Stable Isotope Labeling (SIL) Natural Products Analysis Service
Track metabolic fate, elucidate biosynthetic pathways, and link natural products to their gene clusters using LC-HRMS/MS-based stable isotope labeling.
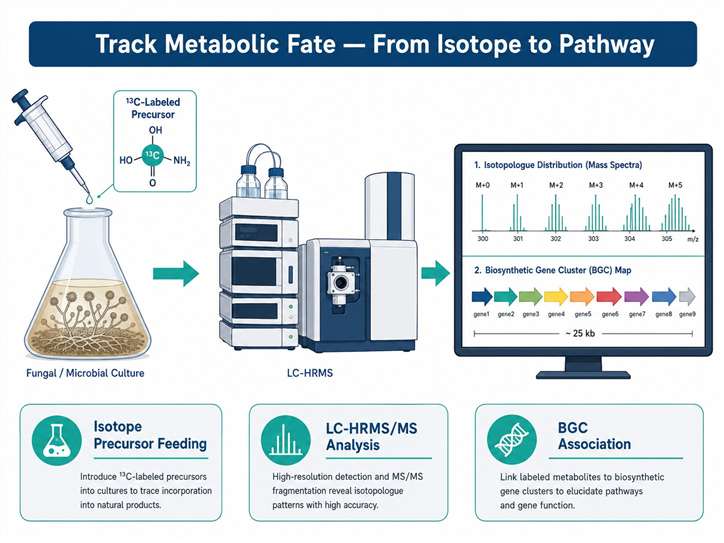
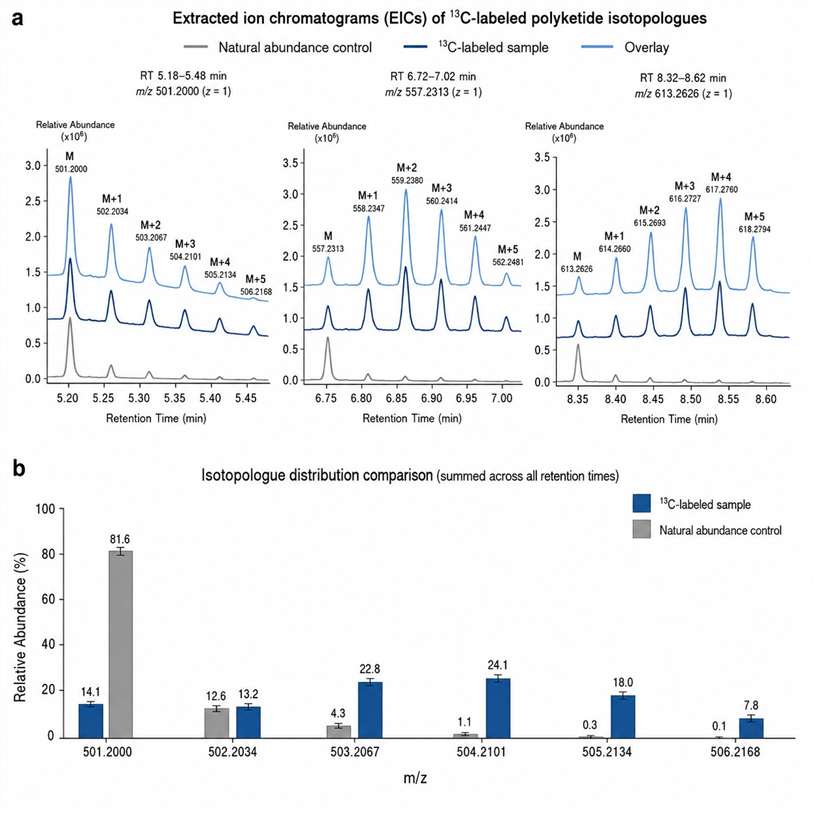
Stable isotope labeling (SIL) introduces non-radioactive isotopic tracers — ¹³C, ¹⁵N, or ²H — into biological systems to track the fate of specific precursors through metabolic networks. Coupled with high-resolution LC-HRMS/MS, SIL enables researchers to monitor isotopologue distributions, quantify metabolic flux, and establish direct precursor–product relationships in complex biological matrices.
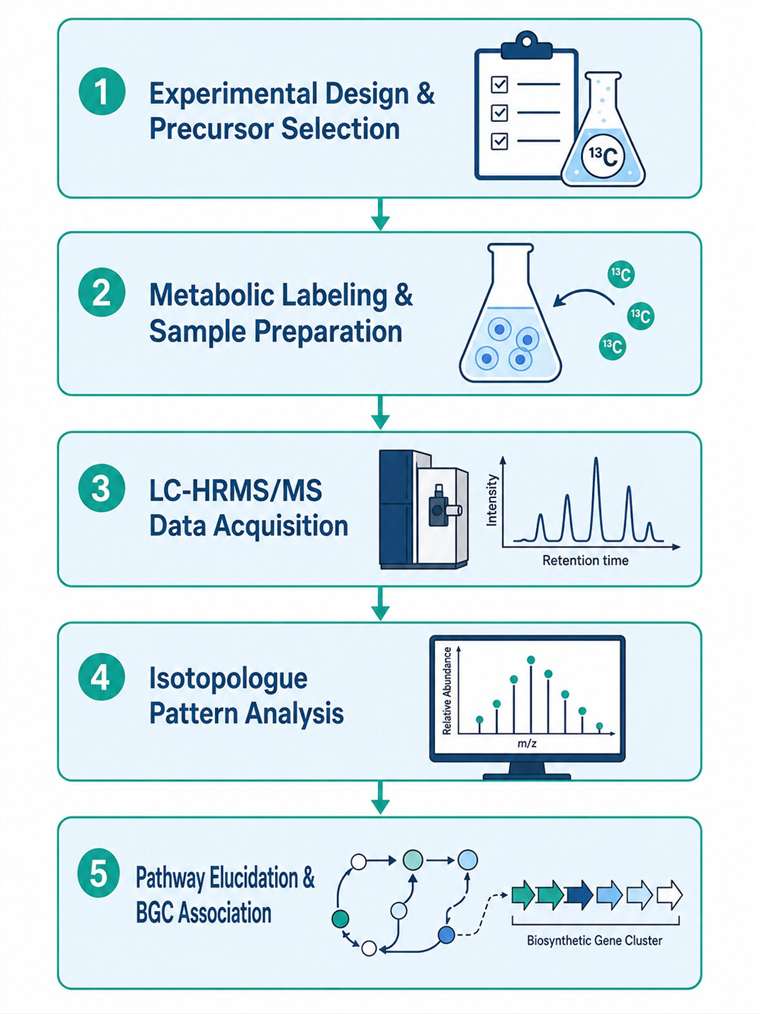
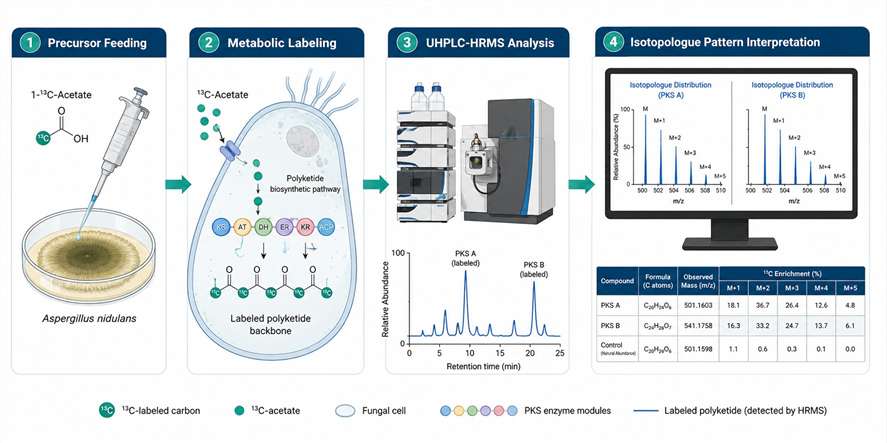
At MassTarget (a service brand of Creative Proteomics), our Stable Isotope Labeling (SIL) Natural Products Analysis Service covers the full pipeline: from isotope precursor selection and experimental design, through metabolic labeling and sample preparation, to high-resolution MS acquisition, isotopologue pattern analysis, and biological interpretation. We combine natural product chemistry expertise with advanced LC-HRMS/MS instrumentation and bioinformatics tools — including IsoAnalyst-compatible data analysis — to deliver actionable results from your labeling experiments.
Whether you are studying microbial polyketide biosynthesis, plant specialized metabolism, or fungal natural product pathways, our platform provides the analytical rigor to move your research forward.
Key Advantages:
- Natural product-specific expertise in chemistry, biosynthesis, and metabolomics.
- BGC association capability via parallel SIL strategies compatible with IsoAnalyst.
- Comprehensive precursor library including ¹³C-glucose, ¹³C-acetate, ¹³C/¹⁵N-amino acids, and custom precursors.
- Advanced LC-HRMS/MS platform with high-resolution Orbitrap and Q-TOF mass spectrometers.
- Integrated bioinformatics pipeline for isotopologue analysis, flux estimation, and BGC association.
- Full-service support from experimental design through final data interpretation.