This literature-supported case study is based on Shamshurin et al., In situ activity-based protein profiling of serine hydrolases in E. coli. It is not a Creative Proteomics customer case.
Background
The study addressed whether activity-based protein profiling could monitor functional enzyme-family states directly in intact cells. The authors focused on serine hydrolases because members of this family share catalytic features that can be targeted by fluorophosphonate-based activity probes.
The paper also highlights a common limitation of gel-based ABPP: visible gel bands do not always identify the labeled enzyme. The authors therefore used LC-MS/MS after enrichment to identify active serine hydrolases more directly.
Methods
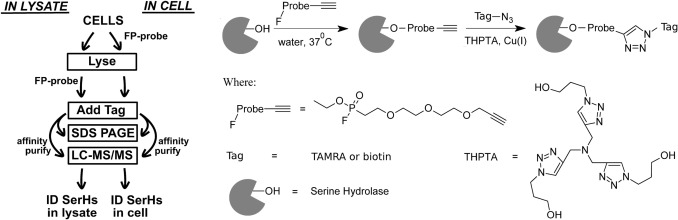
The authors used E. coli K12 cells and a fluorophosphonate-alkyne activity probe to label active serine hydrolases. The probe could react with active serine hydrolases either in intact cells or after cell lysis.
After labeling, the alkyne handle was connected to biotin-azide or TAMRA-azide using CuAAC click chemistry. TAMRA-tagged samples were analyzed by SDS-PAGE fluorescence, while biotin-tagged proteins were enriched with streptavidin and identified by LC-MS/MS.
The study compared in-cell labeling and in-lysate labeling. Biological duplicates were prepared for each labeling condition. Figure 1 shows the experimental design, including in-cell and in-lysate FP-alkyne labeling, click tagging, affinity enrichment, and LC-MS/MS identification. Figure 2 shows fluorescence and Coomassie-stained gel patterns after FP-TAMRA labeling. Table 1 lists the serine hydrolases identified by ABPP enrichment.
Results
The authors first showed that FP-TAMRA labeling generated detectable protein bands in both in-cell and in-lysate samples. In Figure 2, Coomassie-stained total protein patterns were similar across the in-cell, in-lysate, and control samples. This suggested comparable protein loading and no gross disruption of cellular protein composition by probe treatment.
The LC-MS/MS data provided the main family-level result. The authors reported that whole-cell proteomic analysis detected 44 serine hydrolases in E. coli. The FP-alkyne ABPP workflow identified 30 of these serine hydrolases after enrichment, which corresponds to about 68% of the detectable serine hydrolase family in that system.
The study also reported broader proteomic depth. Using 2D-LC-MS, the authors identified 2,496 proteins with two or more unique peptides, representing about 60% of the 4,141 proteins encoded by the E. coli genome. The false-positive rates for ABPP LC-MS/MS searches ranged from 0.7% to 0.93%. The dataset was deposited as PXD000241.
The comparison between in-cell and in-lysate labeling also showed that labeling context matters. The authors observed qualitative differences between the in-cell and lysate labeling patterns. This supports the point that family-specific ABPP results depend on sample state, probe access, and cellular context.
Conclusion
The authors concluded that FP-alkyne ABPP can profile active serine hydrolases in situ and identify a substantial portion, but not all, of the detectable family members. The study shows why enzyme-family-specific ABPP should be interpreted with probe coverage, sample context, enrichment, and LC-MS/MS evidence in mind.
For enzyme-family ABPP projects, this case supports four practical lessons. First, a family-directed probe can produce a focused activity profile across many related enzymes. Second, LC-MS/MS identification provides information that gel-based readouts alone cannot provide. Third, in-cell and lysate labeling can differ, so sample context should be chosen based on the biological question. Fourth, non-detection should not be over-interpreted as lack of enzyme activity because probe coverage is substantial but not complete.