This literature-supported case study is based on the English open-access article ABPP-HT*—Deep Meets Fast for Activity-Based Profiling of Deubiquitylating Enzymes Using Advanced DIA Mass Spectrometry Methods. It is not a Creative Proteomics customer case.
Background
Deubiquitylating enzymes, or DUBs, regulate ubiquitination and are important in cellular processes such as protein degradation and signaling. The paper notes that DUB inhibitors are being developed for diseases including Parkinson's disease and cancer. Because DUB inhibitor selectivity can be affected by potency, permeability, reversibility, stability, and off-target reactivity, the authors used ABPP with LC-MS/MS to measure active DUB profiles in a cellular matrix.
The study also addressed a practical problem in ABPP drug discovery workflows. Earlier high-depth ABPP could identify more than 70 DUBs in MCF-7 breast cancer cells but required low-pH C18 HPLC pre-fractionation and multiple 60-minute LC-MS/MS runs. A later high-throughput ABPP-HT workflow reached about 100 samples per day, but its depth dropped to about 15–25 DUBs. The authors therefore aimed to retain high throughput while improving DUBome depth and data completeness.
Methods
The authors used MCF-7 cell lysates and the HA-Ub-PA activity-based probe, which contains an HA tag for immunoprecipitation, ubiquitin for DUB specificity, and a propargylamine warhead that binds active-site cysteine residues. Lysates were diluted to 3.33 mg/mL, with 250 μg of protein per reaction unless otherwise stated. Inhibitors or DMSO control were incubated with lysate for 1 hour at 37°C, followed by HA-Ub-PA probe labeling for 45 minutes at 37°C.
The workflow used automated immunoprecipitation on an Agilent Bravo AssayMAP liquid handling platform, tryptic digestion, Evosep/timsTOF LC-MS/MS, and comparison of DDA and DIA data acquisition strategies. The authors compared Fragpipe and MaxQuant for DDA analysis and DIA-NN and MaxDIA for DIA analysis. They also tested known DUB inhibitors, including FT827, HBX41108, P22077, and the pan-DUB inhibitor PR619, to evaluate whether the workflow could support inhibitor selectivity profiling.
Results
Figure 1 shows that DIA data analyzed with DIA-NN quantified more DUBs than DDA data analyzed with Fragpipe for the same ABPP-HT samples. DIA-NN identified an additional 10 DUBs that were not quantified by Fragpipe. The authors also compared HA-Ub-PA probe samples with a no-probe control. In Fragpipe analysis, all quantified DUBs except USP4 were enriched more than 10-fold. In DIA-NN analysis, all DUBs except USP15, USP34, USP38, USP48, and OTUD7B were enriched more than 10-fold in the presence of HA-Ub-PA.
The same figure also illustrates why background review matters in ABPP data interpretation. DIA-NN quantified 912 total proteins compared with 218 total proteins for Fragpipe, reflecting higher sensitivity but also increased detection of co-immunoprecipitated or non-specific background proteins. The authors therefore emphasized the value of no-probe controls when interpreting enriched proteins.
Figure 2 evaluates reproducibility and missing values. Both DIA/DIA-NN and DDA/Fragpipe showed average DUBome CV values below 10%. Fragpipe and DIA-NN with match-between-runs had no missing values in this dataset. The authors noted that this is important because, in inhibitor profiling, the absence of a signal at high inhibitor concentration could otherwise be confused with a missing value.
Figure 3 tests sensitivity using peptide injection titration after immunoprecipitation from 100 μg of lysate. Without match-between-runs, both DIA-NN and Fragpipe detected a representative panel of 10–15 DUBs with only 50 ng of peptides injected. DIA data analyzed with DIA-NN and match-between-runs was the most sensitive across the titration.
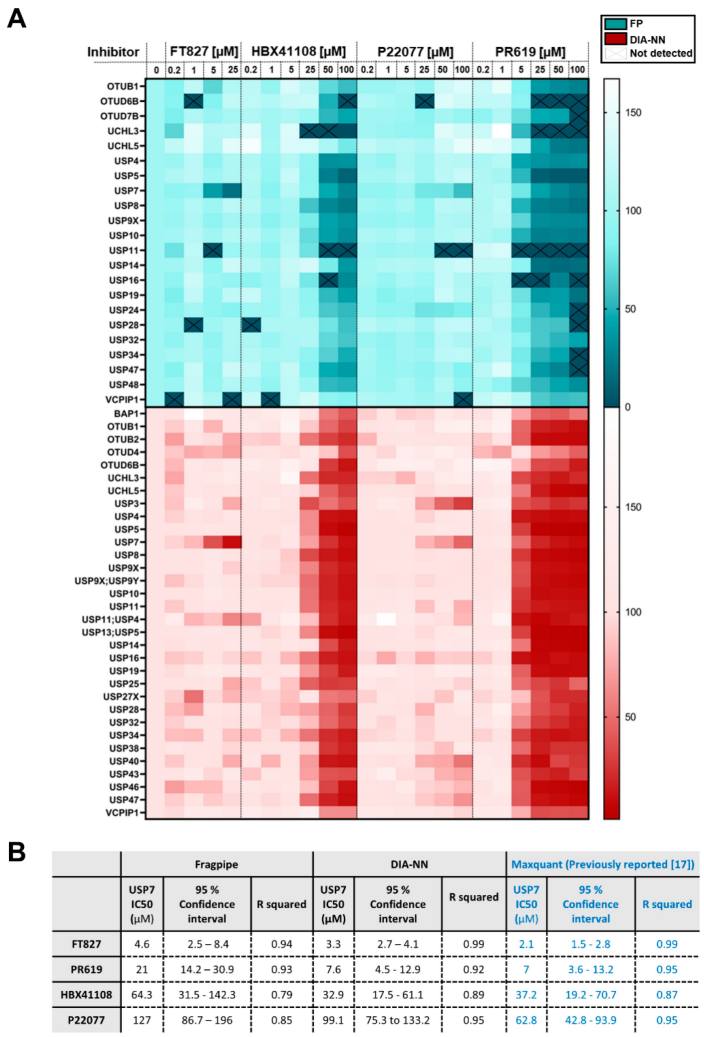
Figure 4 demonstrates inhibitor profiling. After filtering low-intensity DUBs that did not show concentration-dependent behavior, DIA/DIA-NN identified 29 DUBs, compared with 22 DUBs identified by DDA/Fragpipe. Both workflows confirmed known inhibitor profiles across the DUBome. FT827 showed high specificity for USP7, while PR619 showed broad reactivity across the DUB panel. The authors also reported that DIA-NN showed fewer missing values than Fragpipe across the larger inhibitor dataset, especially at high PR619 concentrations.
Conclusion
The authors concluded that optimizing data acquisition and search-engine selection increased the depth of a high-throughput ABPP workflow by about 50% while maintaining fast sample processing. Their ABPP-HT* method combined DIA-MS and DIA-NN analysis to improve sensitivity, reduce missing values, and support more reliable DUB inhibitor selectivity profiling.
For Global ABPP service planning, this English literature case supports several EEAT points. First, ABPP can provide functional enzyme activity evidence that is not captured by abundance proteomics alone. Second, LC-MS/MS acquisition and data-analysis strategy can strongly affect depth, missing values, and interpretation. Third, no-probe controls, replicate consistency, enrichment review, and inhibitor concentration-response behavior are essential for distinguishing real activity-linked signals from background.