Single-cell MS Drug Profiling Service
Reveal drug action at single-cell resolution — directly measure drug uptake, metabolism, and phenotypic response from individual cells using high-sensitivity mass spectrometry.
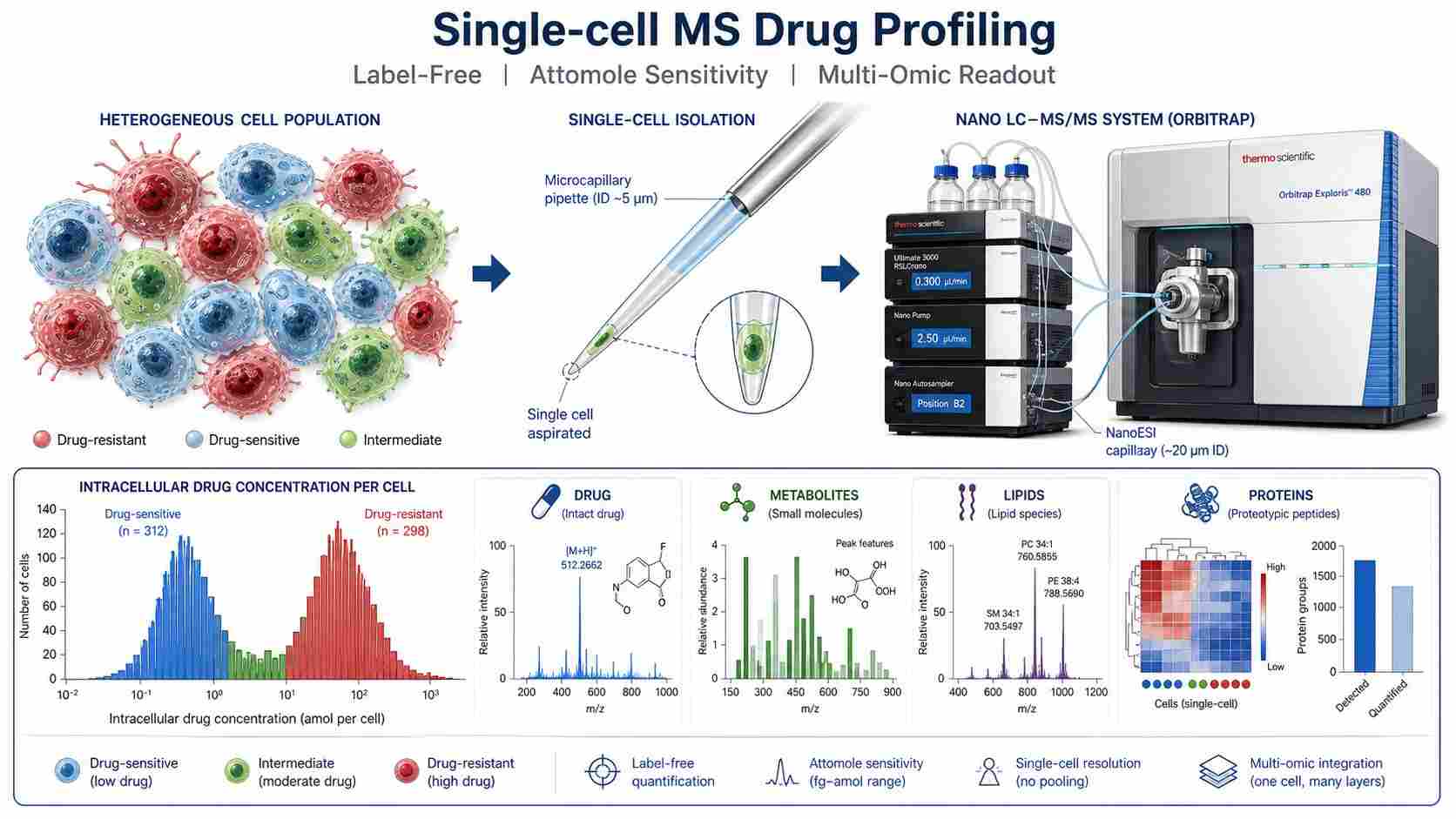
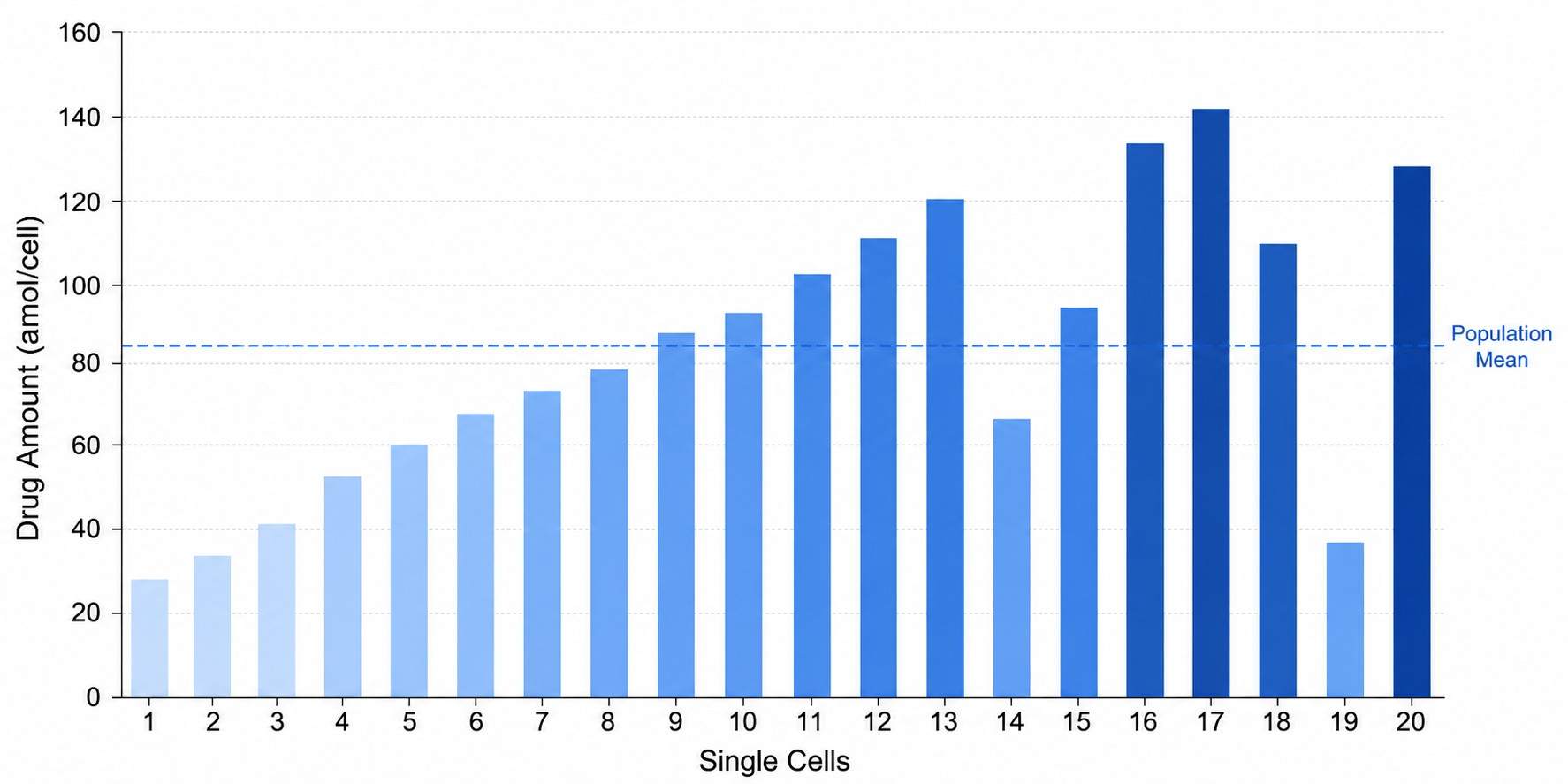
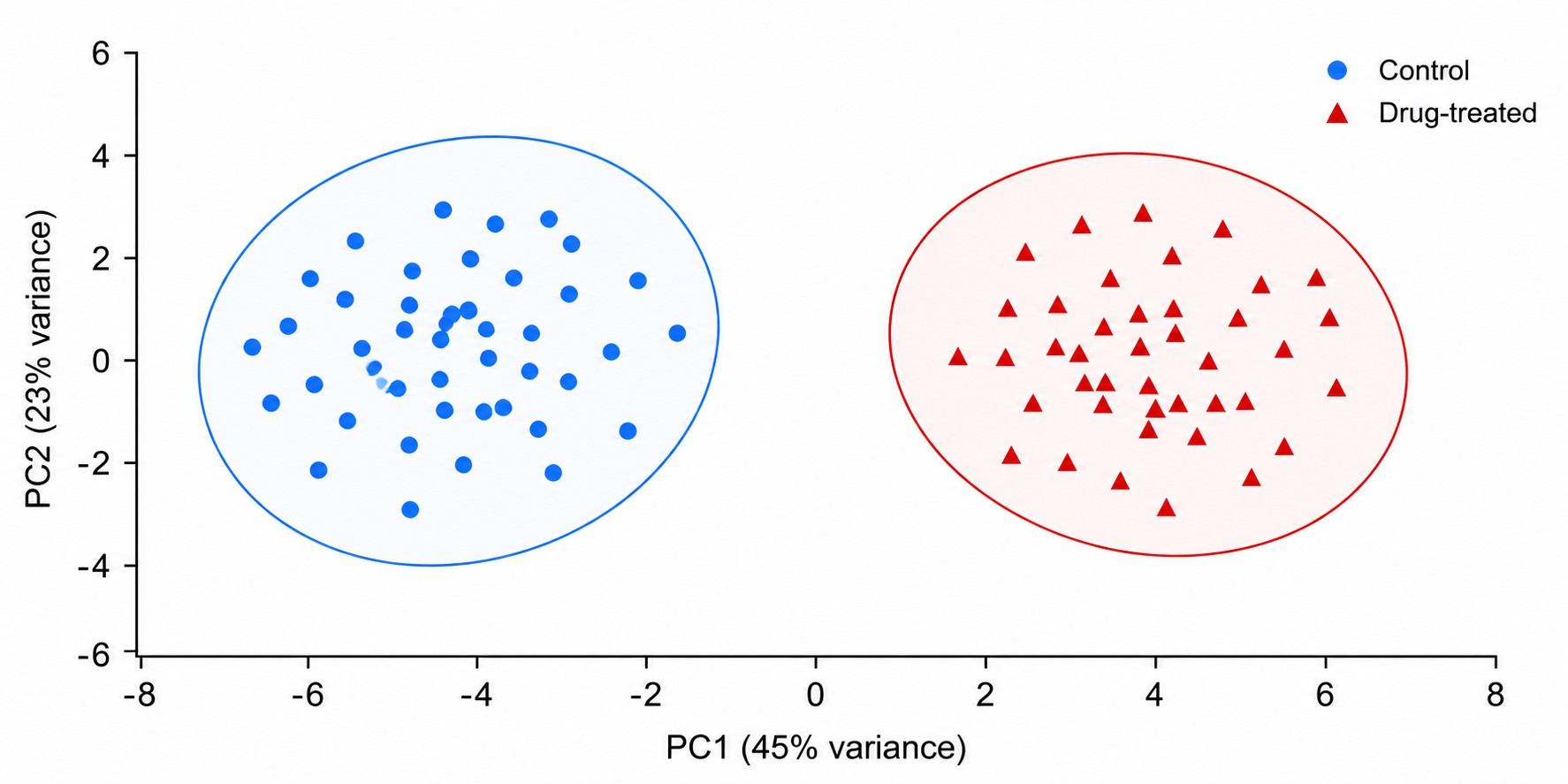
Single-cell mass spectrometry (scMS) drug profiling lets you quantify drugs, metabolites, lipids, and proteins from individual cells — not just population averages. While bulk assays report a single mean value that masks what individual cells are doing, scMS reveals the true heterogeneity of drug action: the drug-resistant subpopulation hiding in plain sight, the cell that metabolizes the drug differently from its neighbors, the rare cell that escapes target engagement.
At Creative Proteomics, our MassTarget™ platform combines advanced single-cell isolation with high-sensitivity nanoLC-MS/MS to deliver label-free drug profiling at single-cell resolution. Whether you are investigating chemoresistance mechanisms, mapping cell-specific pharmacokinetics, or building a multi-omic picture of drug response, our service provides the analytical depth to support informed discovery decisions.
Key Advantages:
- Attomole-level drug detection from individual mammalian cells
- Multi-omic readout: drugs, metabolites, lipids, and proteins from the same cell
- Label-free — no antibody panels, genetic reporters, or radioactive tracers
- Validated across adherent, suspension, and tissue-derived cell types