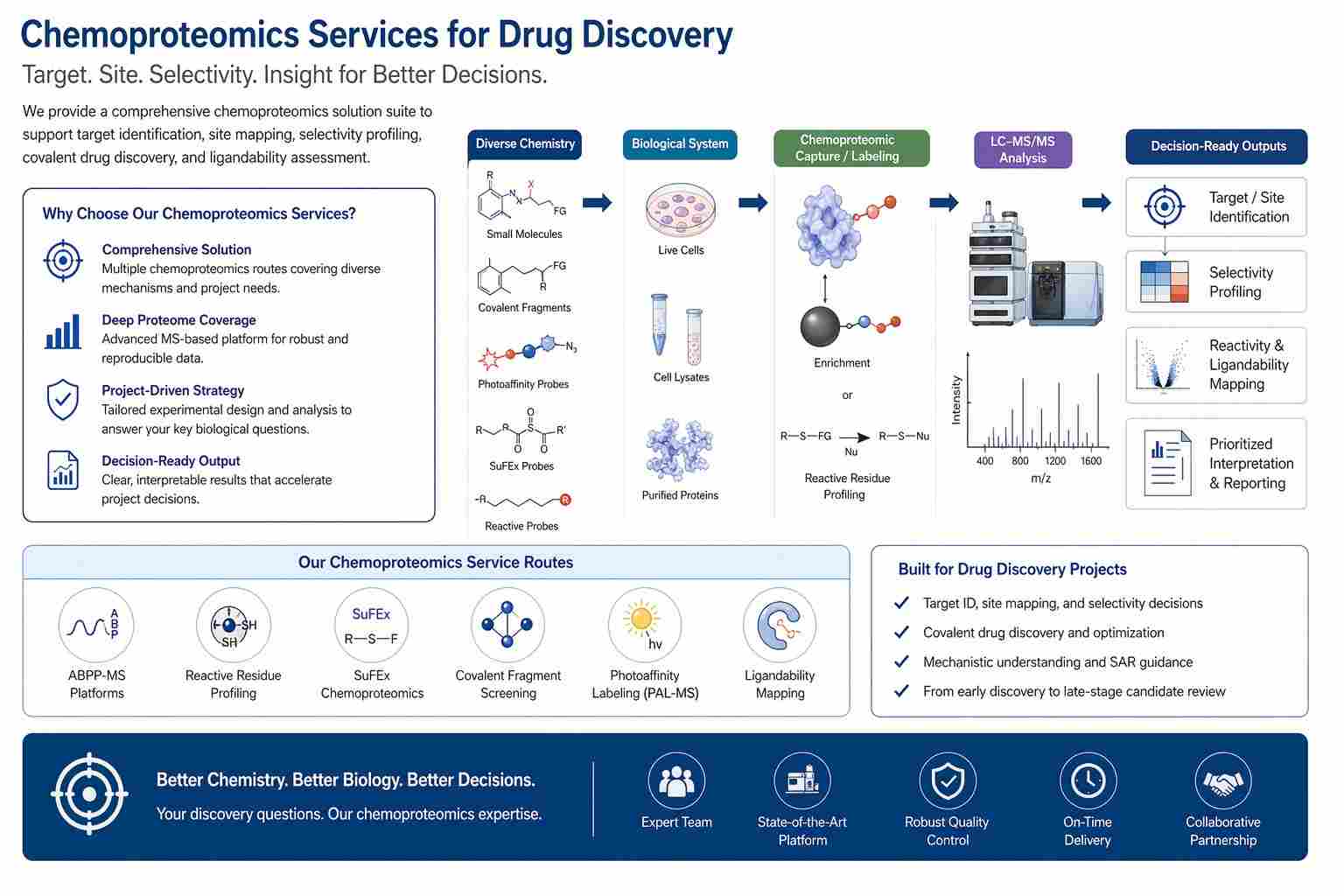
What Chemoproteomics Can Help You Solve in Drug Discovery
Chemoproteomics is most useful when you need to move beyond indirect biology and into direct molecular evidence. In practical terms, it helps answer questions such as: Which protein is my compound engaging? Which residue is involved? How selective is this chemistry across the proteome? Are there ligandable or reactive sites worth pursuing in a difficult target class? For many discovery teams, those are exactly the questions that determine whether a program moves forward, changes direction, or expands into a broader profiling effort.
That makes chemoproteomics valuable across target identification, site-level mechanism work, selectivity assessment, and early covalent discovery. It is not one fixed assay. It is a family of mass spectrometry-based strategies that can be routed according to the project goal, from activity-state profiling to photoaffinity capture to residue-centric mapping.
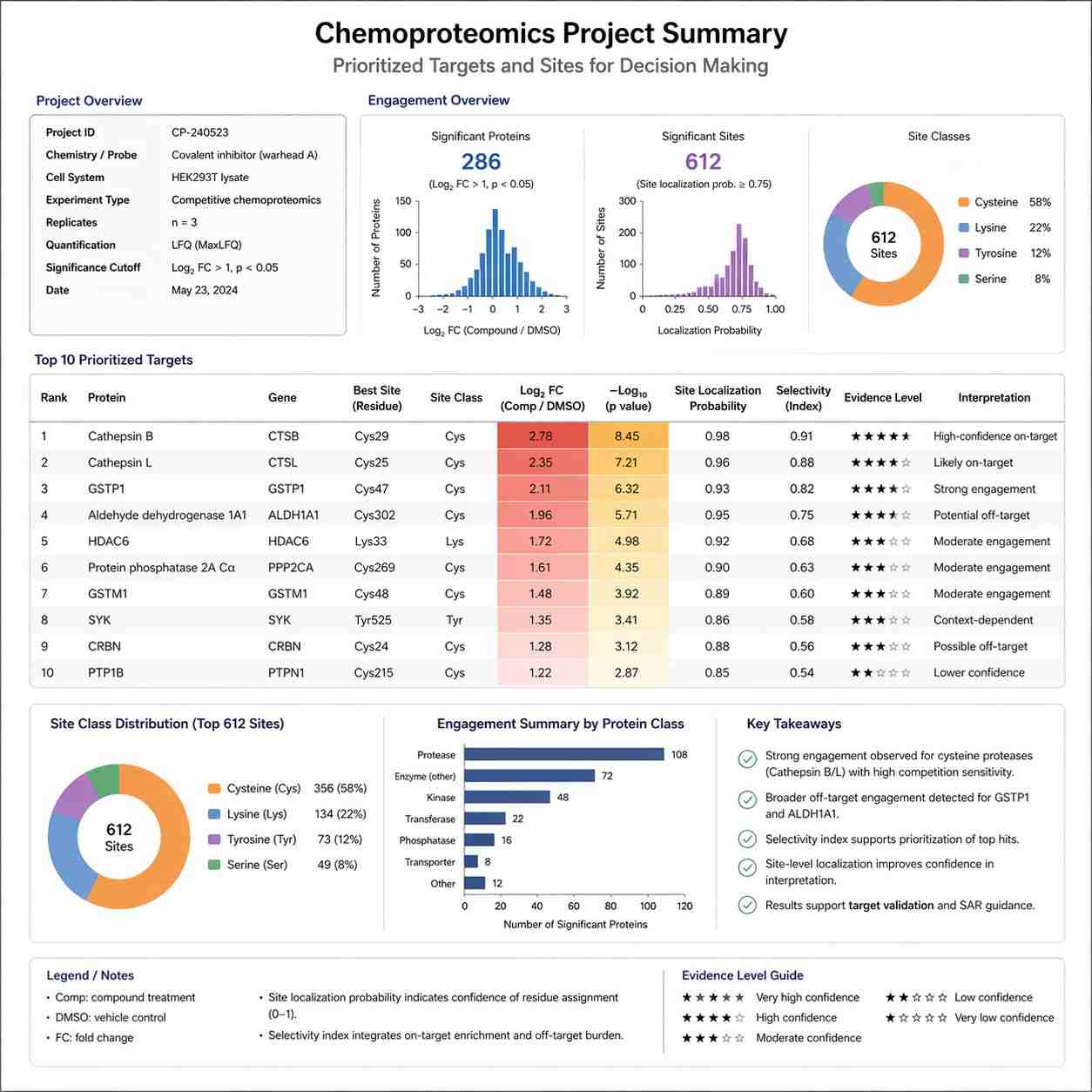
The main advantage is not simply that chemoproteomics generates more data. The real advantage is that it generates the right kind of data for the next decision. Instead of staying at the level of phenotype, pathway change, or indirect target hypotheses, your team can work with target-level, site-level, and competition-aware outputs that are far easier to use in a drug discovery setting.
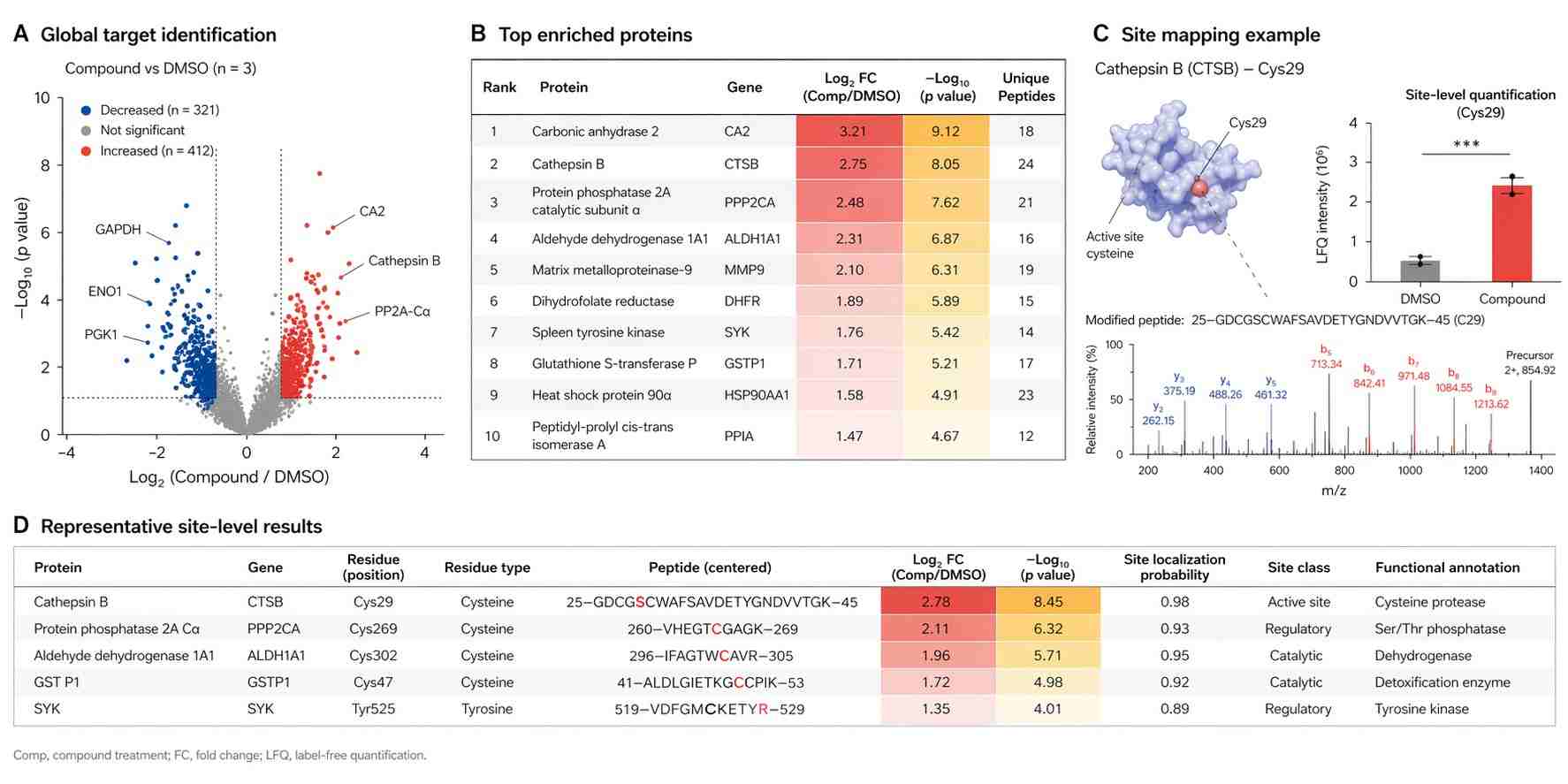
From target identification to site-level evidence
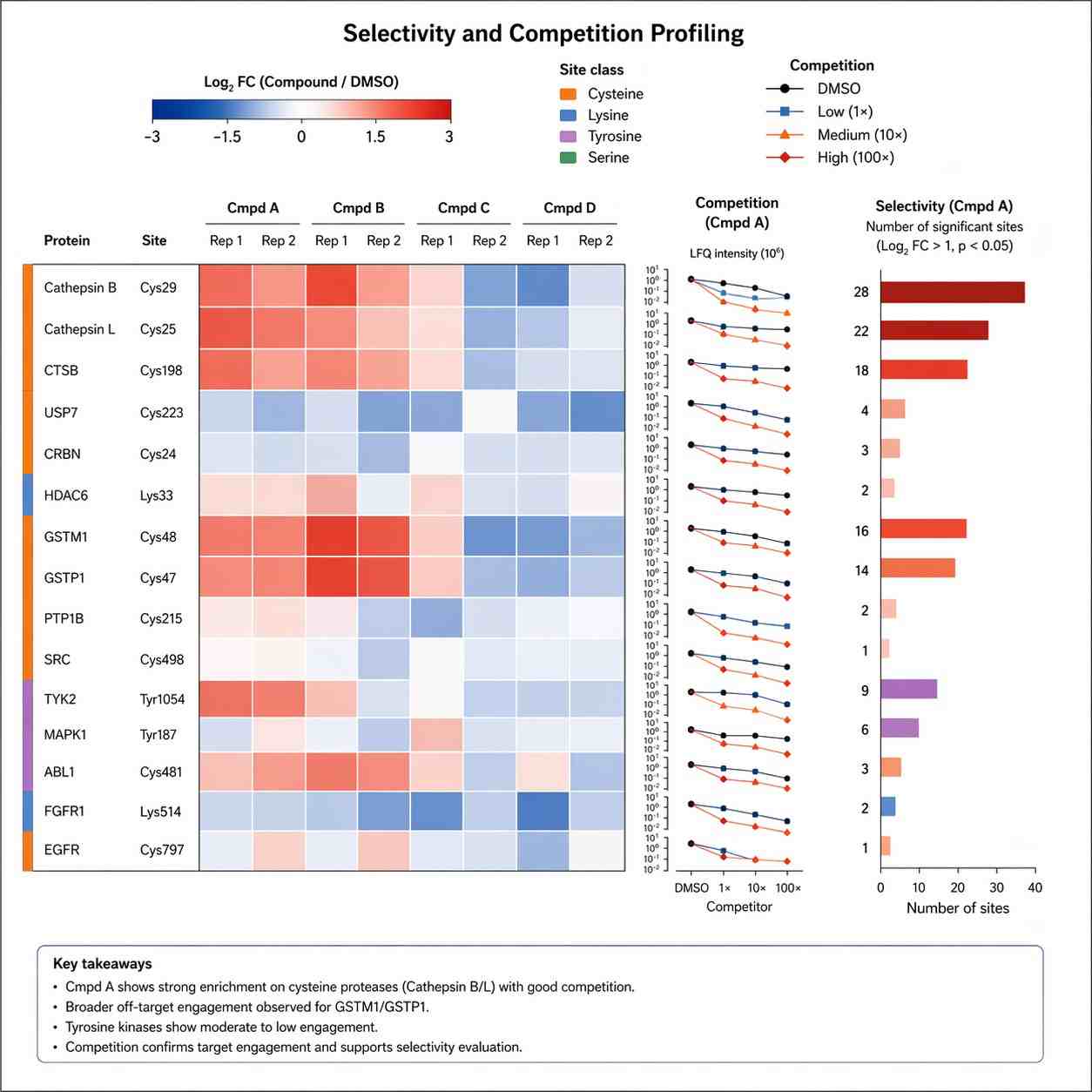
Some projects need to know which protein is engaged. Others need to know where the interaction occurs or whether one reactive chemotype behaves differently from another. Chemoproteomics gives us a route for both types of questions, which is why it is especially valuable for complex discovery programs.
Why chemoproteomics matters when functional data is not enough
A biological response does not automatically prove direct target engagement. At the same time, a weak phenotypic readout does not necessarily mean the chemistry is not worth pursuing. Chemoproteomics adds a direct molecular layer that helps separate those possibilities.
Where it fits in modern drug discovery
Chemoproteomics is especially relevant when you are working with covalent inhibitors, probes, fragment-derived chemotypes, target deconvolution problems, or projects where selectivity and ligandability matter as much as simple activity readouts.