MS2LDA (Substructure Discovery) Service — Uncover Hidden Molecular Patterns from MS/MS Data
Go beyond spectral library matching. Automatically discover recurring fragmentation patterns that reveal the molecular substructures within your unknowns.
Every untargeted metabolomics experiment generates thousands of MS/MS spectra, yet standard library matching typically annotates fewer than 10% of features. The rest remain structurally uncharacterised — not because the data lacks information, but because the fragmentation patterns that encode substructure information have not been systematically extracted.
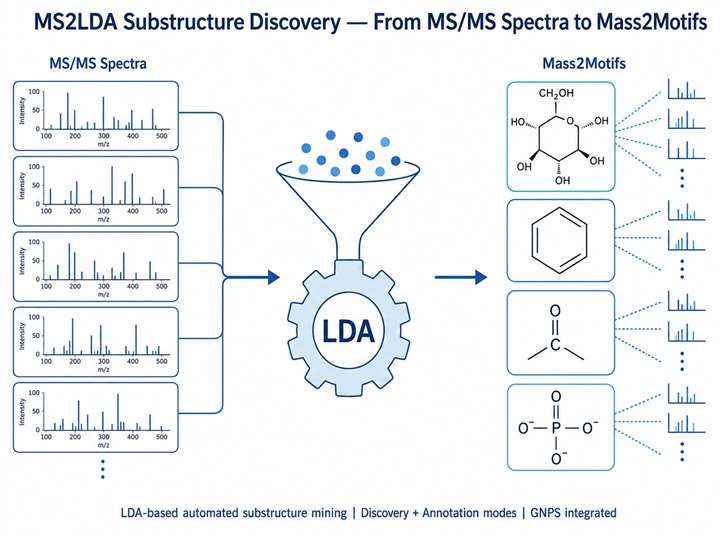
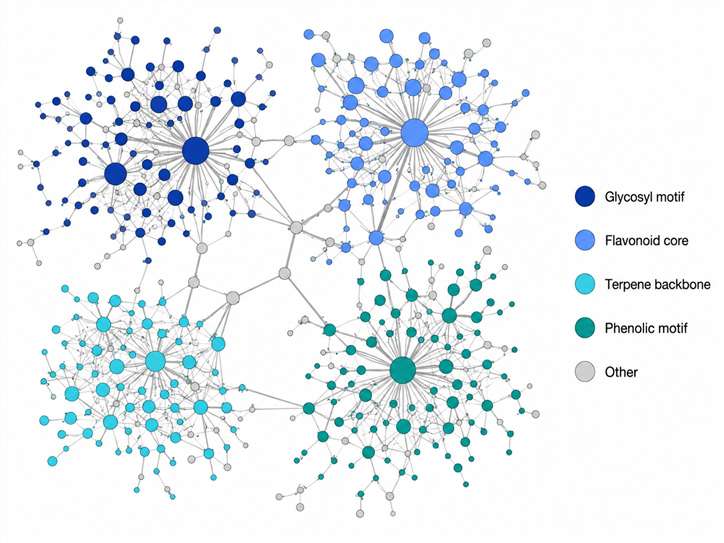
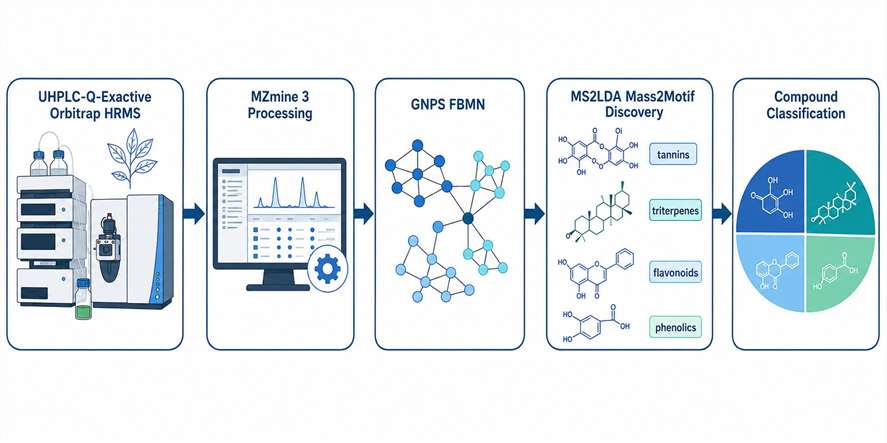
Our MS2LDA substructure discovery service applies Latent Dirichlet Allocation (LDA) topic modeling — a machine learning technique originally developed for text mining — to your MS/MS data. It automatically identifies Mass2Motifs: sets of co-occurring fragment peaks and neutral losses that represent chemically meaningful molecular substructures such as glycosyl groups, aromatic rings, carbonyl moieties, and other recurring scaffolds. Integrated with feature-based molecular networking and GNPS, this approach transforms raw fragmentation data into a chemically interpretable map of the substructures present in your sample.
Key Advantages:
- Discovers substructures without requiring a reference spectral library
- Reveals the substructure composition of every feature in your dataset
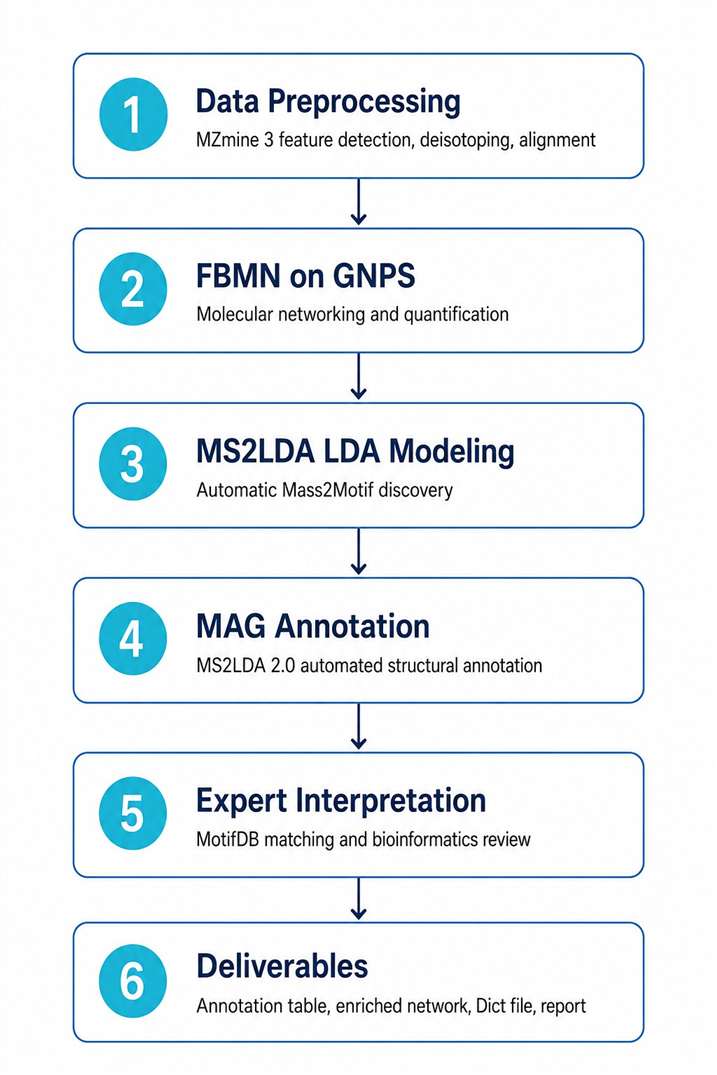
- Integrates seamlessly with FBMN and GNPS molecular networking workflows
- Supports both discovery mode (de novo motif mining) and annotation mode (MotifDB matching)
- MS2LDA 2.0 delivers up to 14× faster processing with automated Mass2Motif Annotation Guidance (MAG)