UltraID Proximity Labeling Proteomics Service
Map proximal protein neighborhoods with compact biotin ligase–based proximity labeling and LC-MS/MS.
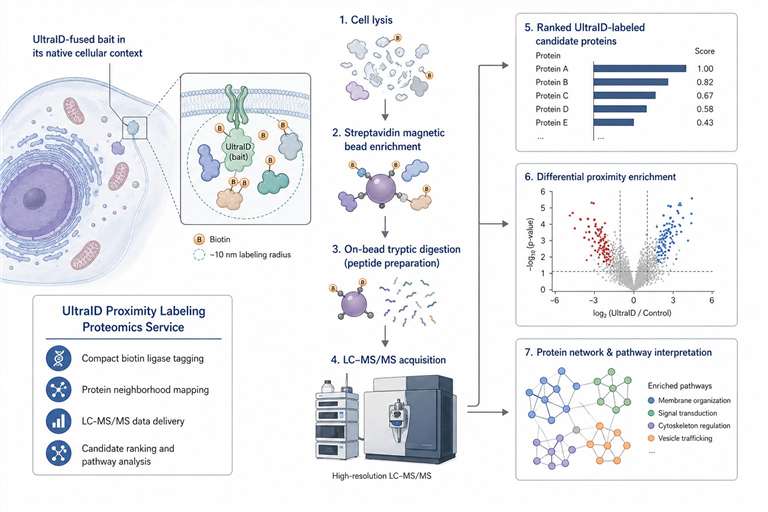
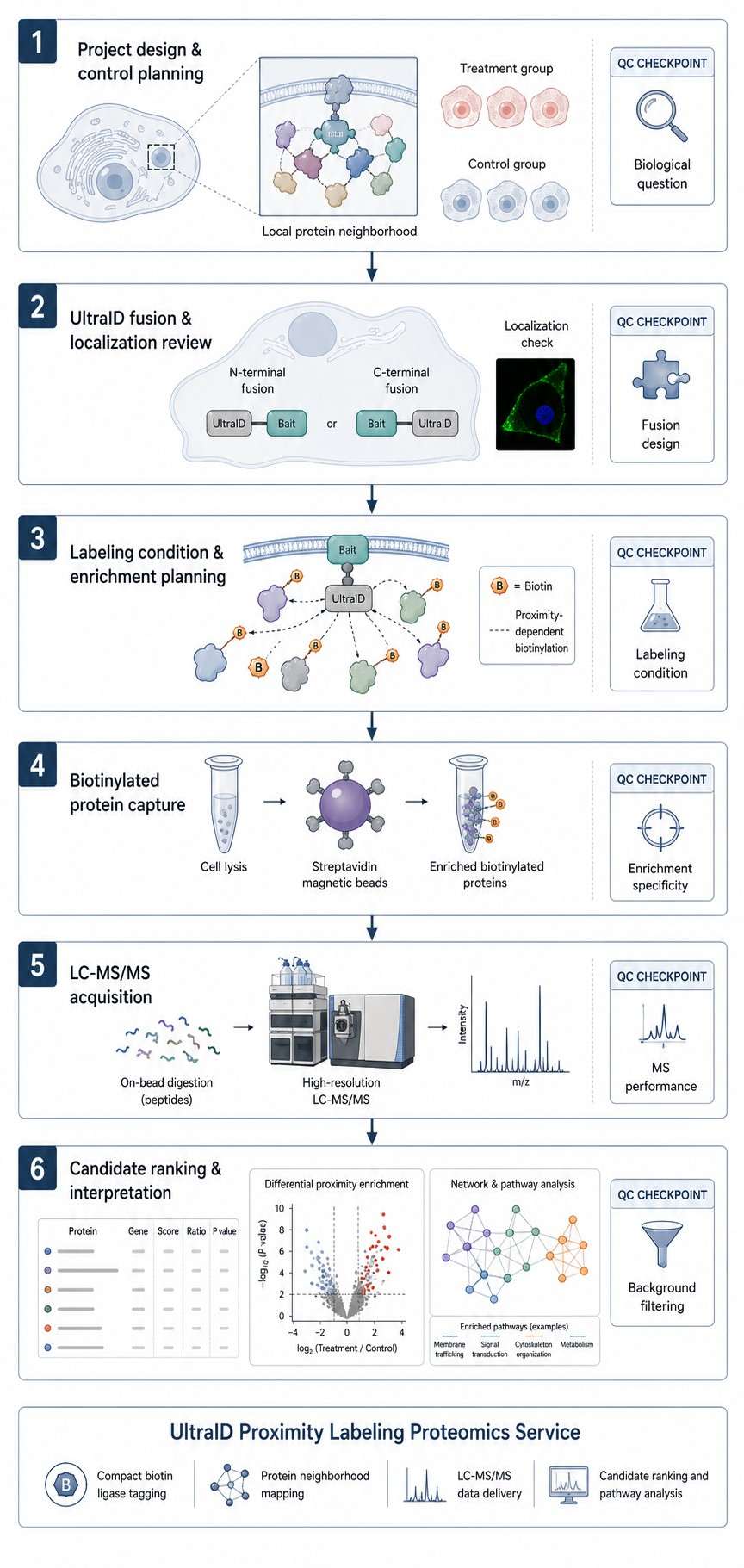
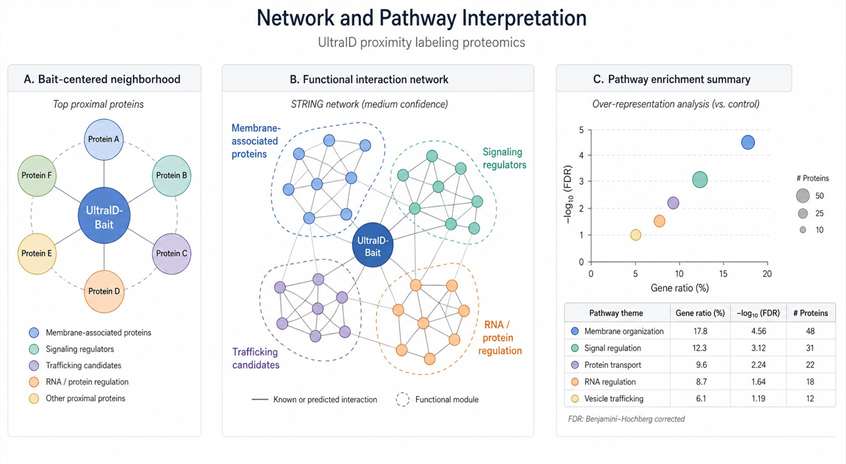
Creative Proteomics provides UltraID Proximity Labeling Proteomics Service for researchers who need compact biotin ligase–based protein-neighborhood mapping. We help you review UltraID fusion design, plan controls, enrich biotinylated proteins, acquire LC-MS/MS data, and interpret proximal candidates across bait-centered, membrane-associated, pathway, and perturbation studies.
UltraID is useful when you want to study proteins located near a bait, compartment, membrane-associated complex, or condition-dependent pathway state in living cells. Our service connects UltraID proximity labeling with enrichment proteomics, QC review, and bioinformatics interpretation, so you receive more than a protein list.
Service strengths:
- Map proximal protein neighborhoods around bait proteins or local cellular regions.
- Support compact UltraID fusion designs for proximity-dependent biotinylation.
- Review localization, controls, labeling conditions, and background risk.
- Analyze enriched biotinylated proteins by LC-MS/MS.
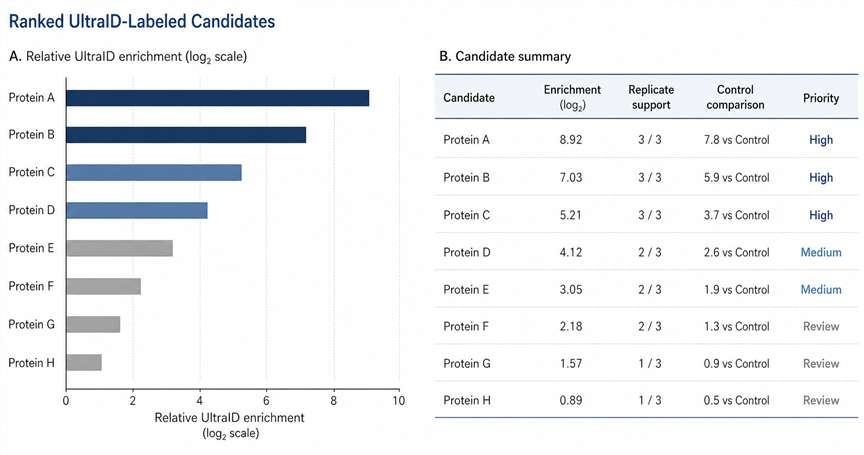
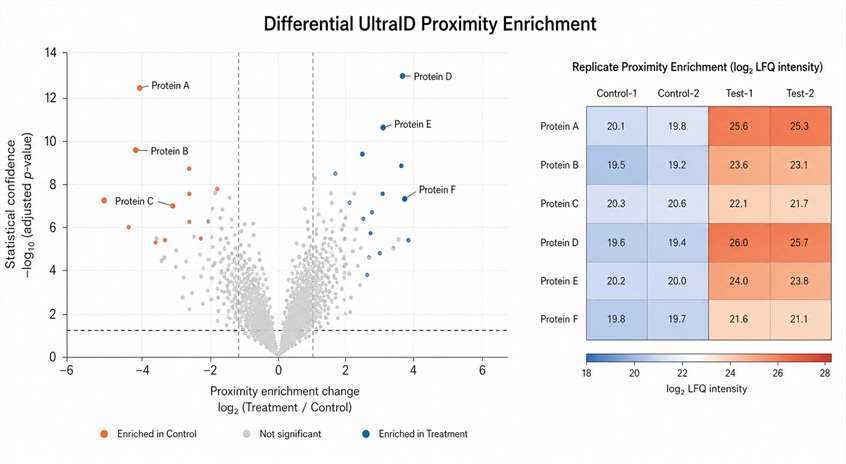
- Deliver candidate ranking, QC summaries, and pathway interpretation.