Protease MS Substrate/Product Assays — Label-Free Enzyme Activity Screening by Mass Spectrometry
Accelerating protease inhibitor discovery and enzyme characterization through direct, interference-free MS-based detection of substrate depletion and product formation.
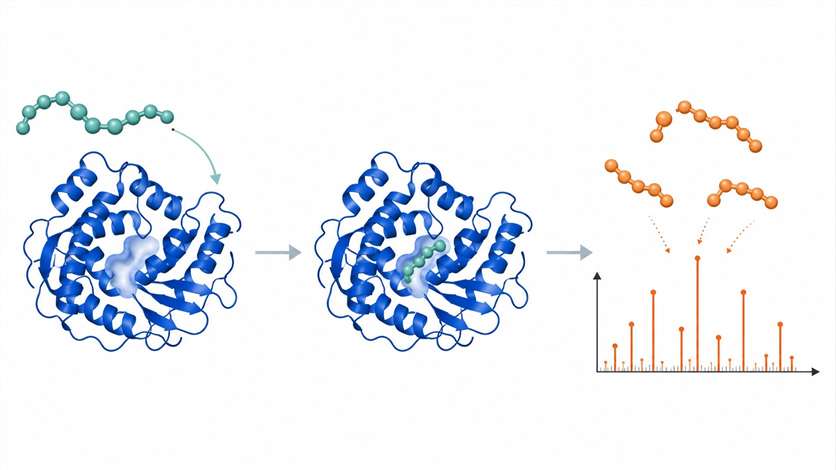
Proteases represent one of the largest and most therapeutically important enzyme families, yet traditional fluorescence-based activity assays bring persistent challenges: compound interference, limited availability of suitable fluorogenic substrates, and an inability to monitor more than one substrate per well. Mass spectrometry (MS)-based protease substrate/product assays solve these problems at their root by detecting substrates and cleavage products directly — by their mass — without any fluorescent label at all.
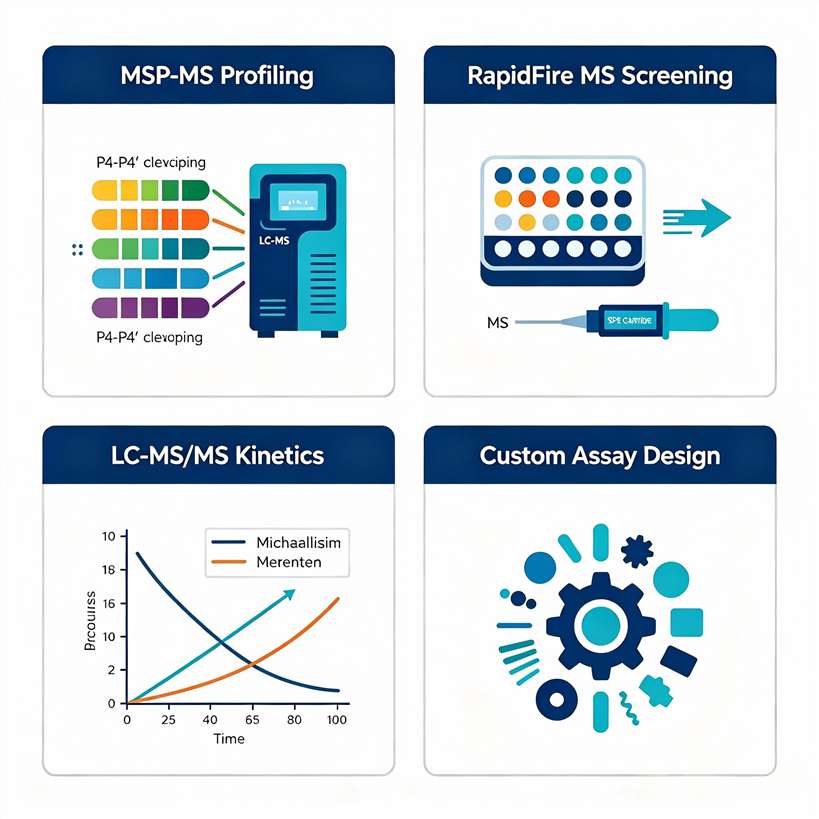
We built our protease MS assay platform around three complementary technologies — RapidFire MS, LC-MS/MS, and MALDI-TOF — to deliver high-throughput inhibitor screening, multiplex substrate specificity profiling (MSP-MS), and full kinetic characterization across serine, cysteine, metallo, aspartyl, and threonine proteases.
Key Advantages:
- Label-free direct detection of substrates and products by accurate mass — no fluorophore required.
- Compatible with serine, cysteine, metallo, aspartyl, and threonine proteases.
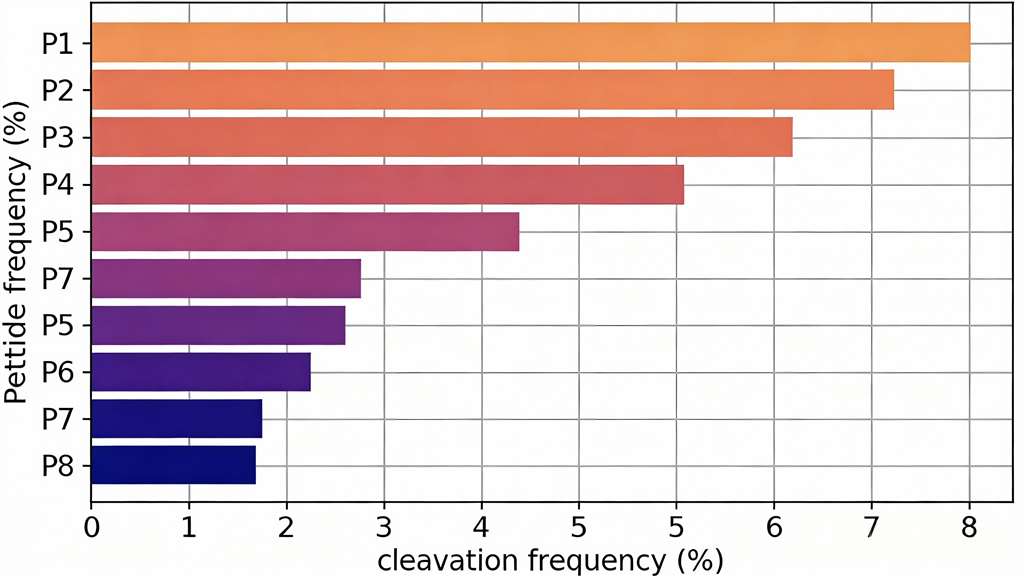
- Multiplex substrate profiling (MSP-MS) for comprehensive specificity fingerprinting.
- RapidFire MS and MALDI-TOF platforms for medium-to-high-throughput inhibitor screening.