HDAC Activity MS
Label-free mass spectrometry-based histone deacetylase activity assays for direct, quantitative measurement of HDAC inhibition and selectivity.
At Creative Proteomics, our HDAC MS activity assay service is designed to address the challenges researchers face with conventional HDAC screening: compound interference in optical readouts, limited substrate flexibility, and the need for detailed kinetic characterisation alongside routine IC₅₀ determination.
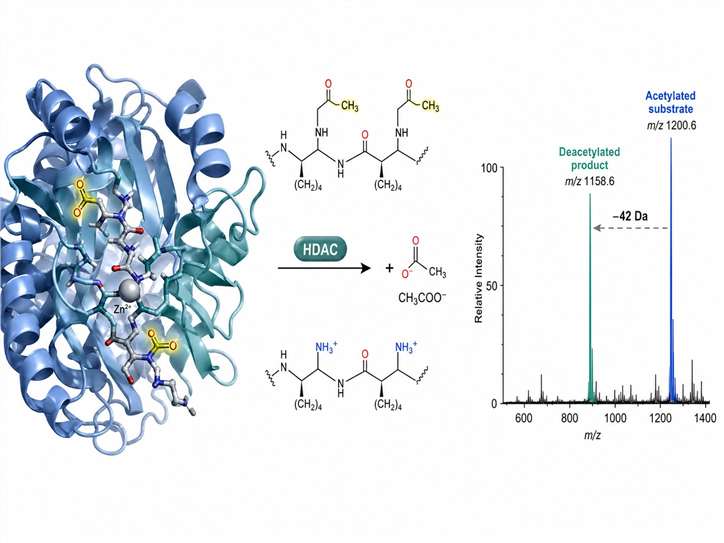
Our platform detects HDAC-catalysed deacetylation directly by MS, measuring the characteristic mass shift (−42 Da) as acetylated substrates convert to deacetylated products. No fluorescent labels, no colorimetric reagents, no coupled enzyme systems — just clean, interference-free data for your HDAC inhibitor screening programmes.
Key Advantages:
- Label-free detection — no compound autofluorescence or quenching interference
- Direct deacetylation readout — measure deacetylated product formation without coupled enzymes
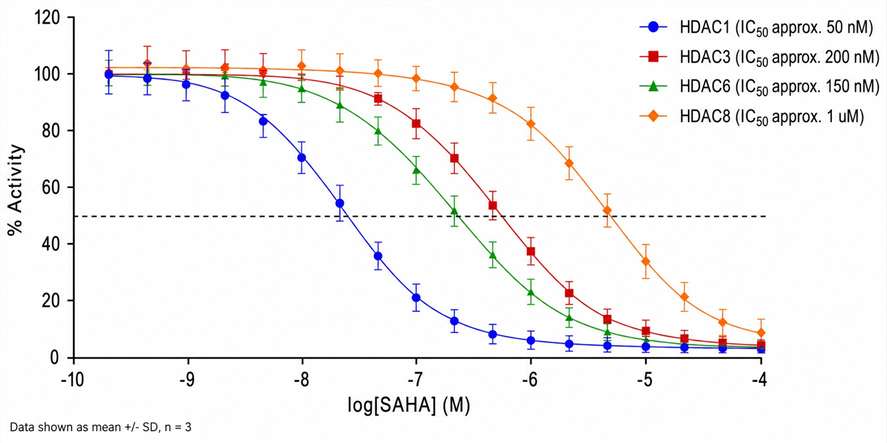
- IC₅₀, Kₘ, and mechanism-of-action from a single MS workflow
- Class I/II HDAC isoforms (HDAC1–11) supported
- Peptide and full-length protein substrates, including non-histone targets
- DMSO-tolerant at standard screening concentrations