GNPS Molecular Networking Service
Transform complex MS/MS data into actionable chemical insights with expert GNPS molecular networking — from raw spectra to publication-ready molecular networks.
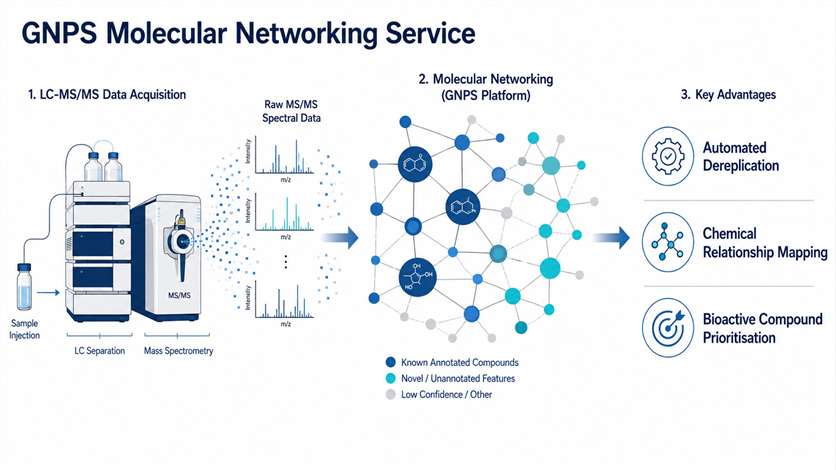
Natural product extracts, microbial fermentation broths, and complex biological mixtures routinely generate thousands of MS/MS spectra that are impractical to annotate manually. GNPS (Global Natural Products Social Molecular Networking) organises this data into molecular networks based on spectral similarity, giving you a visual map of every compound in your sample — and how they relate to each other.
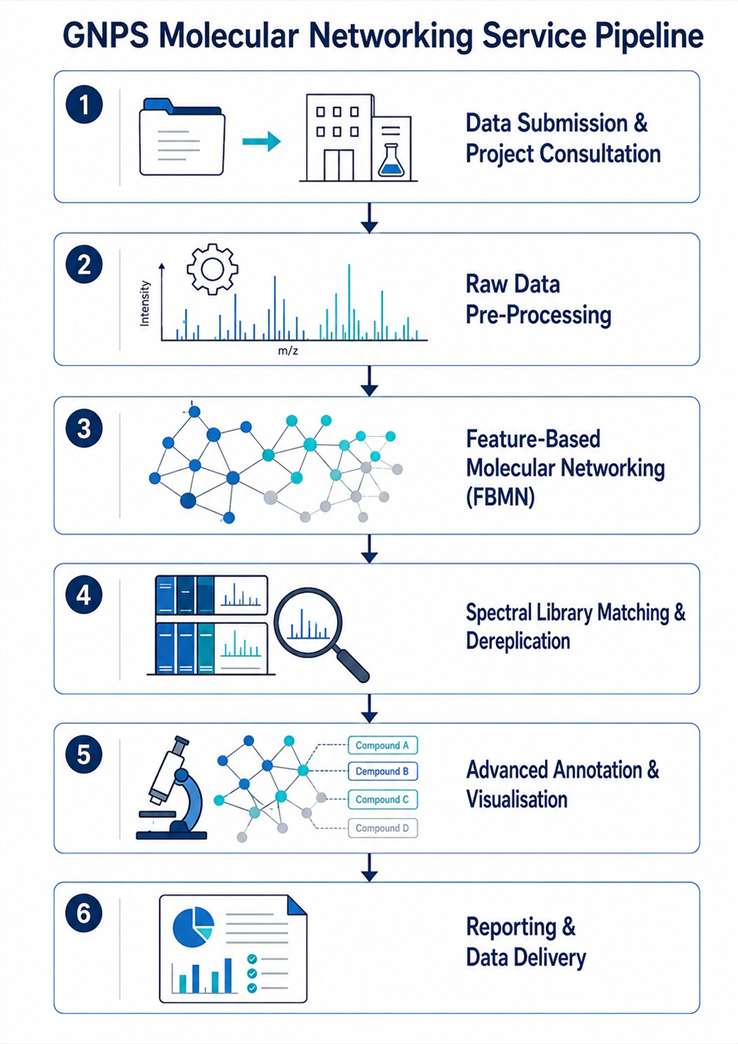
We handle the full computational pipeline: raw data pre-processing, feature-based molecular networking (FBMN), spectral library matching against GNPS and in-house databases, and expert interpretation of the resulting networks. The output is a fully annotated molecular network with a comprehensive dereplication report — no software installation or bioinformatics expertise required on your end. We work with plant extracts, microbial cultures, marine organism samples, fractionated libraries, and bioassay-guided fractionation sets, across Orbitrap, Q-TOF, and timsTOF platforms.
For deeper dereplication capabilities, explore our dedicated LC-HRMS/MS dereplication service. For a complete discovery pipeline, see our Natural Product MS Discovery Service.