Metabolic Pathway Drug-Response Mapping by LC-MS
Elucidating drug mechanism of action through pathway-level metabolomic profiling.
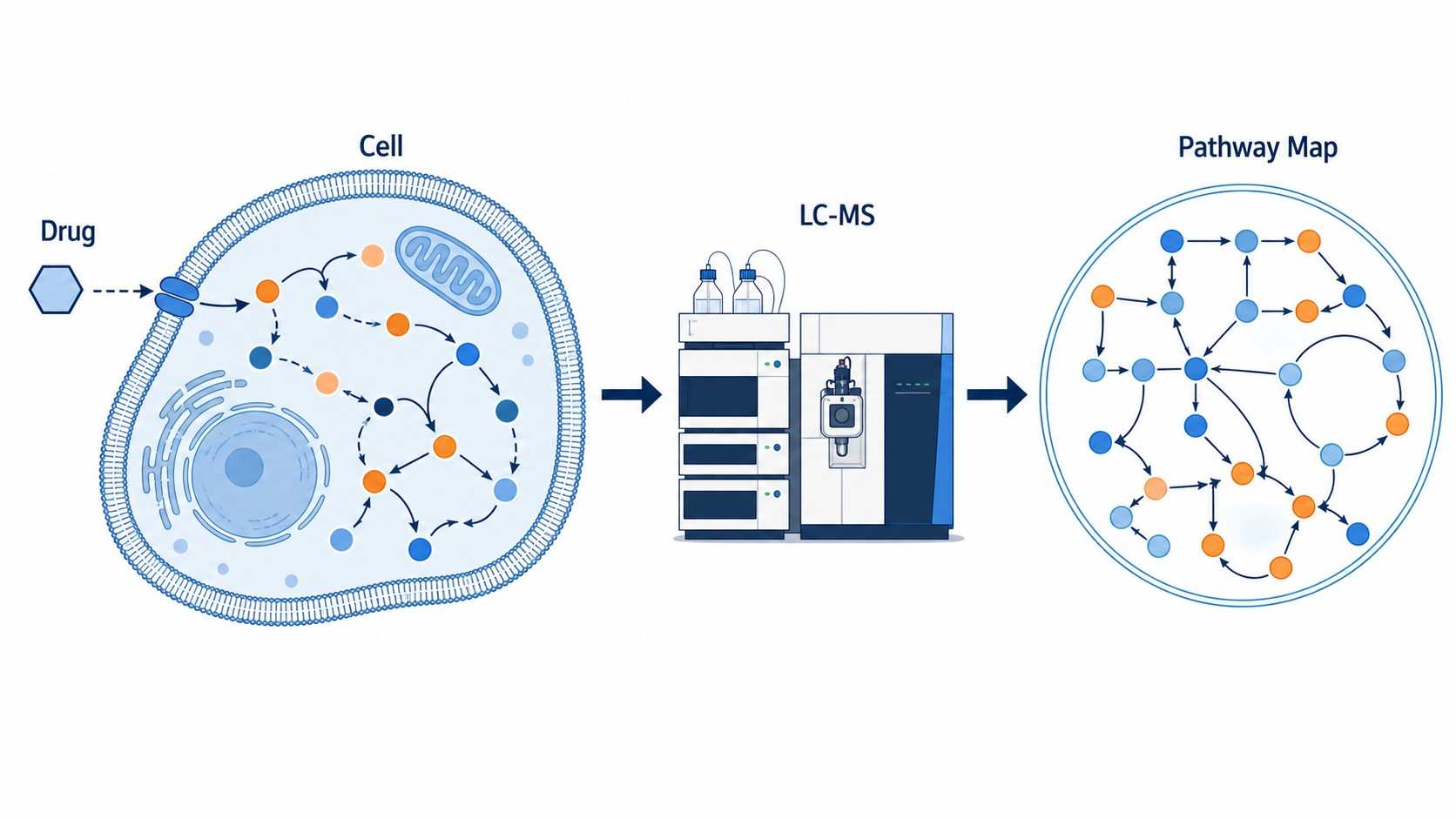
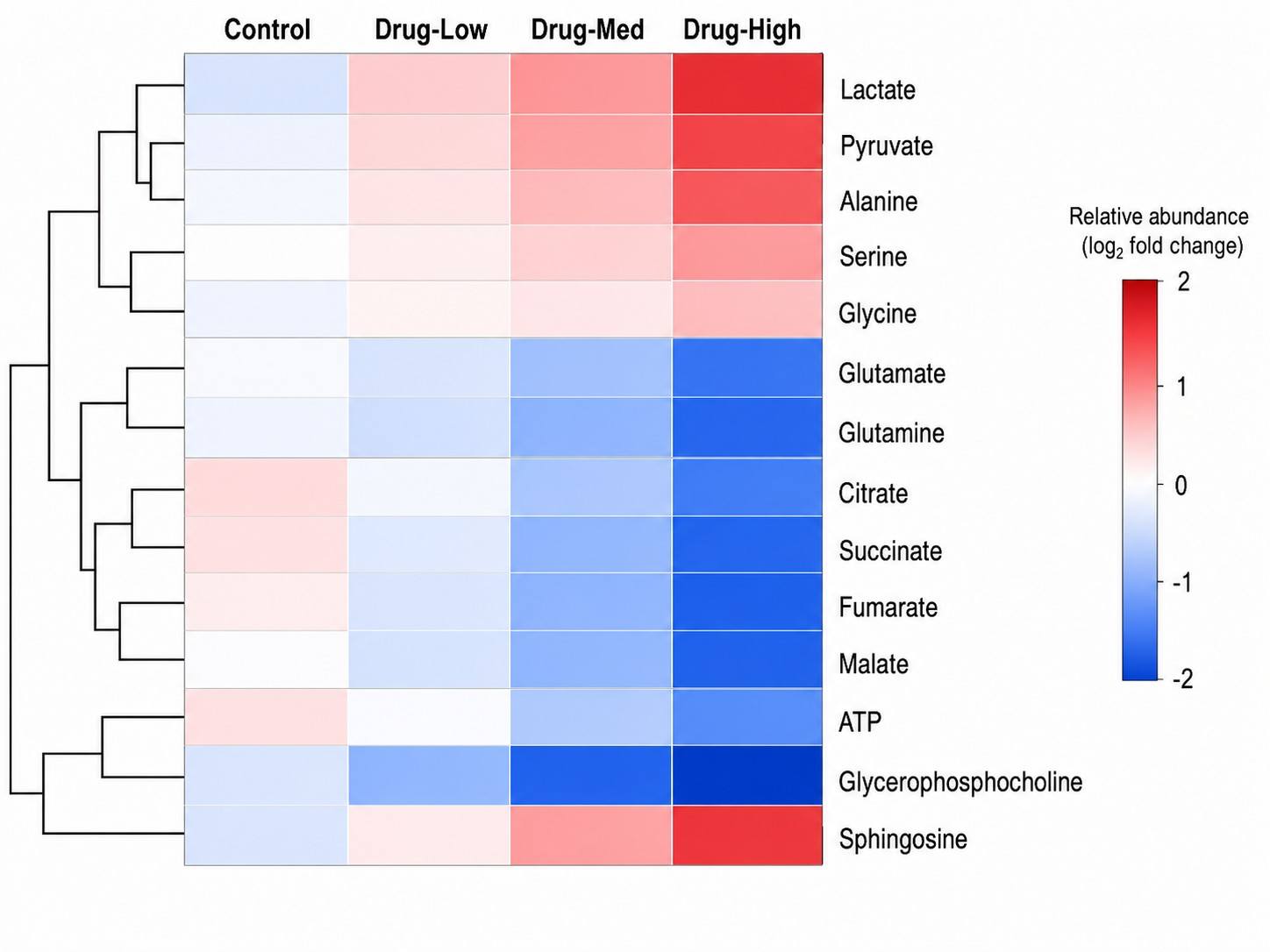
Metabolic Pathway Drug-Response Mapping is an LC-MS-based metabolomics service that profiles global metabolite changes in response to drug treatment, maps these changes onto metabolic pathways, and identifies the specific pathways and metabolites modulated by the compound.
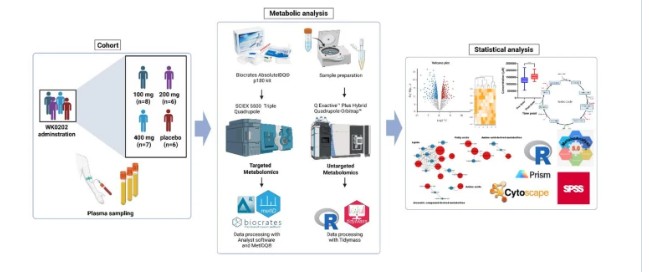
At Creative Proteomics, our MassTarget™ platform integrates high-resolution LC-MS metabolomics with curated pathway analysis pipelines (KEGG, HMDB, MetaboAnalyst) to provide a complete picture of drug-induced metabolic rewiring. Whether you are characterizing a novel compound's MoA, investigating metabolic resistance in oncology models, or screening for off-target metabolic liabilities, our service transforms raw metabolomics data into actionable pathway-level conclusions.
Key Capabilities:
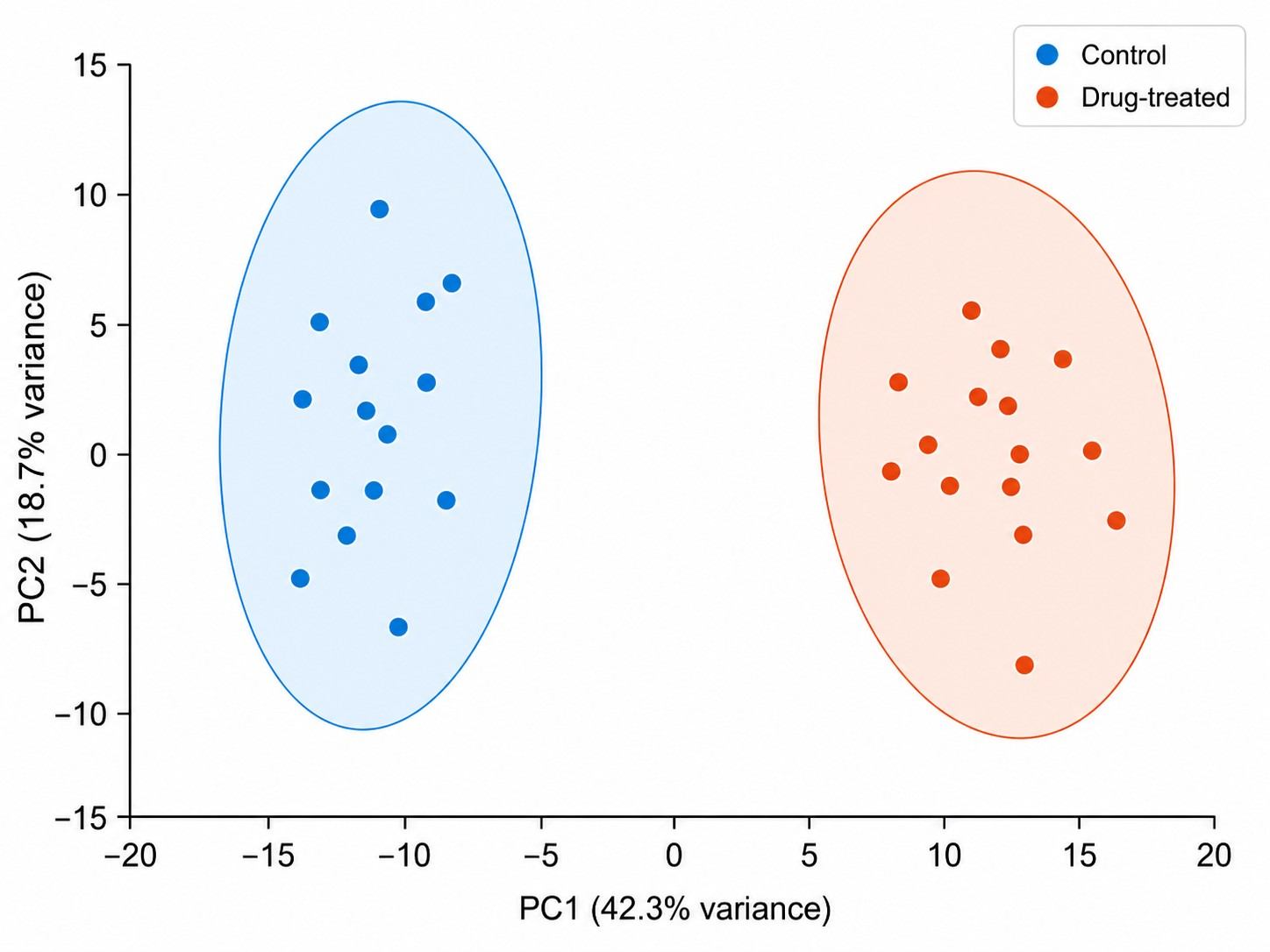
- Untargeted LC-MS metabolomics covering polar metabolites, central carbon metabolites, amino acids, nucleotides, and lipid species.
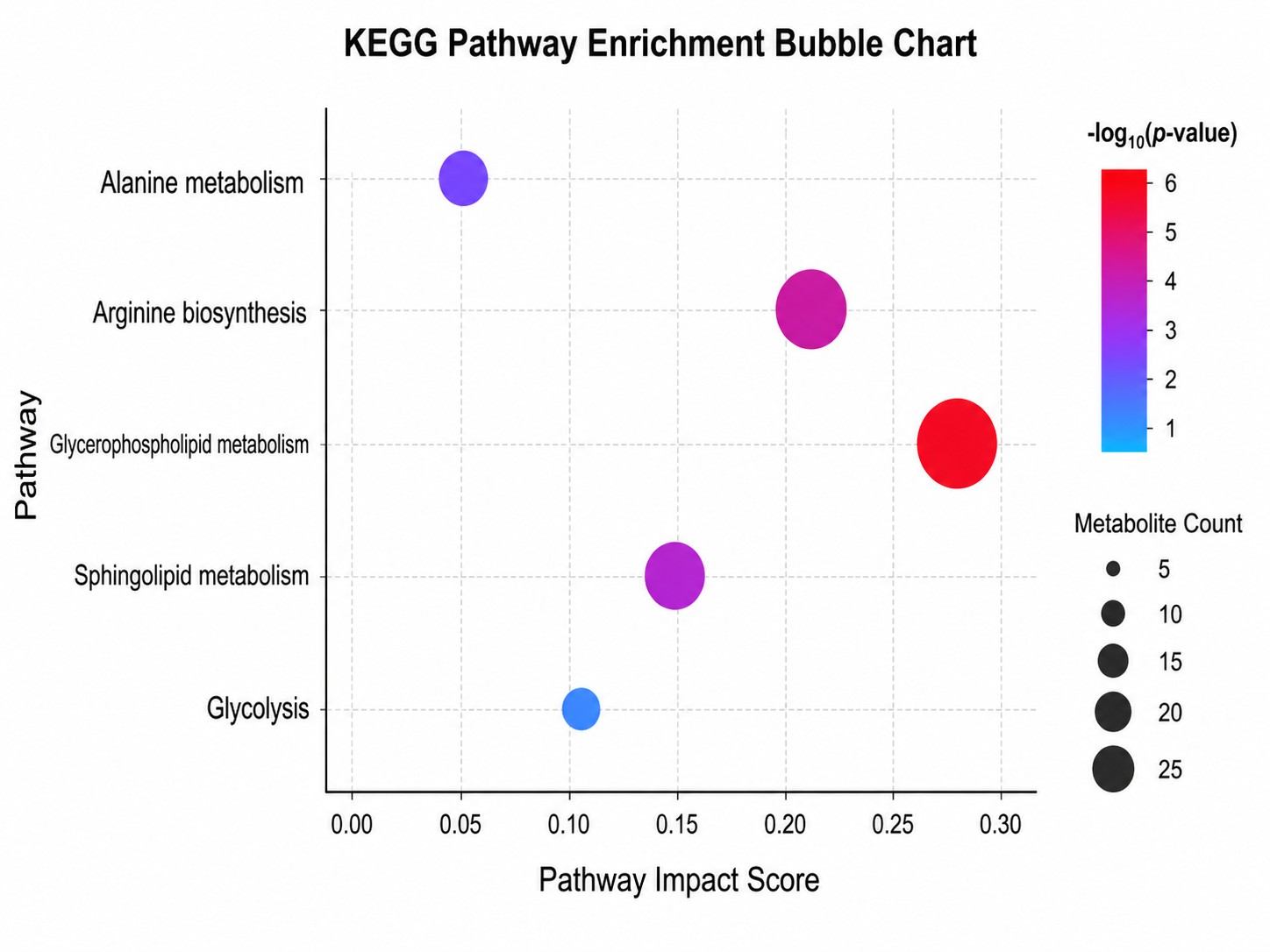
- Comprehensive pathway enrichment analysis with KEGG and HMDB mapping.
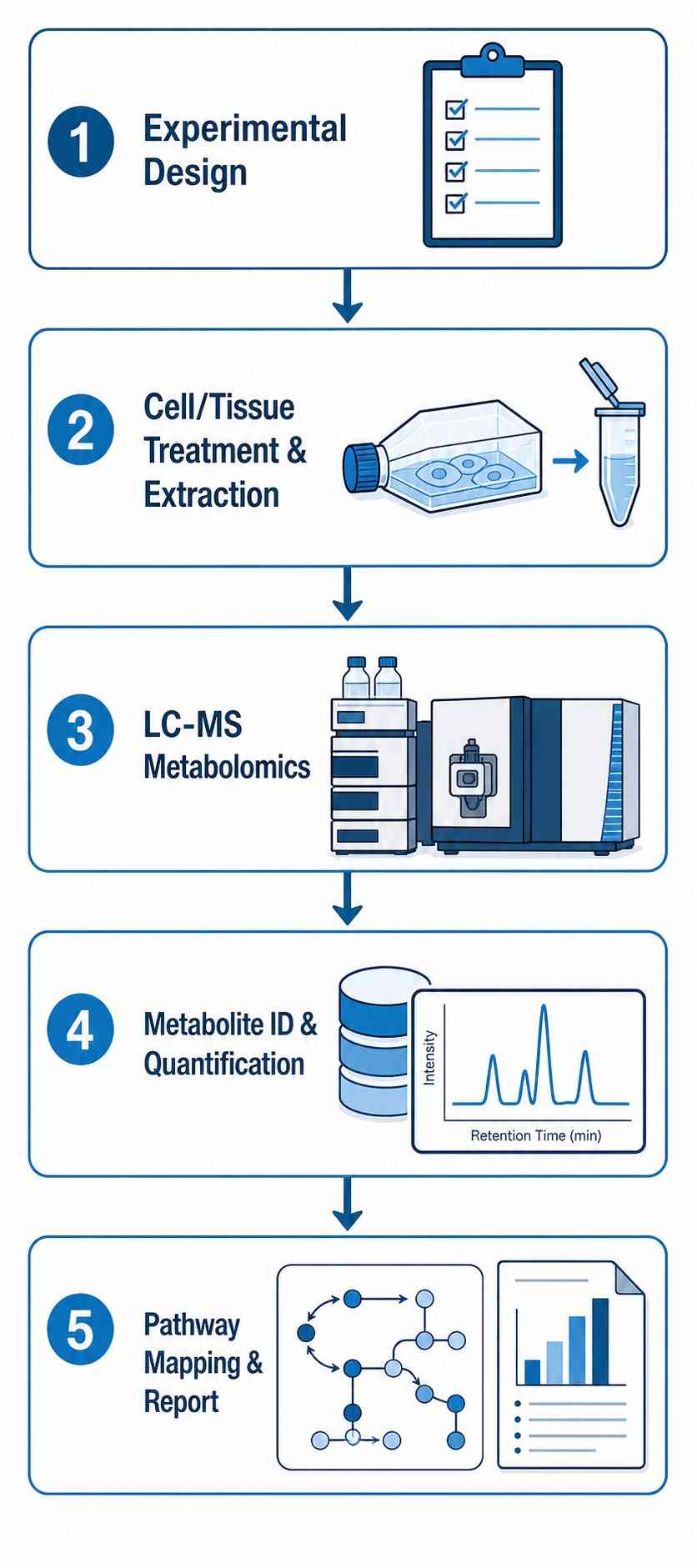
- Time-course and dose-response experimental design expertise.
- Rigorous QC with pooled QC samples, internal standards, and batch correction.
- Integration with proteomics and lipidomics for multi-omics pathway analysis.