Cellular Lipidomics Drug Profiling
Decode drug-induced lipid remodeling at cellular resolution — comprehensive LC-MS/MS lipidomics profiling of drug-treated cells for MOA elucidation, lipid toxicity assessment, and resistance mechanism discovery.
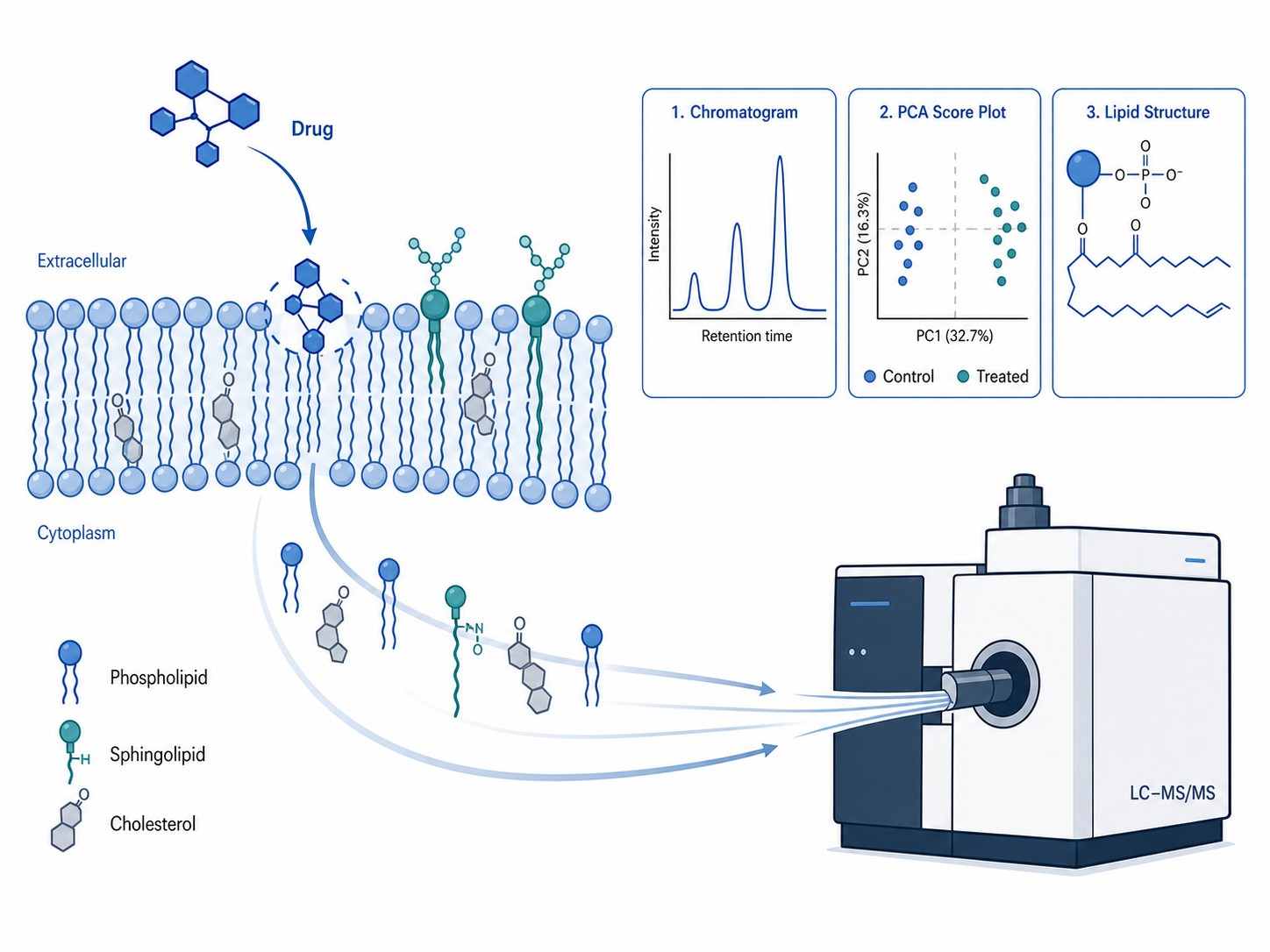
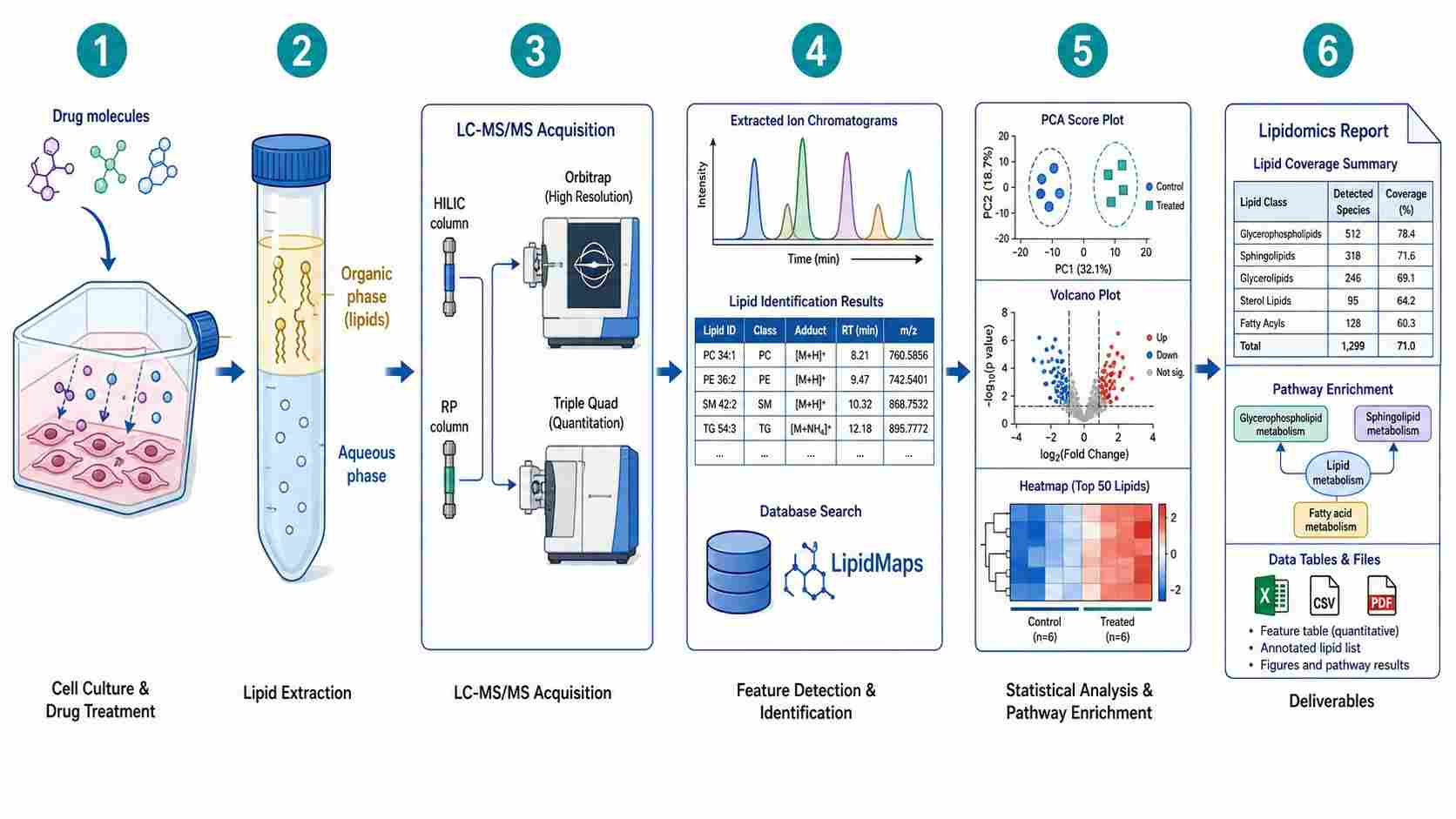
Cellular lipidomics drug profiling is a mass spectrometry-based analytical approach that comprehensively characterizes lipid remodeling in living cells following drug treatment. Unlike conventional biochemical assays that measure single lipid targets or bulk lipid content, cellular lipidomics captures the full lipid landscape — hundreds of individual lipid species across multiple structural classes — providing a systems-level view of how drugs perturb cellular lipid homeostasis.
Lipids are not merely structural components of cell membranes. They function as signaling molecules, energy storage reservoirs, and mediators of protein localization and activity. Drug-induced alterations in lipid composition can reveal mechanism of action (MOA), predict toxicity liabilities such as phospholipidosis and steatosis, and uncover adaptive resistance mechanisms that cancer cells deploy to survive chemotherapy. Despite this biological significance, lipidomics remains underutilized in early-stage drug discovery, largely because of the analytical complexity and the lack of CRO partners with dedicated drug-centric lipidomics workflows.
Key Capabilities:
- 500+ lipid species across 15+ classes per project
- Isomer-level resolution for major phospholipid and sphingolipid classes
- Relative (untargeted) and absolute (targeted, pmol/10⁶ cells) quantification
- Validated for cell lines, primary cells, 3D spheroids, and organoids
- Pathway-centric bioinformatics with lipid-specific KEGG and Reactome mapping
- Integrated with Cell-based MS Drug Screening platform (16 sub-services)