BioID2 Mass Spectrometry Service for Target Discovery
Pinpoint novel protein interactions for your toughest targets.
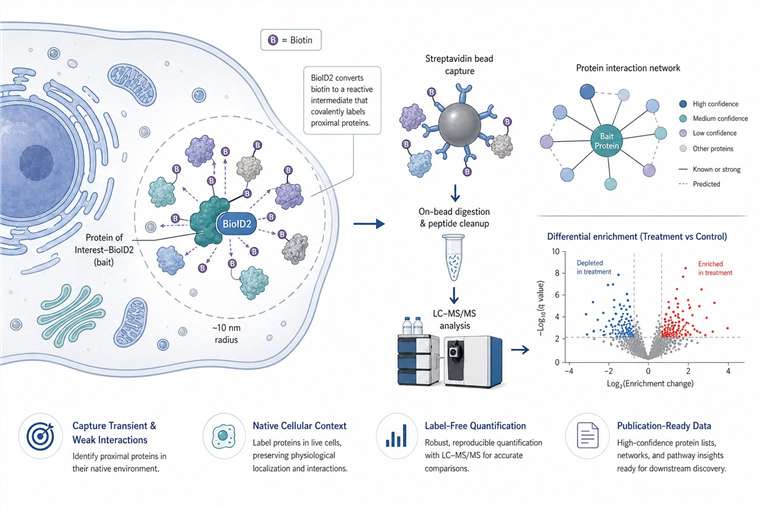
Our BioID2 proximity labeling proteomics service captures transient, weak, and membrane-associated PPIs in their native context.
From construct design to MS analysis, we deliver clean, publication-ready interactome maps to accelerate your drug discovery pipeline.
If you've ever struggled to pull down a real protein interactor—getting back nothing but the same high‑abundance contaminants, or losing that one weak, transient binding event that actually matters—you're not alone. Traditional co‑immunoprecipitation works beautifully for rock‑solid complexes. It's the faint, fleeting, and membrane‑buried interactions that slip through. That's where proximity‑dependent biotinylation changes the game, and BioID2 is one of the most forgiving ways to do it.
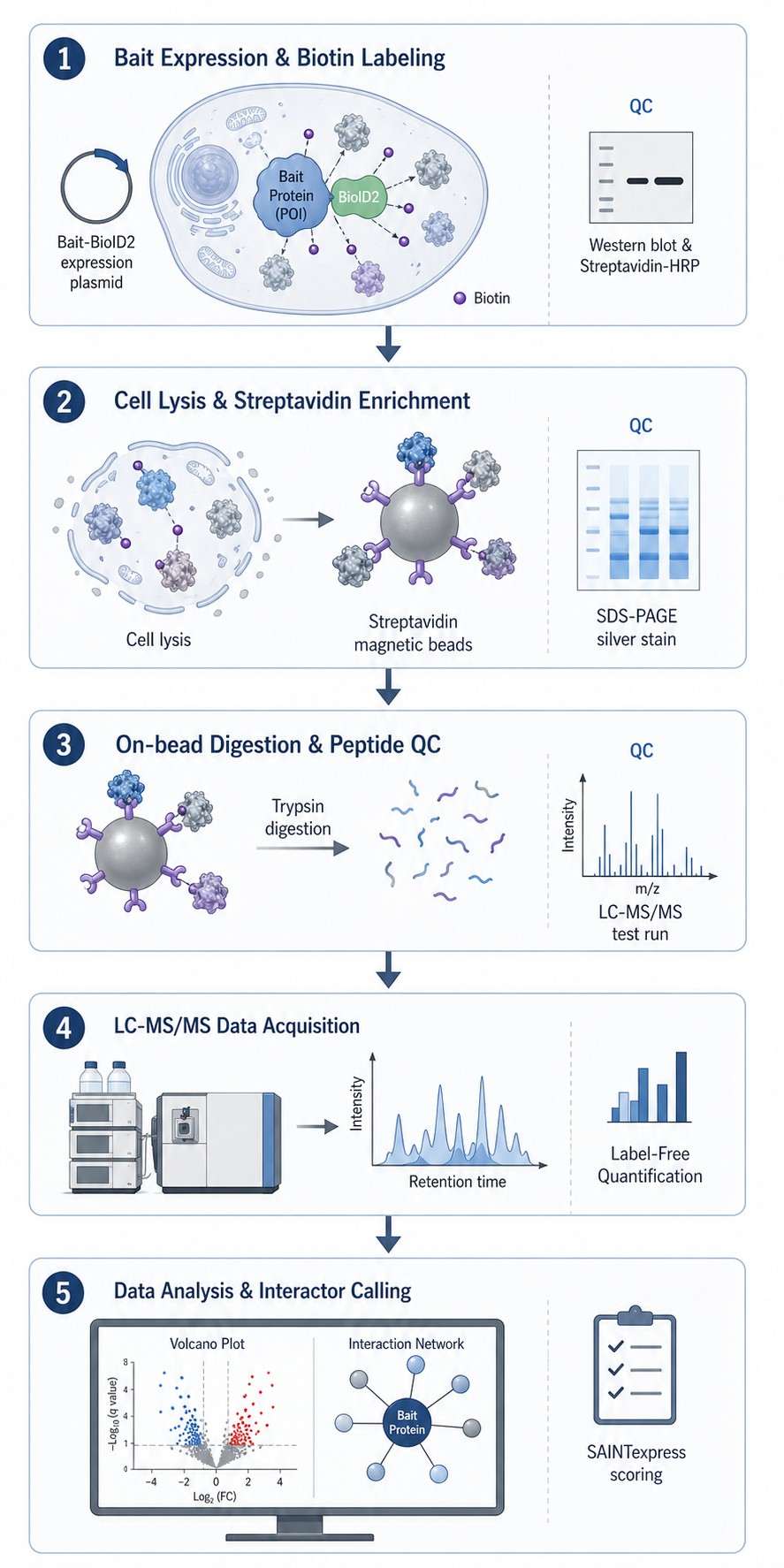
Here, we walk you through how we run BioID2 proximity labeling proteomics, what the data look like, and what you can expect from the service. The short version: we handle everything from construct strategy to the final interaction network, with clear QC gates along the way so you can trust the list of candidates you get back.
Key Advantages:
- Capture elusive protein interactions.
- End-to-end service from construct to data.
- Rigorous QC and negative control design.
- Deliverables tailored for publication.