Background
APEX-based proximity labeling is valuable for subcellular proteomics because it can provide strong spatial and temporal control. However, conventional APEX2 activation can face background labeling and substrate-related concerns in some biological systems. This is especially important when the target compartment is small, difficult to isolate, or surrounded by proteins that can create nonspecific signal.
Sroka and colleagues addressed this challenge by developing iAPEX, an enzymatic cascade designed to locally generate hydrogen peroxide and improve specificity in APEX-based proximity labeling. The study tested this strategy across challenging biological contexts, including primary cilia proteomics.
Methods
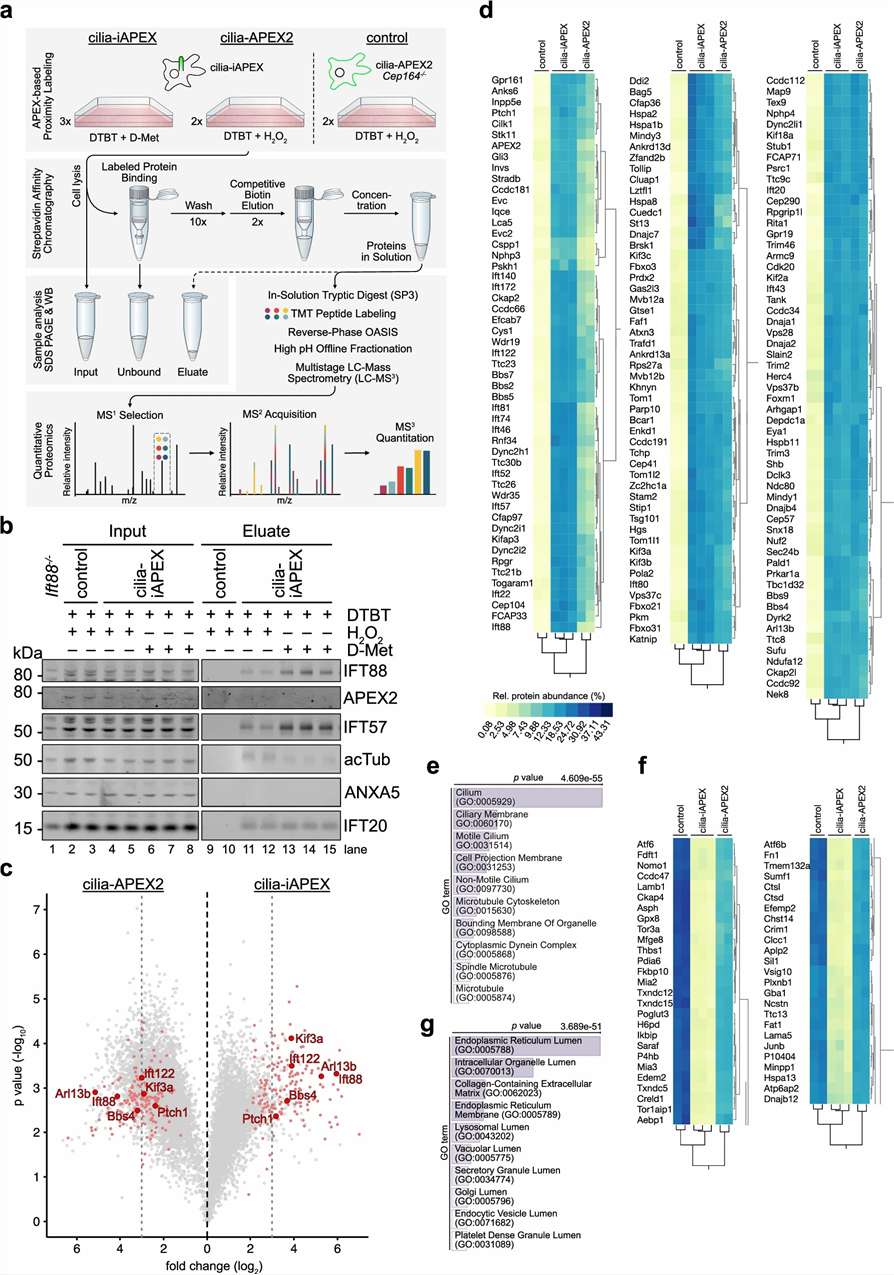
The authors combined APEX2 with D-amino acid oxidase to activate labeling locally. In the primary cilia experiment, they applied the workflow in IMCD3 cells, enriched labeled proteins, digested proteins, labeled peptides with TMT, and analyzed samples using LC-MS-based quantitative proteomics.
The study compared iAPEX with conventional APEX2-based labeling under related experimental conditions. It also used downstream filtering and GO-based interpretation to evaluate specificity and enrichment quality.
Results
Figure 4 reports quantitative primary cilia proteomics using iAPEX compared with a conventional APEX2-based system in IMCD3 cells. The study quantified 5,982 proteins in this experiment. Under stringent enrichment filtering, iAPEX identified 175 high-confidence candidate cilia proteins, while the conventional APEX2-based system produced 799 putative cilia proteins under the same cutoff.
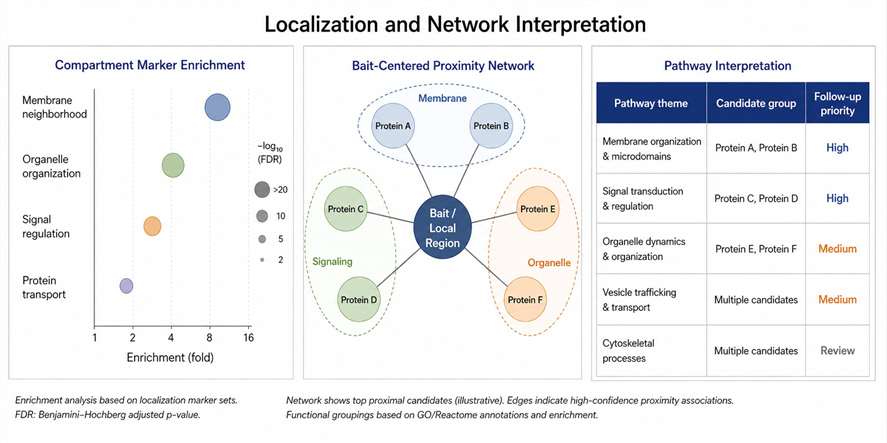
The authors showed that iAPEX separated known ciliary proteins from non-ciliary proteins more clearly than the conventional APEX2-based system. GO enrichment analysis also supported improved specificity for cilia-associated terms.
Conclusion
This case illustrates an important lesson for APEX proximity labeling projects: the labeling strategy, activation conditions, controls, and filtering logic strongly affect interpretability. For clients planning APEX/APEX2 proteomics, this supports the value of feasibility review, careful QC design, and bioinformatics filtering before drawing biological conclusions from candidate lists.