MS‑Based Proteome‑Wide Thermal Stability Profiling Service
Unlock drug‑target insights without licensing constraints.
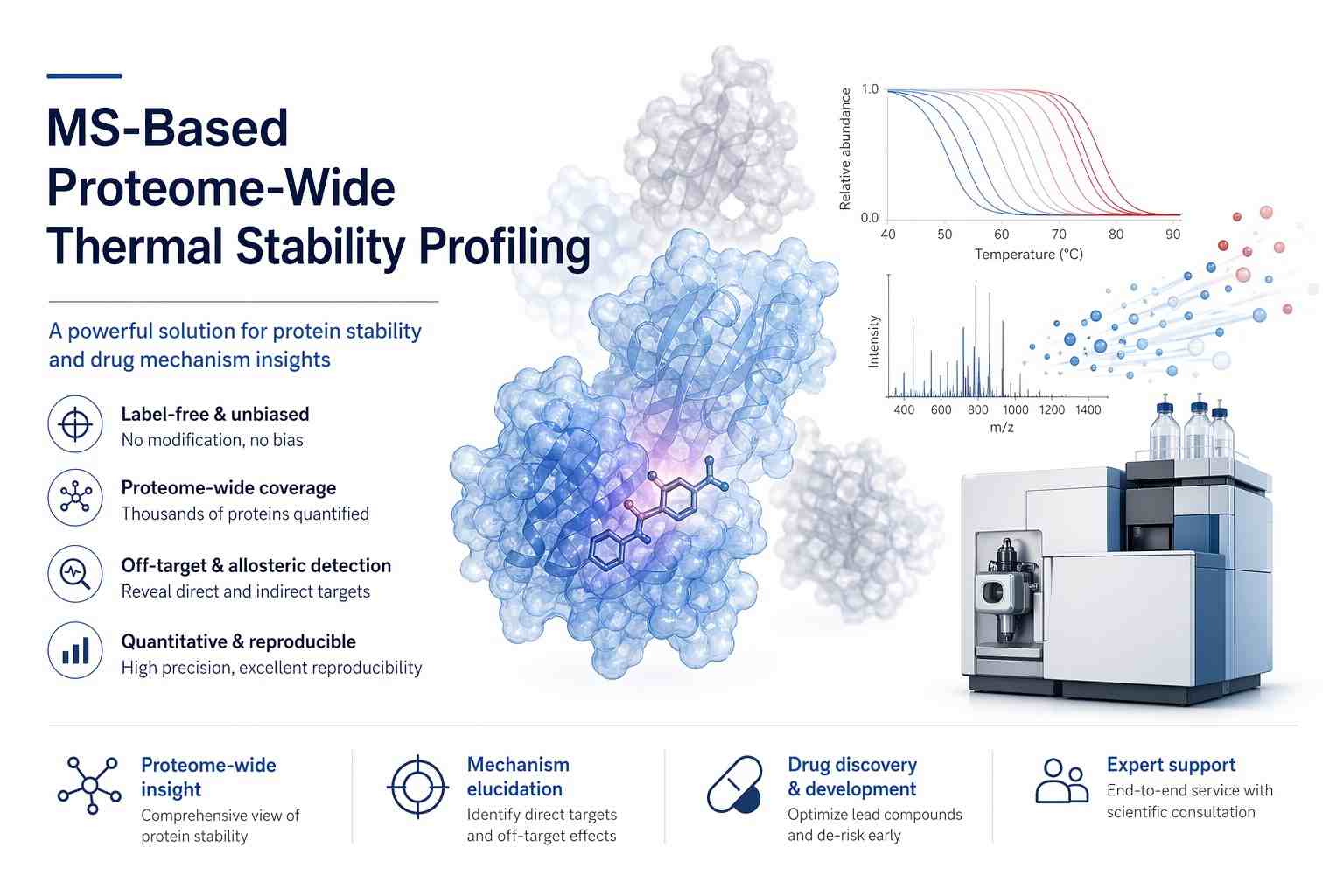
Our proteome‑wide thermal stability profiling combines precise thermal or proteolytic perturbations with high‑resolution mass spectrometry to monitor stability shifts across thousands of proteins. This label‑free approach reveals direct binding, off‑target effects and allosteric modulation in native cellular contexts.
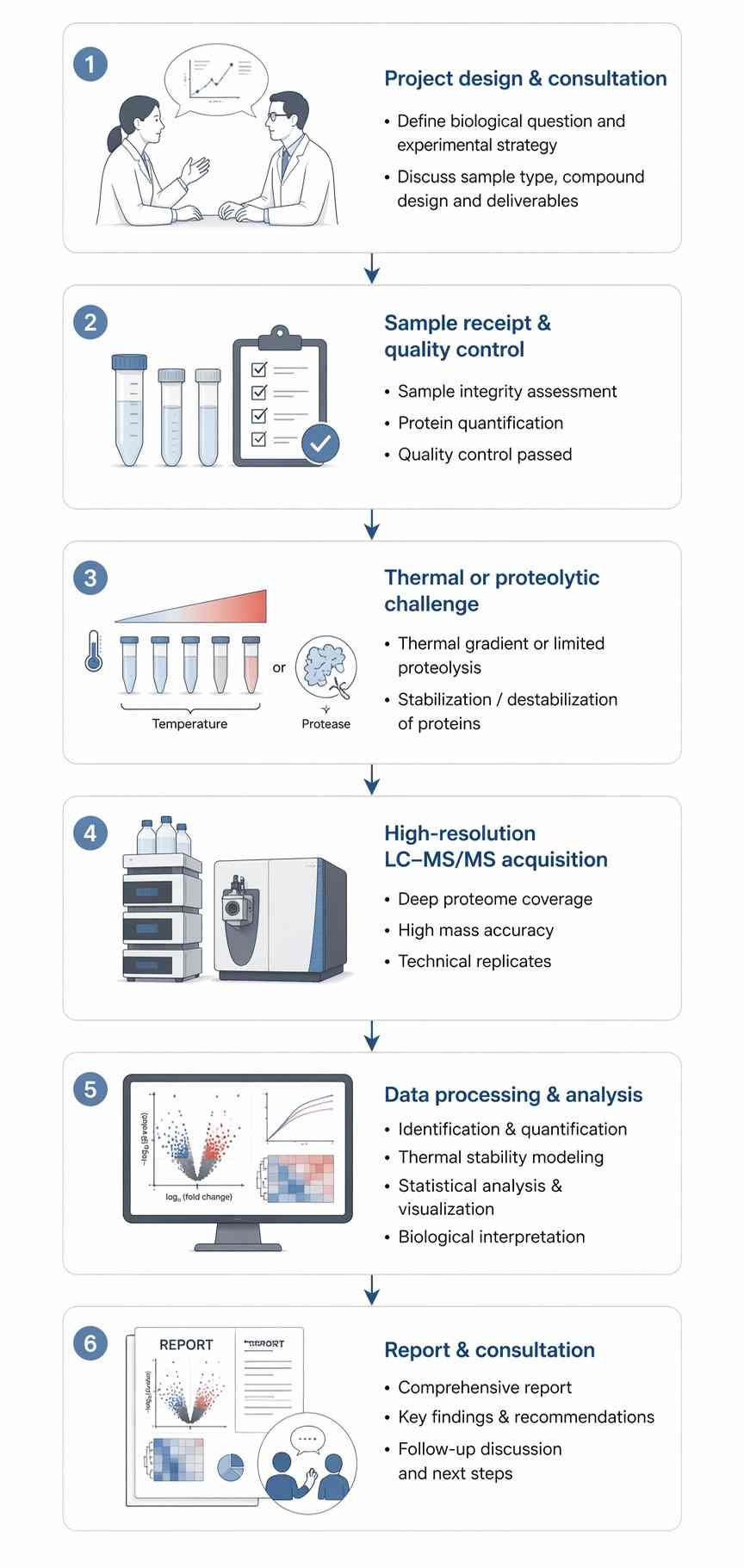
By partnering with Creative Proteomics, you gain access to a turnkey workflow for phenotypic hit deconvolution, off‑target profiling and allosteric site discovery when functional assays are unavailable or purification is challenging.
Key Advantages:
- Label‑free, proteome‑wide coverage
- Detect off‑targets & allosteric binders
- High‑resolution MS & advanced analytics
- Flexible sample compatibility (cells, tissues, complexes)
- Optional integration with LiP‑MS & dose‑response profiling