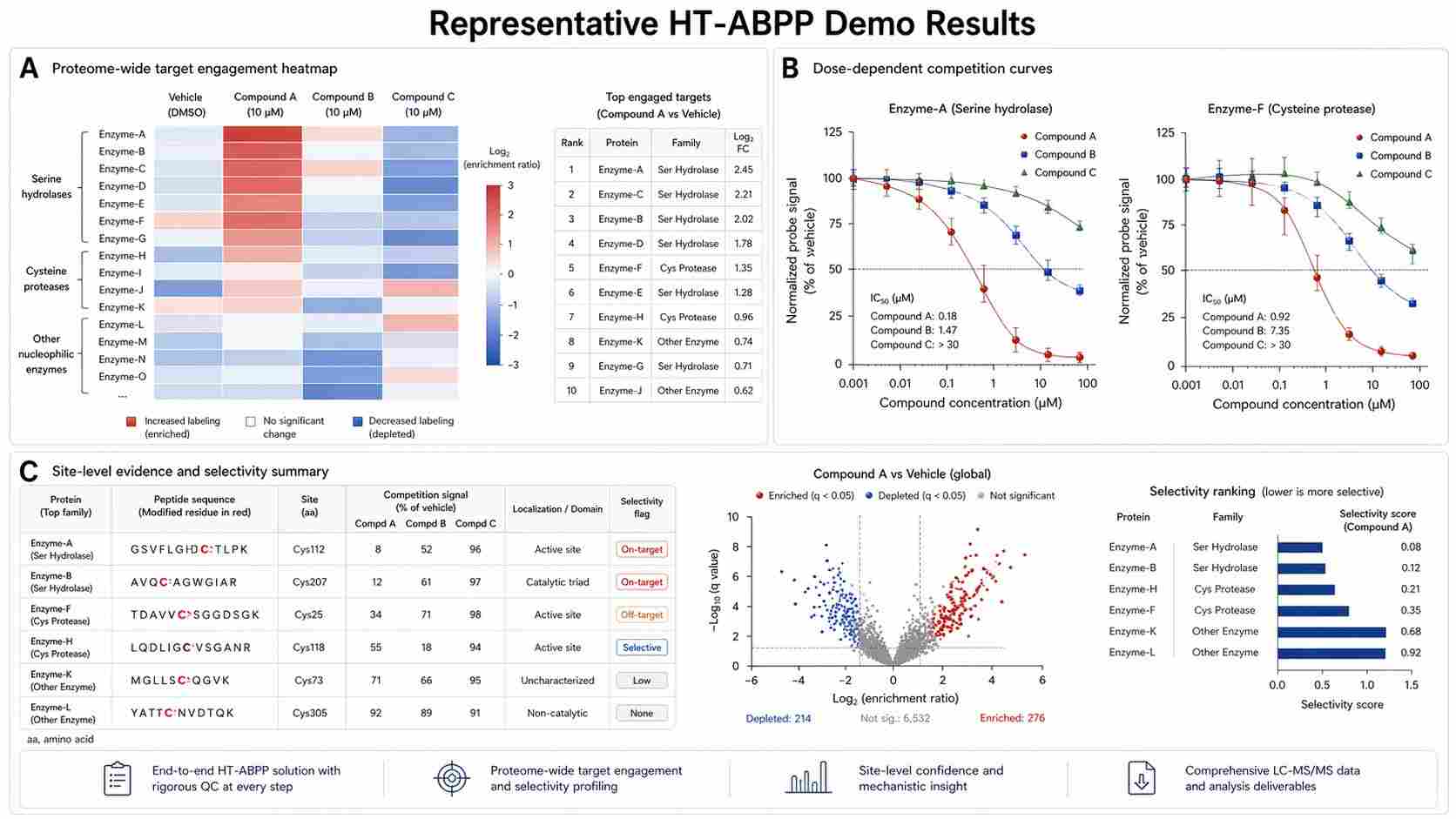
The authors used FT671, a selective USP7 inhibitor, and NEM, a broad cysteine modifier, to test whether the workflow could distinguish selective and non-selective inhibition patterns. They observed that NEM inhibited USP7 in both cells and brain in a non-selective way, while FT671 retained a selective USP7 profile. The study also tested multiple compounds and concentrations, including four USP7 inhibitors, two USP30 inhibitors, and two broad cysteine modifiers. The results were summarized in heat maps for USP7 inhibitors, USP30 inhibitors, and non-selective modifiers.
Conclusion
This study shows why HT-ABPP project design must balance throughput, target coverage, QC, and interpretability. A faster workflow is valuable only when it still supports meaningful enzyme or target profiling, clear control filtering, and selectivity interpretation. For covalent ligand discovery projects, the same principle applies: sample design, probe strategy, enrichment, LC-MS/MS acquisition, and data analysis should be planned together before screening begins.