Natural Product & Complex Mixture MS Discovery Service
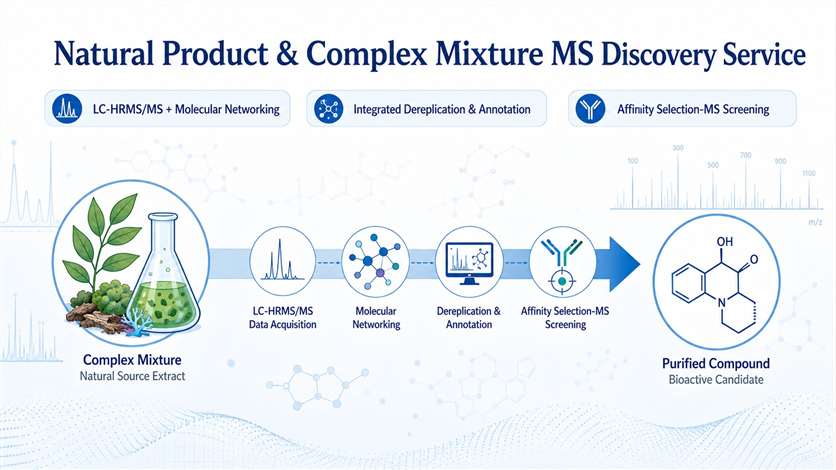
Accelerate natural product hit discovery with integrated MS-based dereplication, molecular networking, and affinity selection. From complex extracts to active compounds — faster, smarter, and without extensive purification.
Natural products remain one of the most prolific sources of novel drug leads, yet their inherent chemical complexity poses a formidable challenge for traditional discovery pipelines. Crude extracts, microbial fermentation broths, and fractionated libraries can contain hundreds to thousands of unique compounds, making the isolation and identification of bioactive molecules a time-intensive and resource-heavy endeavor.
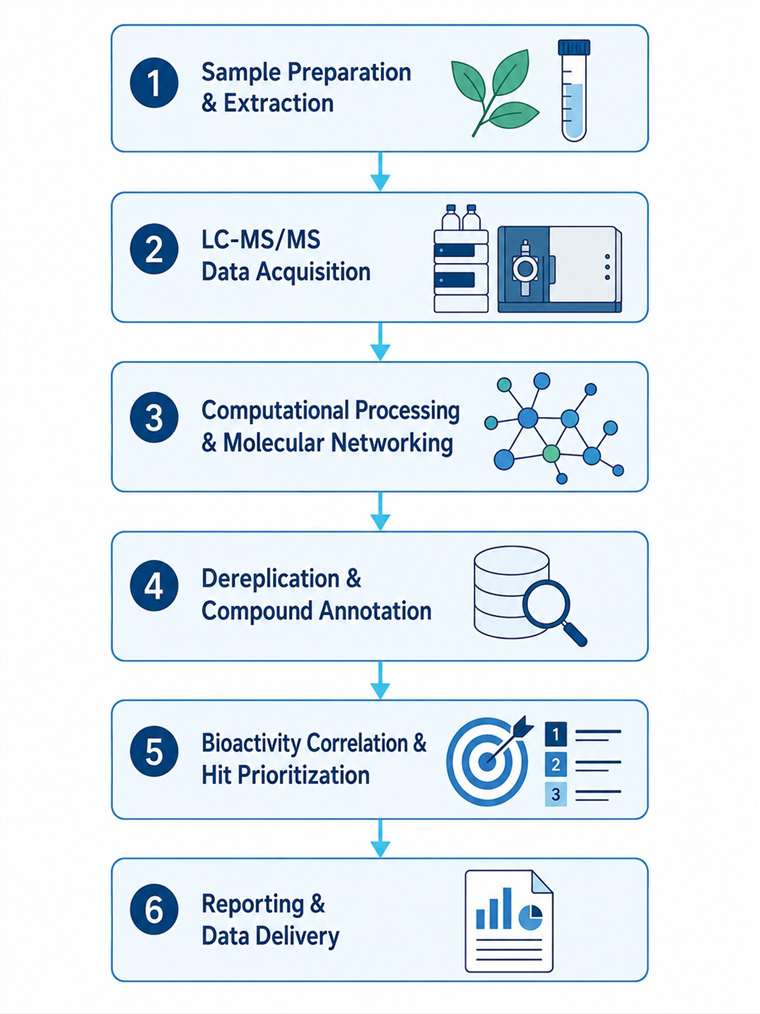
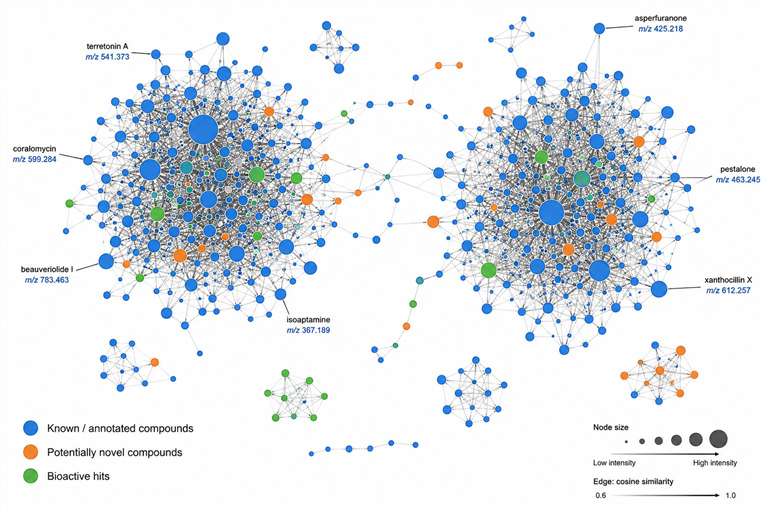
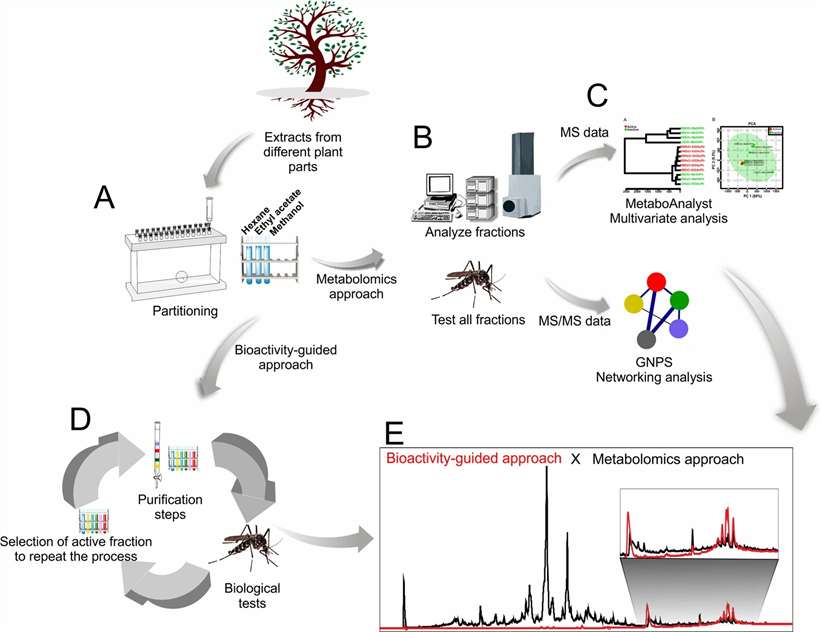
Our Natural Product & Complex Mixture MS Discovery Service addresses this bottleneck head-on. By integrating high-resolution LC-MS/MS, ion mobility MS, and affinity selection-MS with advanced computational workflows — including GNPS molecular networking, feature-based molecular networking, MS2LDA substructure discovery, and SIRIUS/CSI:FingerID — we enable rapid dereplication, structural annotation, and bioactive compound prioritization directly from complex mixtures. This MS-centric approach drastically reduces the iterative fractionation cycles required by traditional bioassay-guided methods, accelerating the journey from extract to active compound.
Whether you are working with plant extracts, marine organism samples, microbial fermentation broths, or synthetic complex mixtures, our platform delivers publication-quality data with comprehensive bioinformatics support.
- LC-HRMS/MS with GNPS molecular networking
- Integrated dereplication and structural annotation
- Affinity selection-MS for bioactive compound identification
- Multi-platform MS (Orbitrap, Q-TOF, Ion Mobility)